Abstract
The HtrA protein family represents an important class of serine proteases that are widely distributed across taxa. These evolutionarily conserved proteins are crucial for survival and function as monitors of protein synthesis during various stresses. Here, we performed gene expression analysis of the entire set of putative serine protease genes in Halothece sp. PCC7418 under salt stress conditions. The gene-encoding HtrA2 (H3553) was highly upregulated. This gene was cloned and functionally characterized, and its sub-cellular localization was determined. The recombinant H3553 protein (rH3553) displayed a pH optimum of 8.0, remained stable at 45 °C, and its proteolytic activity was not affected by salts. H3553 completely degraded the unfolded model protein, β-casein. In contrast, the folded model substrates (lysozyme or BSA) were not degraded by rH3553. Denaturation of BSA at a high temperature significantly increased its degradation by rH3553. H3553 was detected in the soluble protein fraction as well as the plasma membrane and thylakoid membrane fractions. Interestingly, the majority of H3553 was present in the plasma membrane under salt and heat stress conditions. Thus, H3553 resides in multiple sub-cellular locations and its localization drastically changes after exposure to stresses. Taken together, H3553 underpins protein quality-control process and is involved in the response and adaptation to salinity and heat stresses.







Similar content being viewed by others
References
Becker SH, Darwin KH (2017) Bacterial proteasomes: mechanistic and functional insights. Microbiol Mol Biol Rev 81:e00036–e116. https://doi.org/10.1128/MMBR.00036-16
Block MA, Grossman AR (1988) Identification and purification of a derepressible alkaline phosphatase from Anacystis nidulans R2. Plant Physiol 86:1179–1184. https://doi.org/10.1104/pp.86.4.1179
Cecarini V, Gee J, Fioretti E, Amici M, Angeletti M, Eleuteri AM, Keller JN (2007) Protein oxidation and cellular homeostasis: emphasis on metabolism. Biochim Biophys Acta Mol Cell Res 1773:93–104. https://doi.org/10.1016/j.bbamcr.2006.08.039
Cheregi O, Wagner R, Funk C (2016) Insights into the cyanobacterial Deg/HtrA proteases. Front Plant Sci 7:694. https://doi.org/10.3389/fpls.2016.00694
Dahl JU, Gray MJ, Jakob U (2015) Protein quality control under oxidative stress conditions. J Mol Biol 427:1549–1563. https://doi.org/10.1016/j.jmb.2015.02.014
DasSarma S, DasSarma P (2015) Halophiles and their enzymes: negativity put to good use. Curr Opin Microbiol 25:120–126. https://doi.org/10.1016/j.mib.2015.05.009
Dikic I (2017) Proteasomal and autophagic degradation systems. Annu Rev Biochem 86:193–224. https://doi.org/10.1146/annurev-biochem-061516-044908
Fukaya F, Promden W, Hibino T, Tanaka Y, Nakamura T, Takabe T (2009) An Mrp-like cluster in the halotolerant cyanobacterium Aphanothece halophytica functions as a Na+/H+ antiporter. Appl Environ Microbiol 75:6626–6629. https://doi.org/10.1128/AEM.01387-09
Guan Y, Zhu Q, Huang D, Zhao S, Lo LJ, Peng J (2015) An equation to estimate the difference between theoretically predicted and SDS PAGE-displayed molecular weights for an acidic peptide. Sci Rep 5:13370. https://doi.org/10.1038/srep13370
Huang F, Hedman E, Funk C, Kieselbach T, Schröder WP, Norling B (2004) Isolation of outer membrane of Synechocystis sp. PCC 6803 and its proteomic characterization. Mol Cell Proteom 3:586–595. https://doi.org/10.1074/mcp.M300137-MCP200
Huerta-Cepas J, Serra F, Bork P (2016) ETE 3: reconstruction, analysis, and visualization of phylogenomic data. Mol Biol Evol 33:1635–1638. https://doi.org/10.1093/molbev/msw046
Huesgen PF, Miranda H, Lam X, Perthold M, Schuhmann H, Adamska I, Funk C (2011) Recombinant Deg/HtrA proteases from Synechocystis sp. PCC 6803 differ in substrate specificity, biochemical characteristics and mechanism. Biochem J 435:733–742. https://doi.org/10.1042/BJ20102131
Ibrahim HR, Higashiguchi S, Juneja LR, Kim M, Yamamoto T (1996) A structural phase of heat-denatured lysozyme with novel antimicrobial action. J Agric Food Chem 44:1416–1423. https://doi.org/10.1021/jf9507147
Jansén T, Kidron H, Taipaleenmäki H, Salminen T, Mäenpää P (2005) Transcriptional profiles and structural models of the Synechocystis sp. PCC 6803 Deg proteases. Photosynth Res 84:57–63. https://doi.org/10.1007/s11120-005-0475-x
Jiang J, Zhang X, Chen Y, Wu Y, Zhou ZH, Chang Z, Su SF (2008) Activation of DegP chaperone-protease via formation of large cage-like oligomers upon binding to substrate proteins. Proc Natl Acad Sci USA 105:11939–11944. https://doi.org/10.1073/pnas.0805464105
Kageyama H, Tripathi K, Rai AK, Cha-um S, Waditee-Sirisattha R, Takabe T (2011) An alkaline phosphatase/phosphodiesterase, PhoD, induced by salt stress and secreted out of the cells of Aphanothece halophytica, a halotolerant cyanobacterium. Appl Environ Microbiol 77:5178–5183. https://doi.org/10.1128/AEM.00667-11
Kanesaki Y, Suzuki I, Allakhverdiev SI, Mikami K, Murata N (2002) Salt stress and hyperosmotic stress regulate the expression of different sets of genes in Synechocystis sp. PCC 6803. Biochem Biophys Res Commun 290:339–348. https://doi.org/10.1006/bbrc.2001.6201
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kim KI, Park SC, Kang SH, Cheong GW, Chung CH (1999) Selective degradation of unfolded proteins by the self-compartmentalizing HtrA protease, a periplasmic heat shock protein in Escherichia coli. J Mol Biol 294:1363–1374. https://doi.org/10.1006/jmbi.1999.3320
Kim S, Song I, Eom G, Kim S (2018) A small periplasmic protein with a hydrophobic C-terminal residue enhances DegP proteolysis as a suicide activator. J Bacteriol 200:e00519–e617. https://doi.org/10.1128/JB.00519-17
Krojer T, Sawa J, Schäfer E, Saibil HR, Ehrmann M, Clausen T (2008) Structural basis for the regulated protease and chaperone function of DegP. Nature 453:885–890. https://doi.org/10.1038/nature07004
Latifi A, Ruiz M, Zhang CC (2009) Oxidative stress in cyanobacteria. FEMS Microbiol Rev 33:258–278. https://doi.org/10.1111/j.1574-6976.2008.00134.x
Meltzer M, Hasenbein S, Mamant N, Merdanovic M, Poepsel S, Hauske P, Kaiser M, Huber R, Krojer T, Clausen T, Ehrmann M (2009) Structure, function and regulation of the conserved serine proteases DegP and DegS of Escherichia coli. Microbiol Res 160:660–666. https://doi.org/10.1016/j.resmic.2009.07.012
Mizushima N (2007) Autophagy: process and function. Genes Dev 21:2861–2873. https://doi.org/10.1101/gad.1599207
Omata T, Murata N (1984) Isolation and characterization of three types of membranes from the cyanobacterium (blue-green alga) Synechocystis PCC 6714. Arch Microbiol 139:113–116. https://doi.org/10.1007/BF00401984
Page MJ, Di Cera E (2008) Serine peptidases: classification, structure and function. Cell Mol Life Sci 65:1220–1236. https://doi.org/10.1007/s00018-008-7565-9
Patipong T, Ngoennet S, Honda M, Hibino T, Waditee-Sirisattha R, Kageyama H (2019) A class I fructose-1,6-bisphosphate aldolase is associated with salt stress tolerance in a halotolerant cyanobacterium Halothece sp. PCC 7418. Arch Biochem Biophys 672:108059. https://doi.org/10.1016/j.abb.2019.07.024
Price MN, Dehal PS, Arkin AP (2009) FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol Biol Evol 26:1641–1650. https://doi.org/10.1093/molbev/msp077
Rai N, Ramaswamy A (2015) Temperature dependent dynamics of DegP-trimer: a molecular dynamics study. Comput Struct Biotechnol J 13:329–338. https://doi.org/10.1016/j.csbj.2015.04.004
Ramos V, Castelo-Branco R, Leao PN, Martins J, Carvalhal-Gomes S, Sobrinho da Silva F, Mendonça Filho JG, Vasconcelos VM (2017) Cyanobacterial diversity in microbial mats from the hypersaline lagoon system of Araruama, Brazil: an in-depth polyphasic study. Front Microbiol 8:1233. https://doi.org/10.3389/fmicb.2017.01233
Reed CJ, Lewis H, Trejo E, Winston V, Evilia C (2013) Protein adaptations in archaeal extremophiles. Archaea 2013:1–14. https://doi.org/10.1155/2013/373275
Roberts IN, Lam XT, Miranda H, Kieselbach T, Funk C (2012) Degradation of PsbO by the Deg protease HhoA is thioredoxin dependent. PLoS ONE 7:e45713. https://doi.org/10.1371/journal.pone.0045713
Skorko-Glonek J, Zurawa D, Kuczwara E, Wozniak M, Wypych Z, Lipinska B (1999) The Escherichia coli heat shock protease HtrA participates in defense against oxidative stress. Mol Gen Genet 262:342–350. https://doi.org/10.1007/s004380051092
Skórko-Glonek J, Żurawa D, Tanfani F, Scirè A, Wawrzynów A, Narkiewicz J, Bertoli E, Lipińska B (2003) The N-terminal region of HtrA heat shock protease from Escherichia coli is essential for stabilization of HtrA primary structure and maintaining of its oligomeric structure. Biochim Biophys Acta Proteins Proteom 1649:171–182. https://doi.org/10.1016/s1570-9639(03)00170-5
Waditee R, Hibino T, Tanaka Y, Nakamura T, Incharoensakdi A, Takabe T (2001) Halotolerant cyanobacterium Aphanothece halophytica contains an Na+/H+ antiporter, homologous to eukaryotic ones, with novel ion specificity affected by C-terminal tail. J Biol Chem 276:36931–36938. https://doi.org/10.1074/jbc.M103650200
Waditee R, Tanaka Y, Aoki K, Hibino T, Jikuya H, Takano J, Takabe T, Takabe T (2003) Isolation and functional characterization of N-Methyltransferases that catalyze betaine synthesis from glycine in a halotolerant photosynthetic organism Aphanothece halophytica. J Biol Chem 278:4932–4942. https://doi.org/10.1074/jbc.M210970200
Wu FG, Jiang YW, Sun HY, Luo JJ, Yu ZW (2015) Complexation of lysozyme with sodium poly (styrenesulfonate) via the two-state and non-two-state unfoldings of lysozyme. J Phys Chem B 119:14382–14392. https://doi.org/10.1021/acs.jpcb.5b07277
Xiong J, Sun Y, Yang Q, Tian H, Zhang H, Liu Y, Chen M (2017) Proteomic analysis of early salt stress responsive proteins in alfalfa roots and shoots. Proteome Sci 15:19. https://doi.org/10.1186/s12953-017-0127-z
Yildirim V, Baltaci MO, Ozgencli I, Sisecioglu M, Adiguzel A, Adiguzel G (2017) Purification and biochemical characterization of a novel thermostable serine alkaline protease from Aeribacillus pallidus C10: a potential additive for detergents. J Enzyme Inhib Med Chem 32:468–477. https://doi.org/10.1080/14756366.2016.1261131
Acknowledgements
T.P. thanks the Development and Promotion of Science and Technology Talented Project (DPST) for her Ph.D. Scholarship. This research was funded by Institute for Fermentation (grant number: G-2019-3-002) (to H.K.) and the Graduate School of Chulalongkorn University to commemorate the 72nd anniversary of his Majesty King Bhumibol Adulyadej, and the 90th Anniversary of the Chulalongkorn University Fund (Ratchadaphiseksomphot Endowment) (grant number: GCUGR1125623033D) (to T.P. and R.W.S.), respectively.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by S. Albers.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
792_2020_1162_MOESM1_ESM.pdf
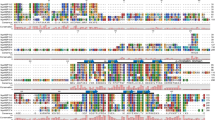
Supplementary file1 (PDF 1043 kb) Table S1 Primers used in this study. Figure S1 Gene expression analysis of 30 serine protease genes from Halothece 7418 grown under high salinity stress. Cells were collected at 0, 6 and 24 hours after exposure to salt stress (2.0 M NaCl). The total RNA was extracted and used as a template for RT-PCR analysis as described in the Materials and Methods section. The PCR products were subjected to electrophoresis and the relative amounts of the DNA fragments were quantitated using Image LabTM 3.0 software. The AprnpB gene was used as an internal control. The values at time zero for each gene were set to 1. The data are represented as the mean ± SEM of three independent experiments. The asterisks indicate significant differences from the value at time zero (one-sample t-test, p<0.05) Figure S2 Phylogenetic analysis of 30 serine protease paralogs from Halothece 7418. The amino acid sequences of the serine protease paralogs were retrieved from the KEGG database. The tree was generated with the maximum-likelihood method using the FastTree v2.1.8 software. The bars represent evolutionary distance. The scale bar comprises 0.5 expected changes per amino acid site (0.5 substitutions/site). Bootstrap probabilities are shown at the nodes. Figure S3 Alignment of the amino acid sequences of three HtrA2 proteins from Halothece 7418 (H3553, H2358, and H0395). The sequences were aligned using ClustalW (https://www.genome.jp/tools-bin/clustalw). (A) the catalytic triad of H3553 (Histidine:H140, aspartic acid:D170, and serine:S247) is shown by highlighted green. (B) predicted Trypsin and PDZ domains for H3553 H3553, H2358, and H0395 were obtained via SMART (https://smart.embl.de/). Figure S4 The effect of inhibitors on rH3553 activity. Proteolytic activity was measured using β-casein as a substrate. The activity value in the absence of inhibitor was set to 100%. Each value represents an average of three independent measurements. The data are represented as the mean ± SD of three independent experiments. Figure S5 Complex formation of H3553 in vivo. Total soluble protein extracts from control and salt acclimation conditions were subjected to gel filtration chromatography followed by Western blotting using anti-rH3553 antibody. The asterisks indicate the peaks of standard proteins and their native molecular size. Figure S6 Adsorption spectra of Halothece 7418 extracts after fractionation by sucrose gradient ultracentrifugation. (A) Sucrose gradient fractions. (B) plasma membrane fraction and (C) thylakoid membrane fraction
Rights and permissions
About this article
Cite this article
Patipong, T., Hibino, T., Kageyama, H. et al. The evolutionarily conserved HtrA is associated with stress tolerance and protein homeostasis in the halotolerant cyanobacterium Halothece sp. PCC7418. Extremophiles 24, 377–389 (2020). https://doi.org/10.1007/s00792-020-01162-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-020-01162-4