Abstract
The study of translation initiation in prokaryotes assumes that there should be a mechanism different from the canonical model, which postulates the formation of the pre-initiation complex through the interaction of the Shine-Dalgarno sequence (SD) at the 5′-end of mRNA and the anti-Shine-Dalgarno site at the 3′-end of 16S rRNA. In this paper we’ve studied the effect of TPS (Translation-initiation Promoting Site) on β-glucuronidase expression in E. coli cells at different cultivation temperatures. The examined leader sequences were cloned into the pET23c plasmid upstream the β-glucuronidase gene; protein expression was performed in E. coli BL21 (DE3) cells. β-glucuronidase activity was measured in bacterial cell extracts via paranitrophenyl b-d-glucuronide assay. The quantity of expressed protein was measured by Western blotting with following densitometry. It was shown that TPS increases the level of protein expression at stressful conditions (10 °C and 44 °C) 5–8 times compared to control. The combination of TPS and SD sites in the 5′-leader sequence of the mRNA created an enhancer that increased the expression level 2–3.6 times compared to a single SD-sequence. Based on the obtained data and the computer modeling of interaction between 16S rRNA and TPS, we proposed an alternative variation of prokaryotic translation initiation.



Similar content being viewed by others
References
Shine, J., & Dalgarno, L. (1975). Determinant of cistron specificity in bacterial ribosomes. Nature,254, 34–38.
Ivanov, L., Alexsandrova, R., Dragulev, B., & Abou-Haidar, M. (1995). A second putative mRNA binding site on the E. coli ribosome. Gene,160, 75–79.
Wu, C., & Janssen, G. (1997). Expression of streptomycete leaderless mRNA encoding chloramphenicol acetyltransferase in Escherichia coli. Journal of Bacteriology,179, 6824–6830.
Yurovsky, A., Amin, M. R., Gardin, J., Chen, Y., Skiena, S., et al. (2018). Prokaryotic coding regions have little if any specific depletion of Shine-Dalgarno motifs. PLoS ONE,13(8), e0202768.
Chang, B., Halgamuge, S., & Tang, S. L. (2006). Analysis of SD sequences in completed microbial genomes: Non-SD-led genes are as common as SD-led genes. Gene,373, 90–99.
Amin, M. R., Yurovsky, A., Chen, Y., Skiena, S., & Futcher, B. (2018). Re-annotation of 12,495 prokaryotic 16S rRNA 3' ends and analysis of Shine-Dalgarno and anti-Shine-Dalgarno sequences. PLoS ONE,13(8), e0202767.
Nakagawa, S., Niimura, Y., & Gojobori, T. (2017). Comparative genomic analysis of translation initiation mechanisms for genes lacking the Shine-Dalgarno sequence in prokaryotes. Nucleic Acids Research,45(7), 3922–3931.
Vesper, O., Amitai, S., Belitsky, M., Byrgazov, K., Kaberdina, A. C., et al. (2011). Selective translation of leaderless mRNAs by specialized ribosomes generated by MazF in Escherichia coli. Cell,147(1), 147–157.
Duval, M., Korepanov, A., Fuchsbauer, O., Fechter, P., Haller, A., et al. (2013). Escherichia coli ribosomal protein S1 unfolds structured mRNAs onto the ribosome for active translation initiation. PLoS Biology,11(12), e1001731.
Komarova, A. V., Tchufistova, L. S., Dreyfus, M., & Boni, I. V. (2005). AU-Rich sequences within 5’-untranslated leaders enhance translation and stabilize mRNA in Escherichia coli. Journal of Bacteriology,187(4), 1344–1349.
Thanaraj, T. A., & Pandit, M. W. (1989). An additional ribosome-binding site on mRNA of highly expressed genes and a bifunctional site on the colicin fragment of 16S rRNA from Escherichia coli: Important determinants of the efficiency of translation–initiation. Nucleic Acids Research,17, 2973–2985.
Thanaraj, T. A., & Pandit, M. W. (1990). Translation-initiation promoting site on transcripts of highly expressed genes from Saccharomyces cerevisiae and the role of hairpin stems to position the site near the initiation codon. Journal of Biomolecular Structure and Dynamics,7, 1279–1289.
Cervera, M. (2002). Histochemical and fluorometric assays for uidA (GUS) gene detection. In L. Peña (Ed.), Methods in molecular biology, transgenic plants: methods and protocols (pp. 203–212). Totowa, NJ: Humana.
Bradford, M. M. (1976). A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Analytical Biochemistry,72, 248–254.
Aich, S., Delbaere, L. T. J., & Chen, R. (2001). Continuous spectrophotometric assay for β-glucuronidase. BioTechniques,30, 846–850.
Schägger, H. (2006). Tricine–SDS-PAGE. Nature Protocols,1, 16–22.
Nakagawa, S., & Cuthill, I. C. (2007). Effect size, confidence interval and statistical significance: A practical guide for biologists. Biological Reviews of the Cambridge Philosophical Society,82, 591–605.
Ron, E. Z., & Davis, B. D. (1971). Growth rate of Escherichia coli at elevated temperatures: Limitation by methionine. Journal of Bacteriology,107, 391–396.
Morgan, G. J., Burkhardt, D. H., Kelly, J. W., & Powers, E. T. (2018). Translation efficiency is maintained at elevated temperature in Escherichia coli. Journal of Biological Chemistry,293(3), 777–793.
Farewell, A., & Neidhardt, F. C. (1998). Effect of temperature on in vivo protein synthetic capacity in Escherichia coli. Journal of Bacteriology,180, 4704–4710.
Zhang, Y., Burkhardt, D. H., Rouskin, S., Li, G. W., Weissman, J. S., et al. (2018). A stress response that monitors and regulates mRNA structure is central to cold shock adaptation. Molecular Cell,70, 274–286.
Osterman, I. A., Evfratov, S. A., Sergiev, P. V., & Dontsova, O. A. (2013). Comparison of mRNA features affecting translation initiation and reinitiation. Nucleic Acids Research,41, 474–486.
Romilly, C., Deindl, S., & Wagner, E. G. H. (2019). The ribosomal protein S1-dependent standby site in tisB mRNA consists of a single-stranded region and a 5′ structure element. PNAS,116(32), 15901–15906.
de Smit, M. H., & van Duin, J. (2003). Translational standby sites: How ribosomes may deal with the rapid folding kinetics of mRNA. Journal of Molecular Biology,331, 737–743.
Del Campo, C., Bartholomäus, A., Fedyunin, I., & Ignatova, Z. (2015). Secondary structure across the bacterial transcriptome reveals versatile roles in mRNA regulation and function. PLoS Genetics,11(10), e1005613.
Acknowledgements
This study was supported by the Kazakhstan Science Committee grants with the Project Numbers AP05130800 and AP05132532.
Author information
Authors and Affiliations
Contributions
AN and BI conceived and designed the experiments. AN, MS and AZ performed the experiments. AN analyzed and/or interpreted the data. AN wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors Nizkorodova A., Suvorova M., Zhigailov A., and Iskakov B. declare that they have no conflicts of interest related to the subject matter or materials discussed in this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary file1 (TIF 1341 kb)
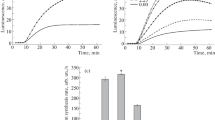
Fig. S1 β-glucuronidase activity at temperature stress condition relative to that at 37°C. GUS-activity in cells cultivated at 37°C was accepted as 100% for each construction; the relative GUS-activity values for stress temperatures are shown. The asterisk indicates the data sets significantly (p ≤ 0.05) differing from those at 37°C for each of the constructions.
Supplementary file2 (TIF 1191 kb)
Fig. S2 Graphical representation of suggested and canonical models of prokaryotic translation initiation. Standard conformation of small ribosomal subunit 30S is shown in beige color and conformation that appears under the cold shock condition – in purple color (with changed position of anti-Shine-Dalgarno sequence).
Supplementary file3 (TIF 765 kb)
Fig. S3 Secondary structure of leaders ‘TPSfar’ (a), ‘TPS__SD’ (b), ‘TPSlongSD’ (c). Start codon is shown in red, TPS – in blue, SD – in green.
Rights and permissions
About this article
Cite this article
Nizkorodova, A., Suvorova, M., Zhigailov, A. et al. The Effect of Translation Promoting Site (TPS) on Protein Expression in E. coli Cells. Mol Biotechnol 62, 326–334 (2020). https://doi.org/10.1007/s12033-020-00251-1
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-020-00251-1