Abstract
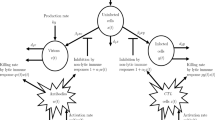
Infection by distinct Dengue virus serotypes and host immunity are intricately linked. In particular, certain levels of cross-reactive antibodies in the host may actually enhance infection severity leading to Dengue hemorrhagic fever (DHF). The coupled immunological and epidemiological dynamics of Dengue calls for a multi-scale modeling approach. In this work, we formulate a within-host model which mechanistically recapitulates characteristics of antibody dependent enhancement in Dengue infection. The within-host scale is then linked to epidemiological spread by a vector–host partial differential equation model structured by host antibody level. The coupling allows for dynamic population-wide antibody levels to be tracked through primary and secondary infections by distinct Dengue strains, along with waning of cross-protective immunity after primary infection. Analysis of both the within-host and between-host systems are conducted. Stability results in the epidemic model are formulated via basic and invasion reproduction numbers as a function of immunological variables. Additionally, we develop numerical methods in order to simulate the multi-scale model and assess the influence of parameters on disease spread and DHF prevalence in the population.







Similar content being viewed by others
References
Ackleh AS, Delcambre ML, Sutton KL, Ennis DG (2014) A structured model for the spread of Mycobacterium marinum: foundations for a numerical approximation scheme. Math Biosci Eng 11(4):679–721
Aguiar M, Stollenwerk N (2017) Mathematical models of dengue fever epidemiology: multi-strain dynamics, immunological aspects associated to disease severity and vaccines. Commun Biomath Sci 1(1):1–12
Aguiar M, Kooi B, Stollenwerk N (2008) Epidemiology of dengue fever: a model with temporary cross-immunity and possible secondary infection shows bifurcations and chaotic behaviour in wide parameter regions. Math Model Nat Phenom 3(4):48–70
Aguiar M, Stollenwerk N, Halstead SB (2016) The risks behind dengvaxia recommendation. Lancet Infect Dis 16(8):882–883
Antia A, Ahmed H, Handel A, Carlson NE, Amanna IJ, Antia R, Slifka M (2018) Heterogeneity and longevity of antibody memory to viruses and vaccines. PLoS Biol 16(8):e2006,601
Barbarossa MV, Röst G (2015) Immuno-epidemiology of a population structured by immune status: a mathematical study of waning immunity and immune system boosting. J Math Biol 71(6–7):1737–1770
Ben-Shachar R, Koelle K (2015) Minimal within-host dengue models highlight the specific roles of the immune response in primary and secondary dengue infections. J R Soc Interface 12(103):20140,886
Browne CJ, Pilyugin SS (2013) Global analysis of age-structured within-host virus model. Discrete Contin Dyn Syst B 18(8):1999–2017
Carrillo J, Colombo R, Gwiazda P, Ulikowska A (2012) Structured populations, cell growth and measure valued balance laws. J Differ Equ 252(4):3245–3277
Clapham HE, Quyen TH, Kien DTH, Dorigatti I, Simmons CP, Ferguson NM (2016) Modelling virus and antibody dynamics during dengue virus infection suggests a role for antibody in virus clearance. PLoS Comput Biol 12(5):e1004,951
Cummings DA, Schwartz IB, Billings L, Shaw LB, Burke DS (2005) Dynamic effects of antibody-dependent enhancement on the fitness of viruses. Proc Natl Acad Sci 102(42):15259–15264
Dejnirattisai W, Jumnainsong A, Onsirisakul N, Fitton P, Vasanawathana S, Limpitikul W, Puttikhunt C, Edwards C, Duangchinda T, Supasa S et al (2010) Cross-reacting antibodies enhance dengue virus infection in humans. Science 328(5979):745–748
Diekmann O, de Graaf W, Kretzschmar M, Teunis P (2018) Waning and boosting: on the dynamics of immune status. J Math Biol 77(6–7):2023–2048
Dowd KA, Pierson TC (2011) Antibody-mediated neutralization of flaviviruses: a reductionist view. Virology 411(2):306–315
Ferguson N, Anderson R, Gupta S (1999) The effect of antibody-dependent enhancement on the transmission dynamics and persistence of multiple-strain pathogens. Proc Natl Acad Sci 96(2):790–794
Ferguson NM, Rodríguez-Barraquer I, Dorigatti I, Mier-y Teran-Romero L, Laydon DJ, Cummings DA (2016) Benefits and risks of the sanofi-pasteur dengue vaccine: modeling optimal deployment. Science 353(6303):1033–1036
Gandolfi A, Pugliese A, Sinisgalli C (2015) Epidemic dynamics and host immune response: a nested approach. J Math Biol 70(3):399–435
Gilchrist MA, Sasaki A (2002) Modeling host-parasite coevolution: a nested approach based on mechanistic models. J Theor Biol 218(3):289–308
Gujarati TP, Ambika G (2014) Virus antibody dynamics in primary and secondary dengue infections. J Math Biol 69(6–7):1773–1800
Gulbudak H (2019) An immuno-epidemiological vector–host model with within-vector viral kinetics. J Biol Syst (forthcoming)
Gulbudak H, Martcheva M (2013) Forward hysteresis and backward bifurcation caused by culling in an avian influenza model. Math Biosci 246(1):202–212
Gulbudak H, Cannataro VL, Tuncer N, Martcheva M (2017) Vector-borne pathogen and host evolution in a structured immuno-epidemiological system. Bull Math Biol 79(2):325–355
Handel A, Rohani P (2015) Crossing the scale from within-host infection dynamics to between-host transmission fitness: a discussion of current assumptions and knowledge. Philos Trans R Soc B Biol Sci 370(1675):20140,302
Katzelnick LC, Montoya M, Gresh L, Balmaseda A, Harris E (2016) Neutralizing antibody titers against dengue virus correlate with protection from symptomatic infection in a longitudinal cohort. Proc Natl Acad Sci 113(3):728–733
Katzelnick LC, Gresh L, Halloran ME, Mercado JC, Kuan G, Gordon A, Balmaseda A, Harris E (2017) Antibody-dependent enhancement of severe dengue disease in humans. Science 358(6365):929–932
Martcheva M, Pilyugin SS (2006) An epidemic model structured by host immunity. J Biol Syst 14(02):185–203
Nguyen NM, Kien DTH, Tuan TV, Quyen NTH, Tran CN, Thi LV, Le Thi D, Nguyen HL, Farrar JJ, Holmes EC et al (2013) Host and viral features of human dengue cases shape the population of infected and infectious Aedes aegypti mosquitoes. Proc Natl Acad Sci 110(22):9072–9077
Nikin-Beers R, Ciupe SM (2015) The role of antibody in enhancing dengue virus infection. Math Biosci 263:83–92
Nikin-Beers R, Ciupe SM (2017) Modelling original antigenic sin in dengue viral infection. Math Med Biol J IMA 35(2):257–272
Nikin-Beers R, Blackwood JC, Childs LM, Ciupe SM (2018) Unraveling within-host signatures of dengue infection at the population level. J Theor Biol 446:79–86
Pugliese A (2011) The role of host population heterogeneity in the evolution of virulence. J Biol Dyn 5(2):104–119
Reich NG, Shrestha S, King AA, Rohani P, Lessler J, Kalayanarooj S, Yoon IK, Gibbons RV, Burke DS, Cummings DA (2013) Interactions between serotypes of dengue highlight epidemiological impact of cross-immunity. J R Soc Interface 10(86):20130,414
Ripoll DR, Wallqvist A, Chaudhury S (2019) Molecular simulations reveal the role of antibody fine specificity and viral maturation state on antibody-dependent enhancement of infection in dengue virus. Front Cell Infect Microbiol 9:200
Salje H, Cummings DA, Rodriguez-Barraquer I, Katzelnick LC, Lessler J, Klungthong C, Thaisomboonsuk B, Nisalak A, Weg A, Ellison D et al (2018) Reconstruction of antibody dynamics and infection histories to evaluate dengue risk. Nature 557(7707):719–723
Stefan MI, Le Novere N (2013) Cooperative binding. PLoS Comput Biol 9(6):e1003,106
Thieme HR (1990) “Integrated semigroups” and integrated solutions to abstract cauchy problems. J Math Anal Appl 152(2):416–447
Tuncer N, Gulbudak H, Cannataro VL, Martcheva M (2016) Structural and practical identifiability issues of immuno-epidemiological vector–host models with application to rift valley fever. Bull Math Biol 78(9):1796–1827
Veliov VM, Widder A (2016) Modelling and estimation of infectious diseases in a population with heterogeneous dynamic immunity. J Biol Dyn 10(1):457–476
Wahala WM, de Silva AM (2011) The human antibody response to dengue virus infection. Viruses 3(12):2374–2395
Wearing HJ, Rohani P (2006) Ecological and immunological determinants of dengue epidemics. Proc Natl Acad Sci 103(31):11802–11807
Yang Y, Meng Y, Halloran ME, Longini IM Jr (2017) Reply to aguiar and stollenwerk. Clin Infect Dis 66(4):642–642
Acknowledgements
The authors thank two anonymous reviewers for their helpful comments and feedback on the manuscript. This work was supported by a grant from the Simons Foundation/SFARI (638193, HG). CJB is partially supported by a U.S. National Science Foundation Grant (DMS-1815095).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix: Numerical convergence rates
Appendix: Numerical convergence rates
In this section, we provide tables showing computed rates and order of convergence for numerical experiments of the finite difference and multi-scale simulation procedure described in Sect. 4. For the numerical tests, we calculate the error in norm between computed solutions of the t, y stepping method at certain step sizes \(\varDelta t,h=\varDelta y\) and reference solutions at some final time \(t=T\). We utilize three different types of reference solutions: (i) the numerically approximated equilibrium given by our derived formula (31), (ii) solution of the numerical scheme with smallest step sizes \(\varDelta t,{\tilde{h}}=\varDelta y\), and (iii) solution of the numerical scheme with step sizes multiplied by factor of 1/2, \(\frac{\varDelta t}{2},\frac{h}{2}\). For each error calculation at step size h, \(e_h\), we form a sequence by successively decreasing step size by 1/2, whereby we compute order of convergence by \(\log _2(e_h/e_{h/2})\). Furthermore, we consider two different scenarios: (a) we start the initial condition where infected vectors, \(I_v^1,I_v^2\) are slightly perturbed from \({\mathcal {E}}_0\) (outbreak scenario) with final time \(T=50\) days, (b) we start the initial condition at numerically calculated equilibrium with final time \(T=500\) days. For the former scenario (a), we do not use numerically calculated equilibrium as a reference solution since this may be far off from simulation at \(t=50\).
We compute the different orders of convergence because there are several sources of error and to test different initial condition scenarios. Our method relies on distinct algorithms in addition to the finite difference scheme, such as Runge-Kutta method for within-host ODE (ode45 in MatLab), interpolation, integration and, in the case of numerical equilibrium formula, nonlinear root-finding. Each routine can produce error, which can also propagate in the form of discontinuities in recovered distribution corresponding to an influx of recovery from primary infected individuals with pre-existent antibody levels at a certain mesh points from the initial susceptible antibody distribution. In order to efficiently reduce error we utilize the trapezoidal integration when integrating with respect to initial susceptible antibody level \(y_0\), but left endpoint integration for other antibody variables since there is small number of mesh points (\(M_0\)) for \(s(\cdot , y_0)\) when compared to the range of antibody levels after infection and waning. We do also provide one numerical test with only left-endpoint integration shown in last two tables, which gives more error than trapezoidal, but has more regular order of convergence pattern. In addition, we include comparisons with a larger step size (\(\varDelta y=0.06\)) for scenarios (a) and the last two tables, which forces a point distribution for susceptible antibody levels, creating different error structure. Overall, from the different numerical tests, we observe convergence to certain error rates within a particular compartment and/or test scenario, ranging from orders that are sub-linear (\(<1\)) to larger than quadratic (\(>2\)). When comparing with reference solutions computed by numerical simulation at smaller step size, the order of convergence is mostly faster than linear (Tables 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16).
Rights and permissions
About this article
Cite this article
Gulbudak, H., Browne, C.J. Infection severity across scales in multi-strain immuno-epidemiological Dengue model structured by host antibody level. J. Math. Biol. 80, 1803–1843 (2020). https://doi.org/10.1007/s00285-020-01480-3
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00285-020-01480-3
Keywords
- Dengue hemorrhagic fever (DHF)
- Antibody dependent enhancement (ADE)
- Multi-scale
- Immuno-epidemiological model
- Size-structured partial differential equation (PDE)
- Invasion stability analysis