Abstract
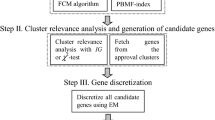
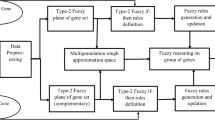
In recent years, microarray technology and gene expression profiles have been widely used to detect, predict, or classify the samples of various diseases. The presence of large genes in these profiles and the small number of samples are known challenges in this field and are widely considered in previous papers. In previous studies, other topics such as the noise of microarray data or the dependence of selected genes on samples have been less considered. Therefore, we have tried to address these two issues by using a fuzzy classifier and stability index of selected genes, respectively. The proposed method is based on the regression function between the genes and class labels which is determined by the self-representing method. This regression function is determined individually for each class of the database. To minimize the effect of noise in microarray data, a fuzzy classifier is applied in the proposed model. Four databases of gene expression profiles are examined in this article, and the results indicate that the proposed model has a relative advantage over the previous methods.

Graphical abstract







Similar content being viewed by others
References
Gregory KB, Momin AA, Coombes KR, Baladandayuthapani V (2014) Latent feature decompositions for integrative analysis of multi-platform genomic data. IEEE/ACM Trans Comput Biol Bioinform 11(6):984–994. https://doi.org/10.1109/TCBB.2014.2325035
Hu Y, Liu J-X, Gao Y-L, Li S-J, Wang J (2019) Differentially expressed genes extracted by the tensor robust principal component analysis (TRPCA) method. Complexity 2019:1–13
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, Coller H, Loh ML, Downing JR, Caligiuri MA, Bloomfield CD, Lander ES (1999) Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286(5439):531–537. https://doi.org/10.1126/science.286.5439.531
Mitra P, Murthy CA, Pal SK (2002) Unsupervised feature selection using feature similarity. IEEE Trans Pattern Anal Mach Intell 24(3):301–312. https://doi.org/10.1109/34.990133
van’t Veer LJ, Dai H, van de Vijver MJ, He YD, AAM H, Mao M, Peterse HL, van der Kooy K, Marton MJ, Witteveen AT, Schreiber GJ, Kerkhoven RM, Roberts C, Linsley PS, Bernards R, Friend SH (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415(6871):530–536. https://doi.org/10.1038/415530a
He X, Cai D, Niyogi P (2005) Laplacian score for feature selection. Paper presented at the Proceedings of the 18th International Conference on Neural Information Processing Systems, Vancouver, British Columbia, Canada
Chang CF, Wai KM, Patterton HG (2004) Calculating the statistical significance of physical clusters of co-regulated genes in the genome: the role of chromatin in domain-wide gene regulation. Nucleic Acids Res 32(5):1798–1807. https://doi.org/10.1093/nar/gkh507
Hanchuan P, Fuhui L, Ding C (2005) Feature selection based on mutual information criteria of max-dependency, max-relevance, and min-redundancy. IEEE Trans Pattern Anal Mach Intell 27(8):1226–1238. https://doi.org/10.1109/TPAMI.2005.159
Guyon I, Weston J, Barnhill S, Vapnik V (2002) Gene selection for cancer classification using support vector machines. Mach Learn 46(1):389–422. https://doi.org/10.1023/A:1012487302797
Zhu P, Zuo W, Zhang L, Hu Q, Shiu SCK (2015) Unsupervised feature selection by regularized self-representation. Pattern Recogn 48(2):438–446. https://doi.org/10.1016/j.patcog.2014.08.006
Zhu P, Zhu W, Wang W, Zuo W, Hu Q (2017) Non-convex regularized self-representation for unsupervised feature selection. Image Vis Comput 60:22–29. https://doi.org/10.1016/j.imavis.2016.11.014
Shang R, Zhang Z, Jiao L, Liu C, Li Y (2016) Self-representation based dual-graph regularized feature selection clustering. Neurocomputing 171:1242–1253. https://doi.org/10.1016/j.neucom.2015.07.068
Liu Y, Liu K, Zhang C, Wang J, Wang X (2017) Unsupervised feature selection via diversity-induced self-representation. Neurocomputing 219:350–363. https://doi.org/10.1016/j.neucom.2016.09.043
Tang C, Cao L, Zheng X, Wang M (2018) Gene selection for microarray data classification via subspace learning and manifold regularization. Med Biol Eng Comput 56(7):1271–1284. https://doi.org/10.1007/s11517-017-1751-6
Yang K, Cai Z, Li J, Lin G (2006) A stable gene selection in microarray data analysis. BMC Bioinformatics 7(1):228–243. https://doi.org/10.1186/1471-2105-7-228
Mahmoodian H, Hamiruce Marhaban M, Abdulrahim R, Rosli R, Saripan I (2011) Using fuzzy association rule mining in cancer classification. Australas Phys Eng Sci Med 34(1):41–54. https://doi.org/10.1007/s13246-011-0054-8
Aziz R, Verma CK, Srivastava N (2016) A fuzzy based feature selection from independent component subspace for machine learning classification of microarray data. Genomics Data 8:4–15. https://doi.org/10.1016/j.gdata.2016.02.012
Paul A, Sil J, Mukhopadhyay CD (2017) Gene selection for designing optimal fuzzy rule base classifier by estimating missing value. Appl Soft Comput 55:276–288. https://doi.org/10.1016/j.asoc.2017.01.046
Mahmoodian H, Ebrahimian L (2016) Using support vector regression in gene selection and fuzzy rule generation for relapse time prediction of breast cancer. Biocybern Biomed Eng 36(3):466–472. https://doi.org/10.1016/j.bbe.2016.03.003
Cetisli B Gene selection by using a linguistic hedged adaptive neuro-fuzzy classifier for cancer classification. In: 2009 IEEE 17th Signal Processing and Communications Applications Conference, 9–11 April 2009 2009. pp 257–260. https://doi.org/10.1109/SIU.2009.5136381
Zhang M, Zhang L, Zou J, Yao C, Xiao H, Liu Q, Wang J, Wang D, Wang C, Guo Z (2009) Evaluating reproducibility of differential expression discoveries in microarray studies by considering correlated molecular changes. Bioinformatics 25(13):1662–1668. https://doi.org/10.1093/bioinformatics/btp295
Kuncheva LI (2007) A stability index for feature selection. Paper presented at the Proceedings of the 25th conference on Proceedings of the 25th IASTED International Multi-Conference: artificial intelligence and applications, Innsbruck, Austria
Kalousis A, Prados J, Hilario M (2007) Stability of feature selection algorithms: a study on high-dimensional spaces. Knowl Inf Syst 12(1):95–116. https://doi.org/10.1007/s10115-006-0040-8
Alon U, Barkai N, Notterman DA, Gish K, Ybarra S, Mack D, Levine AJ (1999) Broad patterns of gene expression revealed by clustering analysis of tumor and normal colon tissues probed by oligonucleotide arrays. Proc Natl Acad Sci 96(12):6745–6750. https://doi.org/10.1073/pnas.96.12.6745
Alter MD, Kharkar R, Ramsey KE, Craig DW, Melmed RD, Grebe TA, Bay RC, Ober-Reynolds S, Kirwan J, Jones JJ, Turner JB, Hen R, Stephan DA (2011) Autism and increased paternal age related changes in global levels of gene expression regulation. PLoS One 6(2):e16715. https://doi.org/10.1371/journal.pone.0016715
Li H-D, Xu Q-S, Liang Y-Z (2017) A phase diagram for gene selection and disease classification. Chemom Intell Lab Syst 167:208–213. https://doi.org/10.1016/j.chemolab.2017.06.008
Zhang L, Zhou W, Wang B, Zhang Z, Li F (2018) Applying 1-norm SVM with squared loss to gene selection for cancer classification. Appl Intell 48(7):1878–1890. https://doi.org/10.1007/s10489-017-1056-3
Latkowski T, Osowski S (2017) Gene selection in autism – comparative study. Neurocomputing 250:37–44. https://doi.org/10.1016/j.neucom.2016.08.123
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Davoudi, A., Mahmoodian, H. Stable gene selection by self-representation method in fuzzy sample classification. Med Biol Eng Comput 58, 1213–1223 (2020). https://doi.org/10.1007/s11517-020-02160-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-020-02160-6