Abstract
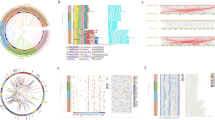
Sterols are important lipid constituents of cellular membranes in plants and other organisms. Sterol homeostasis is under strict regulation in plants because excess sterols negatively impact plant growth. HIGH STEROL ESTER 1 (HISE1) functions as a negative regulator of sterol accumulation. If sterol production exceeds a certain threshold, excess sterols are detoxified via conversion to sterol esters by PHOSPHOLIPID STEROL ACYL TRANSFERASE 1 (PSAT1). We previously reported that the Arabidopsis thaliana double mutant hise1-3 psat1-2 shows 1.5-fold higher sterol content than the wild type and consequently a severe growth defect. However, the specific defects caused by excess sterol accumulation in plants remain unknown. In this study, we investigated the effects of excess sterols on plants by analyzing the phenotypes and transcriptomes of the hise1-3 psat1-2 double mutant. Transcriptomic analysis revealed that 435 genes were up-regulated in hise1-3 psat1-2 leaves compared with wild-type leaves. Gene ontology (GO) enrichment analysis revealed that abiotic and biotic stress-responsive genes including RESPONSIVE TO DESICCATION 29B/LOW-TEMPERATURE-INDUCED 65 (RD29B/LTI65) and COLD-REGULATED 15A (COR15A) were up-regulated in hise1-3 psat1-2 leaves compared with wild-type leaves. Expression levels of senescence-related genes were also much higher in hise1-3 psat1-2 leaves than in wild-type leaves. hise1-3 psat1-2 leaves showed early senescence, suggesting that excess sterols induce senescence of leaves. In the absence of sucrose, hise1-3 psat1-2 exhibited defects in seedling growth and root elongation. Together, our data suggest that excess sterol accumulation disrupts cellular activities of vegetative organs including leaves and roots, resulting in multiple damages to plants.





Similar content being viewed by others
References
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B 57:298–300
Bouvier-Nave P, Berna A, Noiriel A, Compagnon V, Carlsson AS, Banas A, Stymne S, Schaller H (2010) Involvement of the phospholipid sterol acyltransferase1 in plant sterol homeostasis and leaf senescence. Plant Physiol 152:107–119
Carland F, Fujioka S, Nelson T (2010) The sterol methyltransferases SMT1, SMT2, and SMT3 influence Arabidopsis development through nonbrassinosteroid products. Plant Physiol 153:741–756
Chen S, Zhou Y, Chen Y, Gu J (2018) Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890
Clouse SD (2002) Arabidopsis mutants reveal multiple roles for sterols in plant development. Plant Cell 14:1995–2000
Erffelinck ML, Goossens A (2018) Review: endoplasmic reticulum-associated degradation (ERAD)-dependent control of (Tri)terpenoid metabolism in plants. Planta Med 84:874–880
Eulgem T, Somssich IE (2007) Networks of WRKY transcription factors in defense signaling. Curr Opin Plant Biol 10:366–371
Fisher RA (1922) On the interpretation of χ2 from contingency tables, and the calculation of P. J R Stat Soc 85:87–94
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360
Kodaira KS, Qin F, Tran LS, Maruyama K, Kidokoro S, Fujita Y, Shinozaki K, Yamaguchi-Shinozaki K (2011) Arabidopsis Cys2/His2 zinc-finger proteins AZF1 and AZF2 negatively regulate abscisic acid-repressive and auxin-inducible genes under abiotic stress conditions. Plant Physiol 157:742–756
Kopischke M, Westphal L, Schneeberger K, Clark R, Ossowski S, Wewer V, Fuchs R, Landtag J, Hause G, Dormann P, Lipka V, Weigel D, Schulze-Lefert P, Scheel D, Rosahl S (2013) Impaired sterol ester synthesis alters the response of Arabidopsis thaliana to Phytophthora infestans. Plant J 73:456–468
Kurt F, Filiz E (2018) Genome-wide and comparative analysis of bHLH38, bHLH39, bHLH100 and bHLH101 genes in Arabidopsis, tomato, rice, soybean and maize: insights into iron (Fe) homeostasis. Biometals 31:489–504
Lawrence M, Huber W, Pages H, Aboyoun P, Carlson M, Gentleman R, Morgan MT, Carey VJ (2013) Software for computing and annotating genomic ranges. PLoS Comput Biol 9:e1003118
Li H, Durbin R (2009) Fast and accurate short read alignment with burrows-wheeler transform. Bioinformatics 25:1754–1760
Maurey K, Wolf F, Golbeck J (1986) 3-Hydroxy-3-methylglutaryl coenzyme A reductase activity in Ochromonas malhamensis: a system to study the relationship between enzyme activity and rate of steroid biosynthesis. Plant Physiol 82:523–527
Nakamoto M, Schmit AC, Heintz D, Schaller H, Ohta D (2015) Diversification of sterol methyltransferase enzymes in plants and a role for beta-sitosterol in oriented cell plate formation and polarized growth. Plant J 84:860–874
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140
Shimada TL, Shimada T, Okazaki Y, Higashi Y, Saito K, Kuwata K, Oyama K, Kato M, Ueda H, Nakano A, Ueda T, Takano Y, Hara-Nishimura I (2019) HIGH STEROL ESTER 1 is a key factor in plant sterol homeostasis. Nat Plants 5:1154–1166
Sivitz AB, Hermand V, Curie C, Vert G (2012) Arabidopsis bHLH100 and bHLH101 control iron homeostasis via a FIT-independent pathway. PLoS One 7:e44843
Son GH, Wan J, Kim HJ, Nguyen XC, Chung WS, Hong JC, Stacey G (2012) Ethylene-responsive element-binding factor 5, ERF5, is involved in chitin-induced innate immunity response. Mol Plant Microbe Interact 25:48–60
Sun J, Nishiyama T, Shimizu K, Kadota K (2013) TCC: an R package for comparing tag count data with robust normalization strategies. BMC Bioinform 14:219
Takahashi H, Nakagawa A, Kojima S, Takahashi A, Cha BY, Woo JT, Nagai K, Machida Y, Machida C (2012) Discovery of novel rules for G-quadruplex-forming sequences in plants by using bioinformatics methods. J Biosci Bioeng 114:570–575
Yamaguchi-Shinozaki K, Shinozaki K (1993) Characterization of the expression of a desiccation-responsive rd29 gene of Arabidopsis thaliana and analysis of its promoter in transgenic plants. Mol Gen Genet 236:331–340
Yoo SY, Kim Y, Kim SY, Lee JS, Ahn JH (2007) Control of flowering time and cold response by a NAC-domain protein in Arabidopsis. PLoS One 2:e642
Zhu B, Jiang L, Huang T, Zhao Y, Liu T, Zhong Y, Li X, Campos A, Pomeroy K, Masliah E, Zhang D, Xu H (2017) ER-associated degradation regulates Alzheimer's amyloid pathology and memory function by modulating gamma-secretase activity. Nat Commun 8:1472
Acknowledgements
This work was supported by Grants-in-Aid for Scientific Research (JP15H05776 and JP22000014 to I.H.-N.; JP16K18834 and JP19K05809 to T.L.S.; JP18H03330 to H.T.) from the Japan Society for the Promotion of Science (JSPS); Leading Initiative for Excellent Young Researchers (LEADER; J16HJ00026 to T.L.S.) and a Grant-in-Aid for Scientific Research on Innovative Areas (KAKENHI, JP17H05659 to H.T.) from the Ministry of Education, Culture, Sports, Science and Technology in Japan (MEXT); Kato Memorial Bioscience Foundation to T.L.S.; SUNBOR GRANT of Suntory Foundation for life science to T.L.S.; Phytochemical Plant Molecular Science of Strategic Priority Research Promotion Program from Chiba University to T.L.S.; the NIBB Collaborative Research Program (17-301, 17-453, 18-301, 18-412, 19-302 to T.L.S.), and the Hirao Taro Foundation of KONAN GAKUEN for Academic Research to I.H.-N. We thank Miwako Matsumoto (NIBB), Asaka Akita (NIBB), Kazuo Ebine (NIBB), and Mitsumasa Hanaoka (Chiba University) for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shimada, T.L., Yamaguchi, K., Shigenobu, S. et al. Excess sterols disrupt plant cellular activity by inducing stress-responsive gene expression. J Plant Res 133, 383–392 (2020). https://doi.org/10.1007/s10265-020-01181-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-020-01181-4