Abstract
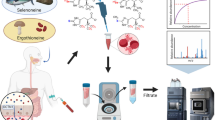
The methionine cycle is a key pathway to provide substrates for many basic biological processes including methylation and redox reactions. Here, we demonstrated a rapid and sensitive liquid chromatography–tandem mass spectrometry (LC–MS/MS) method for quantifying the metabolites and co-factors of the methionine metabolism. The analytes included methionine, S-adenosylmethionine, S-adenosylhomocysteine, 5′-deoxy-5′-(methylthio)adenosine, homocysteine, cystathionine, cysteine, glutathione, 5-methyltetrahydrofolate, vitamins B6, folic acid and vitamin B12. Linearities were obtained in all of the analytes with R2 larger than 0.99. Limits of quantification were in the range of 0.02–0.91 ng/106 cells, respectively. The recoveries of all of the analytes spiked at low, medium and high concentrations in cell lysates ranged from 74 to 117% and the accuracies ranged from 93.5 to 123.4%. The intra-day and inter-day precisions were lower than 20% of the relative standard deviations. This method was specifically designed for determining the intracellular concentrations of these analytes in the porcine small intestinal epithelial cell lines and the pig iliac artery endothelial cell lines. It enables the demonstration of changes in the concentrations of methionine intermediates when the cells are faced with deficient, moderate or excessive methionine. This method is expected to facilitate the understanding of the regulatory mechanism of nutrients on methionine metabolism.




Similar content being viewed by others
References
Brosnan JT, Brosnan ME (1640S) J Nutr 136(6):1636S–1640S
Fontecave M, Atta M, Mulliez E (2004) Trends Biochem Sci 29(5):243–249
Andrade F, Rodríguezsoriano J, Prieto JA, Aguirre M, Ariceta G, Lage S, Azcona I, Prado C, Sanjurjo P, Aldámiz-Echevarría L (2011) Nephrol Dial Transpl 26(1):328–336
Ragione FD, Carteni-Farina M, Gragnaniello V, Schettino MI, Zappia V (1986) J Biol Chem 261(26):12324–12329
McCarty M (2010) Med Hypoth 75(2):141–147
Kalhan SC, Marczewski SE (2012) Rev Endocr Metab Disord 13(2):109–119
Matthews RG, Elmore CL (2007) Clin Chem Lab Med 45(12):1700–1703
Stoll B, Henry J, Reeds PJ, Yu H, Jahoor F, Burrin DG (1998) J Nutr 128(3):606–614
Fang Z, Yao K, Zhang X, Zhao S, Sun Z, Tian G, Yu B, Lin Y, Zhu B, Jia G, Zhang K, Chen D, Wu D (2010) Amino Acids 39(3):633–640
Xia M, Pan Y, Guo L, Wei XX, Xiong J, Wang L, Peng J, Wang C, Peng J, Wei HK (2019) J Anim Sci 97:3487–3497
Longchamp A, Mirabella T, Arduini A et al (2018) Cell 173(1):117–129.e14
Sahin M, Sahin E, Gümüşlü S, Erdoğan A, Gültekin M (2011) Biochem Biophys Res Commun 408(1):145–148
Pan L, Yu G, Huang J, Zheng X, Xu Y (2017) Biosci Rep 37(5):BSR20170860
Kořínek M, Šístek V, Mládková J, Mikeš P, Jiráček J, Selicharová I (2013) Biomed Chromatogr 27(1):111–121
Johansson M, Van Guelpen B, Vollset SE, Hultdin J, Bergh A, Key T, Midttun O, Hallmans G, Ueland PM, Stattin P (2009) Cancer Epidemiol Biomark Prev 18(5):1538–1543
Stevens AP, Dettmer K, Kirovski G, Samejima K, Hellerbrand C, Bosserhoff AK, Oefner PJ (2010) J Chromatogr A 1217(19):3282–3288
Iglesias González T, Cinti M, Montes-Bayón M, Fernández de la Campa MR, Blanco-González E (2015) J Chromatogr A 1393(3):89–95
Borowczyk K, Chwatko G, Kubalczyk P, Jakubowski H, Kubalska J, Głowacki R (2016) Talanta 161:917–924
Mohammadi S, Domeno C, Nerin I, Aznar M, Samper P, Khayatian G, Nerin C (2017) J Pharm Biomed Anal 145:331–338
Glushchenko AV, Jacobsen DW (2007) Antioxid Redox Signal 9(11):1883–1898
Sen CK, Packer L (2000) Am J Clin Nutr 72(2):653S–669S
Persichilli S, Gervasoni J, Iavarone F, Zuppi C, Zappacosta B (2010) J Sep Sci 33(20):3119–3124
Gardner LA, Desiderio DM, Groover CJ, Hartzes A, Yates CR, Zucker-Levin AR, Bloom L, Levin MC (2013) Electrophoresis 34(11):1710–1716
Zhang M, Wang L, Pei P, Bao YH (2018) Anal Methods 10(11):1315–1324
Fu XW, Xu YK, Chan P, Pattengale PK (2013) Jimd Rep 10:69–78
Ghassabian S, Rethwan NSA, Griffiths L, Smit MT (2014) J Chromatogr B Anal Technol Biomed Life Sci 972:14–21
Guiraud SP, Montoliu I, Silva LD, Dayon L, Galindo AN, Corthésy J, Kussmann M, Martin FP (2017) Anal Bioanal Chem 409(1):295–305
Kovac A, Svihlova K, Michalicova A, Novak M (2014) J Chromatogr Sci 53(6):953–958
Da Silva L, Collino S, Cominetti O, Martin FP, Montoliu I, Moreno SO, Corthesy J, Kaput J, Kussmann M, Monteiro JP (2016) Bioanalysis 8(18):1937–1949
Jiang Y, Mistretta B, Elsea S, Sun Q (2017) Clin Chim Acta 464:93–97
Goniewicz ML, Havel CM, Peng MW, Jacob P, Dempsey D, Yu L, Zielinska-Danch W, Koszowski B, Czogala J, Sobczak A, Benowitz NL (2009) Cancer Epidemiol Biomark Prev 18(12):3421–3425
US Department of Health and Human Services FaDA, Center for Drug Evaluation and Research (CDER), Center for Veterinary Medicine (CVM) (2011) Guidance for industry, bioanalytical method validation.
Oosterink JE, Naninck EF, Korosi A, Lucassen PJ, van Goudoever JB, Schierbeek H (2015) J Chromatogr B 998–999:106–113
Zheng LF, Zuo FR, Zhao SJ, He PL, Wei HK, Xiang QH, Pang JM, Peng J (2017) Br J Nutr 117(7):911–922
Shi B, Liu J, Sun Z, Li T, Zhu W, Tang Z (2016) J Appl Anim Res 46(1):74–80
Song T, Lu J, Deng Z, Xu T, Yang Y, Wei H, Li S, Jiang S, Peng J (2018) Int J Obes 42:1812–1820
Acknowledgements
The authors greatly appreciate the State Key Laboratory of Agricultural Microbiology of Huazhong Agricultural University for the LC–MS/MS usage.
Funding
This study was supported by the Fundamental Research Funds for the Central Universities of China (no. 2662018JC009 and no. 2662017PY017); National key Research and Development project of China (no. 2017YFD0502004); China Agriculture Research System (no. CARS-36); Hubei Provincial Creative Team Project of Agricultural Science and Technology (no. 2007-620).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zuo, F., Gu, Q., Peng, J. et al. Simultaneous Quantification of Methionine-Related Metabolites and Co-factors in IPEC-J2 and PIEC Cells by LC–MS/MS. Chromatographia 83, 361–371 (2020). https://doi.org/10.1007/s10337-019-03852-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10337-019-03852-4