Abstract
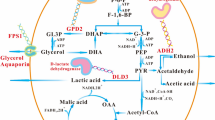
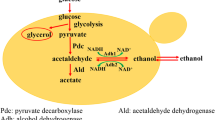
During alcohol fermentation, Saccharomyces cerevisiae produces organic acids, including succinate, acetate, and malate. Since malate contributes to the pleasant flavor of sake (a Japanese alcoholic beverage), various methods for breeding high-malate-producing yeast have been developed. We previously isolated a high-malate-producing strain and found that a missense mutation in GID4 was responsible for the high-malate-producing phenotype. Gid4 is a component of the GID (glucose-induced degradation-deficient) complex and stimulates the catabolic degradation of gluconeogenic enzymes. In this study, the mechanism by which this mutation led to high malate production in yeast cells was investigated. The evaluation of disruptants and mutants of gluconeogenic enzymes revealed that cytosolic malate dehydrogenase (Mdh2) participated in the malate production. Furthermore, target proteome analysis indicated that an increase in malate production resulted from the accumulation of Mdh2 in gid4 disruptant due to the loss of GID complex–mediated degradation. Next, we investigated the effects of GID protein–coding genes (GID1–GID9) on organic acid production and enzyme expression profiles in yeast. The disruptants of GID1, 2, 3, 4, 5, 8, and 9 exhibited high malate production. Comparison of protein abundance among the GID disruptants revealed variations in protein expression profiles, including in glycolysis and tricarboxylic acid cycle–related enzymes. The high-malate-producing disruptants showed the activation of several glycolytic enzymes and a reduction in enzymes involved in the conversion of pyruvate to ethanol. Our results suggest that high-malate-producing disruptants adapt their metabolism to produce malate in excess via the regulation of protein expression in glucose assimilation and ethanol fermentation.
Key Points
-
An increase in malate level of GID4 mutant resulted from the accumulation of Mdh2.
-
The disruptants of GID1, 2, 3, 4, 5, 8, and 9 showed high malate production.
-
The protein expression profiles in the GID disruptants differed from one another.





Similar content being viewed by others
References
Aikawa M, Suizu T, Ichikawa E, Kawato A, Abe Y, Imayasu S (1992) Breeding of higher malic acid-productive mutants from Saccharomyces cerevisiae Kyokai No.7. Hakkokogaku 70:473–477
Akada R, Matsuo K, Aritomi K, Nishizawa Y (1999) Construction of recombinant sake yeast containing a dominant FAS2 mutation without extraneous sequences by a two-step gene replacement protocol. J Biosci Bioeng 87(1):43–48. https://doi.org/10.1016/s1389-1723(99)80006-1
Amerik A, Swaminathan S, Krantz BA, Wilkinson KD, Hochstrasser M (1997) In vivo disassembly of free polyubiquitin chains by yeast Ubp14 modulates rates of protein degradation by the proteasome. EMBO J 16(16):4826–4838. https://doi.org/10.1093/emboj/16.16.4826
Arikawa Y, Kobayashi M, Kodaira R, Shimosaka M, Muratsubaki H, Enomoto K, Okazaki M (1999) Isolation of sake yeast strains possessing various levels of succinate- and/or malate-producing abilities by gene disruption or mutation. J Biosci Bioeng 87(3):333–339. https://doi.org/10.1016/s1389-1723(99)80041-3
Asano T, Kurose N, Tarumi S (2001) Isolation of high-malate-producing sake yeasts from low-maltose-assimilating mutants. J Biosci Bioeng 92(5):429–433. https://doi.org/10.1263/jbb.92.429
Bakker BM, Overkamp KM, van Maris AJ, Kötter P, Luttik MA, van Dijken JP, Pronk JT (2001) Stoichiometry and compartmentation of NADH metabolism in Saccharomyces cerevisiae. FEMS Microbiol Rev 25(1):15–37. https://doi.org/10.1111/j.1574-6976.2001.tb00570.x
Bereman MS, MacLean B, Tomazela DM, Liebler DC, MacCoss MJ (2012) The development of selected reaction monitoring methods for targeted proteomics via empirical refinement. Proteomics 12(8):1134–1141. https://doi.org/10.1002/pmic.201200042
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1006/abio.1976.9999
Braun B, Pfirrmann T, Menssen R, Hofmann K, Scheel H, Wolf DH (2011) Gid9, a second RING finger protein contributes to the ubiquitin ligase activity of the Gid complex required for catabolite degradation. FEBS Lett 585(24):3856–3861. https://doi.org/10.1016/j.febslet.2011.10.038
Chen SJ, Wu X, Wadas B, Oh JH, Varshavsky A (2017) An N-end rule pathway that recognizes proline and destroys gluconeogenic enzymes. Science 355:eaal6323. https://doi.org/10.1126/science.aal3655
Chi Z, Wang ZP, Wang GY, Khan I, Chi ZM (2016) Microbial biosynthesis and secretion of L-malic acid and its applications. Crit Rev Biotechnol 36(1):99–107. https://doi.org/10.3109/07388551.2014.924474
Costenoble R, Picotti RL, Stallmach R, Heinemann M, Sauer U, Aebersold R (2011) Comprehensive quantitative analysis of central carbon and amino-acid metabolism in Saccharomyces cerevisiae under multiple conditions by targeted proteomics. Mol Syst Biol 7:464. https://doi.org/10.1038/msb.2010.122
Dong C, Zhang H, Li L, Tempel W, Loppnau P, Min J (2018) Molecular basis of GID4-mediated recognition of degrons for the pro/N-end rule pathway. Nat Chem Biol 14(5):466–473. https://doi.org/10.1038/s41589-018-0036-1
Eskes E, Deprez MA, Wilms T, Winderickx J (2018) pH homeostasis in yeast; the phosphate perspective. Curr Genet 64(1):155–161. https://doi.org/10.1007/s00294-017-0743-2
Hung GC, Brown CR, Wolfe AB, Liu J, Chiang HL (2004) Degradation of the gluconeogenic enzymes fructose-1,6-bisphosphatase and malate dehydrogenase is mediated by distinct proteolytic pathways and signaling events. J Biol Chem 279(47):49138–49150. https://doi.org/10.1074/jbc.M404544200
Ito H, Fukuda Y, Murata K, Kimura A (1983) Transformation of intact yeast cells treated with alkali cations. J Bacteriol 153(1):163–168
Kamiie J, Ohtsuki S, Iwase R, Ohmine K, Katsukura Y, Yanai K, Sekine Y, Uchida Y, Ito S, Terasaki T (2008) Quantitative atlas of membrane transporter proteins: development and application of a highly sensitive simultaneous LC/MS/MS method combined with novel in-silico peptide selection criteria. Pharm Res 25(6):1469–1483. https://doi.org/10.1007/s11095-008-9532-4
Kosugi S, Kiyoshi K, Oba T, Kusumoto K, Kadokura T, Nakazato A, Nakayama S (2014) Isolation of a high malic and low acetic acid-producing sake yeast Saccharomyces cerevisiae strain screened from respiratory inhibitor 2,4-dinitrophenol (DNP)-resistant strains. J Biosci Bioeng 117(1):39–44. https://doi.org/10.1016/j.jbiosc.2013.06.016
Liu H, Pfirrmann T (2019) The Gid-complex: an emerging player in the ubiquitin ligase league. Biol Chem 400(11):1429–1441. https://doi.org/10.1515/hsz-2019-0139
MacLean B, Tomazela DM, Shulman N, Chambers M, Finney GL, Frewen B, Kern R, Tabb DL, Liebler DC, MacCoss MJ (2010) Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics. 26(7):966–968. https://doi.org/10.1093/bioinformatics/btq054
Matsuda F, Ogura T, Tomita A, Hirano I, Shimizu H (2015) Nano-scale liquid chromatography coupled to tandem mass spectrometry using the multiple reaction monitoring mode based quantitative platform for analyzing multiple enzymes associated with central metabolic pathways of Saccharomyces cerevisiae using ultra fast mass spectrometry. J Biosci Bioeng 119(1):117–120. https://doi.org/10.1016/j.jbiosc.2014.06.010
Matsuda F, Kinoshita S, Nishino S, Tomita A, Shimizu H (2017) Targeted proteome analysis of single-gene deletion strains of Saccharomyces cerevisiae lacking enzymes in the central carbon metabolism. PLoS One 12(2):e0172742. https://doi.org/10.1371/journal.pone.0172742
Menssen R, Schweiggert J, Schreiner J, Kusevic D, Reuther J, Braun B, Wolf DH (2012) Exploring the topology of the Gid complex, the E3 ubiquitin ligase involved in catabolite-induced degradation of gluconeogenic enzymes. J Biol Chem 287(30):25602–25614. https://doi.org/10.1074/jbc.M112.363762
Menssen R, Bui K, Wolf DH (2018) Regulation of the Gid ubiquitin ligase recognition subunit Gid4. FEBS Lett 592(19):3286–3294. https://doi.org/10.1002/1873-3468.13229
Metsalu T, Vilo J (2015) ClustVis: a web tool for visualizing clustering of multivariate data using principal component analysis and heatmap. Nucleic Acids Res 43(W1):W566–W570. https://doi.org/10.1093/nar/gkv468
Motomura S, Horie K, Kitagaki H (2012) Mitochondrial activity of sake brewery yeast affects malic and succinic acid production during alcoholic fermentation. J Inst Brew 118(1):22–26
Negoro H, Kotaka A, Matsumura K, Tsutsumi H, Hata Y (2016) Enhancement of malate-production and increase in sensitivity to dimethyl succinate by mutation of the VID24 gene in Saccharomyces cerevisiae. J Biosci Bioeng 121(6):665–671. https://doi.org/10.1016/j.jbiosc.2015.11.012
Picotti P, Bodenmiller B, Mueller LN, Domon B, Aebersold R (2009) Full dynamic range proteome analysis of S. cerevisiae by targeted proteomics. Cell 138(4):795–806. https://doi.org/10.1016/j.cell.2009.05.051
Rappsilber J, Mann M, Ishihama Y (2007) Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat Protoc 2(8):1896–1906. https://doi.org/10.1038/nprot.2007.261
Regelmann J, Schüle T, Josupeit FS, Horak J, Rose M, Entian KD, Thumm M, Wolf DH (2003) Catabolite degradation of fructose-1,6-bisphosphatase in the yeast Saccharomyces cerevisiae: a genome-wide screen identifies eight novel GID genes and indicates the existence of two degradation pathways. Mol Biol Cell 14(4):1652–1663. https://doi.org/10.1091/mbc.e02-08-0456
Santt O, Pfirrmann T, Braun B, Juretschke J, Kimmig P, Scheel H, Hofmann K, Thumm M, Wolf DH (2008) The yeast GID complex, a novel ubiquitin ligase (E3) involved in the regulation of carbohydrate metabolism. Mol Biol Cell 19(8):3323–3333. https://doi.org/10.1091/mbc.e08-03-0328
Snowdon C, Hlynialuk C, van der Merwe G (2008) Components of the Vid30c are needed for the rapamycin-induced degradation of the high-affinity hexose transporter Hxt7p in Saccharomyces cerevisiae. FEMS Yeast Res 8(2):204–216. https://doi.org/10.1111/j.1567-1364.2007.00327.x
Uchida Y, Tachikawa M, Obuchi W, Hoshi Y, Tomioka Y, Ohtsuki S, Terasaki T (2013) A study protocol for quantitative targeted absolute proteomics (QTAP) by LC-MS/MS: application for inter-strain differences in protein expression levels of transporters, receptors, claudin-5, and marker proteins at the blood-brain barrier in ddY, FVB, and C57BL/6J mice. Fluids Barriers CNS 10(1):21. https://doi.org/10.1186/2045-8118-10-21
Uebayashi K, Shimizu H, Matsuda F (2018) Comparative analysis of fermentation and enzyme expression profiles among industrial Saccharomyces cerevisiae strains. Appl Microbiol Biotechnol 102(16):7071–7081. https://doi.org/10.1007/s00253-018-9128-9
Wach A, Brachat A, Pöhlmann R, Philippsen P (1994) New heterologous modules for classical or PCR-based gene disruptions in Saccharomyces cerevisiae. Yeast 10(13):1793–1808. https://doi.org/10.1002/yea.320101310
Zelle RM, de Hulster E, van Winden WA, de Waard P, Dijkema C, Winkler AA, Geertman JM, van Dijken JP, Pronk JT, van Maris AJ (2008) Malic acid production by Saccharomyces cerevisiae: engineering of pyruvate carboxylation, oxaloacetate reduction, and malate export. Appl Environ Microbiol 74(9):2766–2777. https://doi.org/10.1128/AEM.02591-07
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by Hiroaki Negoro and Fumio Matsuda. The first draft of the manuscript was written by Hiroaki Negoro and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(XLSX 56 kb)
Rights and permissions
About this article
Cite this article
Negoro, H., Matsumura, K., Matsuda, F. et al. Effects of mutations of GID protein–coding genes on malate production and enzyme expression profiles in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 104, 4971–4983 (2020). https://doi.org/10.1007/s00253-020-10573-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10573-4