Abstract
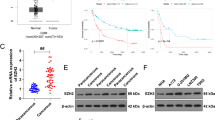
Glioma is characterized by high morbidity, high mortality and poor prognosis. Recent studies exhibited that lncRNA CCAT2 is overexpressed in glioma and promotes glioma progression, but the specific molecular biological mechanism remains to be determined. We performed qRT-PCR to evaluate the expression of related genes, Western blotting analysis to measure protein levels, colony formation assay to detect the proliferative ability of glioma cells, flow cytometry to measure cell apoptosis, bioinformatics analysis and dual luciferase assay to verify the binding sites and the targeted regulatory relationship in A172 and U251 cell lines and tube formation assay to determine endothelial angiogenesis. LncRNA CCAT2 and VEGFA were highly expressed, while miR-424 was expressed at low levels in NHA cells. Furthermore, knockdown of lncRNA CCAT2 decreased cell proliferation, increased cell apoptosis and inhibited endothelial angiogenesis in glioma. Moreover, lncRNA CCAT2 shared a complementary sequence with miR-424 which in turn directly bound to the 3′-UTR of VEGFA. Further investigation indicated that lncRNA CCAT2 promoted cell proliferation and endothelial angiogenesis by inducing the PI3K/AKT signalling pathway in glioma. The oncogenic lncRNA CCAT2 is highly associated with the development of glioma and exerts its function by upregulating VEGFA via miR-424.







Similar content being viewed by others
References
Rich JN, Bigner DD (2004) Development of novel targeted therapies in the treatment of malignant glioma. Nat Rev Drug Discov 3:430–446. https://doi.org/10.1038/nrd1380
Jiang Y, Song Y, Wang R, Hu T, Zhang D, Wang Z, Tie X, Wang M, Han S (2019) NFAT1-mediated regulation of NDEL1 promotes growth and invasion of glioma stem-like cells. Cancer Res 79:2593–2603. https://doi.org/10.1158/0008-5472.CAN-18-3297
Ma J, Benitez JA, Li J, Miki S, Ponte de Albuquerque C, Galatro T, Orellana L, Zanca C, Reed R, Boyer A, Koga T, Varki NM, Fenton TR, Nagahashi Marie SK, Lindahl E, Gahman TC, Shiau AK, Zhou H, DeGroot J, Sulman EP, Cavenee WK, Kolodner RD, Chen CC, Furnari FB (2019) Inhibition of nuclear PTEN tyrosine phosphorylation enhances glioma radiation sensitivity through attenuated DNA repair. Cancer Cell 35:816. https://doi.org/10.1016/j.ccell.2019.04.011
Wang J, Xu SL, Duan JJ, Yi L, Guo YF, Shi Y, Li L, Yang ZY, Liao XM, Cai J, Zhang YQ, Xiao HL, Yin L, Wu H, Zhang JN, Lv SQ, Yang QK, Yang XJ, Jiang T, Zhang X, Bian XW, Yu SC (2019) Invasion of white matter tracts by glioma stem cells is regulated by a NOTCH1-SOX2 positive-feedback loop. Nat Neurosci 22:91–105. https://doi.org/10.1038/s41593-018-0285-z
Malla RR, Gopinath S, Gondi CS, Alapati K, Dinh DH, Gujrati M, Rao JS (2011) Cathepsin B and uPAR knockdown inhibits tumor-induced angiogenesis by modulating VEGF expression in glioma. Cancer Gene Ther 18:419–434. https://doi.org/10.1038/cgt.2011.9
Ping YF, Yao XH, Jiang JY, Zhao LT, Yu SC, Jiang T, Lin MC, Chen JH, Wang B, Zhang R, Cui YH, Qian C, Wang J, Bian XW (2011) The chemokine CXCL12 and its receptor CXCR4 promote glioma stem cell-mediated VEGF production and tumour angiogenesis via PI3K/AKT signalling. J Pathol 224:344–354. https://doi.org/10.1002/path.2908
Yang HK, Chen H, Mao F, Xiao QG, Xie RF, Lei T (2017) Downregulation of LRIG2 expression inhibits angiogenesis of glioma via EGFR/VEGF-A pathway. Oncol Lett 14:4021–4028. https://doi.org/10.3892/ol.2017.6671
Ballester LY, Moghadamtousi SZ, Leeds NE, Huse JT, Fuller GN (2019) Coexisting FGFR3 p.K650T mutation in two FGFR3-TACC3 fusion glioma cases. Acta Neuropathol Commun 7:63. https://doi.org/10.1186/s40478-019-0721-7
Bao S, Wu Q, Sathornsumetee S, Hao Y, Li Z, Hjelmeland AB, Shi Q, McLendon RE, Bigner DD, Rich JN (2006) Stem cell-like glioma cells promote tumor angiogenesis through vascular endothelial growth factor. Cancer Res 66:7843–7848. https://doi.org/10.1158/0008-5472.CAN-06-1010
Zhang Y, Zhang N, Dai B, Liu M, Sawaya R, Xie K, Huang S (2008) FoxM1B transcriptionally regulates vascular endothelial growth factor expression and promotes the angiogenesis and growth of glioma cells. Cancer Res 68:8733–8742. https://doi.org/10.1158/0008-5472.CAN-08-1968
Zhao LN, Wang P, Liu YH, Cai H, Ma J, Liu LB, Xi Z, Li ZQ, Liu XB, Xue YX (2017) MiR-383 inhibits proliferation, migration and angiogenesis of glioma-exposed endothelial cells in vitro via VEGF-mediated FAK and Src signaling pathways. Cell Signal 30:142–153. https://doi.org/10.1016/j.cellsig.2016.09.007
Gao YF, Liu JY, Mao XY, He ZW, Zhu T, Wang ZB, Li X, Yin JY, Zhang W, Zhou HH, Liu ZQ (2019) LncRNA FOXD1-AS1 acts as a potential oncogenic biomarker in glioma. CNS Neurosci Ther 26:66–75. https://doi.org/10.1111/cns.13152
Liu N, Hu G, Wang H, Wang Y, Guo Z (2019) LncRNA BLACAT1 regulates VASP expression via binding to miR-605-3p and promotes giloma development. J Cell Physiol 234:22144. https://doi.org/10.1002/jcp.28778
Quan J, Pan X, Zhao L, Li Z, Dai K, Yan F, Liu S, Ma H, Lai Y (2018) LncRNA as a diagnostic and prognostic biomarker in bladder cancer: a systematic review and meta-analysis. Onco Targets Ther 11:6415–6424. https://doi.org/10.2147/OTT.S167853
Tian T, Wang M, Lin S, Guo Y, Dai Z, Liu K, Yang P, Dai C, Zhu Y, Zheng Y, Xu P, Zhu W, Dai Z (2018) The impact of lncRNA dysregulation on clinicopathology and survival of breast cancer: a systematic review and meta-analysis. Mol Ther Nucleic Acids 12:359–369. https://doi.org/10.1016/j.omtn.2018.05.018
Monnier P, Martinet C, Pontis J, Stancheva I, Ait-Si-Ali S, Dandolo L (2013) H19 lncRNA controls gene expression of the Imprinted Gene Network by recruiting MBD1. Proc Natl Acad Sci USA 110:20693–20698. https://doi.org/10.1073/pnas.1310201110
Zhou J, Yang L, Zhong T, Mueller M, Men Y, Zhang N, Xie J, Giang K, Chung H, Sun X, Lu L, Carmichael GG, Taylor HS, Huang Y (2015) H19 lncRNA alters DNA methylation genome wide by regulating S-adenosylhomocysteine hydrolase. Nat Commun 6:10221. https://doi.org/10.1038/ncomms10221
Wu ZR, Yan L, Liu YT, Cao L, Guo YH, Zhang Y, Yao H, Cai L, Shang HB, Rui WW, Yang G, Zhang XB, Tang H, Wang Y, Huang JY, Wei YX, Zhao WG, Su B, Wu ZB (2018) Inhibition of mTORC1 by lncRNA H19 via disrupting 4E-BP1/Raptor interaction in pituitary tumours. Nat Commun 9:4624. https://doi.org/10.1038/s41467-018-06853-3
Jia P, Cai H, Liu X, Chen J, Ma J, Wang P, Liu Y, Zheng J, Xue Y (2016) Long non-coding RNA H19 regulates glioma angiogenesis and the biological behavior of glioma-associated endothelial cells by inhibiting microRNA-29a. Cancer Lett 381:359–369. https://doi.org/10.1016/j.canlet.2016.08.009
Han Y, Li X, Yan J, Ma C, Zheng X, Zhang J, Zhang D, Meng C, Zhang Z, Ji X, He F (2019) Knockdown of LncRNA PVT1 inhibits glioma progression by regulating miR-424 expression. Oncol Res 27:681–690. https://doi.org/10.3727/096504018X15424939990246
Jin C, Li M, Ouyang Y, Tan Z, Jiang Y (2017) MiR-424 functions as a tumor suppressor in glioma cells and is down-regulated by DNA methylation. J Neurooncol 133:247–255. https://doi.org/10.1007/s11060-017-2438-4
Hua F, Li CH, Chen XG, Liu XP (2018) Long Noncoding RNA CCAT2 knockdown suppresses tumorous progression by sponging miR-424 in epithelial ovarian cancer. Oncol Res 26:241–247. https://doi.org/10.3727/096504017X14953948675412
Gu L, Chen H, Tuo J, Gao X, Chen L (2010) Inhibition of experimental choroidal neovascularization in mice by anti-VEGFA/VEGFR2 or non-specific siRNA. Exp Eye Res 91:433–439. https://doi.org/10.1016/j.exer.2010.06.019
Hu S, Liu Y, You T, Zhu L (2018) Semaphorin 7A promotes VEGFA/VEGFR2-mediated angiogenesis and intraplaque neovascularization in ApoE(-/-) mice. Front Physiol 9:1718. https://doi.org/10.3389/fphys.2018.01718
Wang H, Smith GW, Yang Z, Jiang Y, McCloskey M, Greenberg K, Geisen P, Culp WD, Flannery J, Kafri T, Hammond S, Hartnett ME (2013) Short hairpin RNA-mediated knockdown of VEGFA in Muller cells reduces intravitreal neovascularization in a rat model of retinopathy of prematurity. Am J Pathol 183:964–974. https://doi.org/10.1016/j.ajpath.2013.05.011
Liu Q, Liao F, Wu H, Cai T, Yang L, Fang J (2016) Different expression of miR-29b and VEGFA in glioma. Artif Cells Nanomed Biotechnol 44:1927–1932. https://doi.org/10.3109/21691401.2015.1111237
Zhao P, Chen A, Qi Q, Zhou W, Feng Z, Wang J, Yang N, Li X, Wang J, Huang Q, Huang B (2017) Impact of VEGFA polymorphisms on glioma risk in Chinese. Oncotarget 8:83712–83722. https://doi.org/10.18632/oncotarget.19380
Haryu S, Saito R, Jia W, Shoji T, Mano Y, Sato A, Kanamori M, Sonoda Y, Sampetrean O, Saya H, Tominaga T (2018) Convection-enhanced delivery of sulfasalazine prolongs survival in a glioma stem cell brain tumor model. J Neurooncol 136:23–31. https://doi.org/10.1007/s11060-017-2621-7
Wang X, Yang K, Xie Q, Wu Q, Mack SC, Shi Y, Kim LJY, Prager BC, Flavahan WA, Liu X, Singer M, Hubert CG, Miller TE, Zhou W, Huang Z, Fang X, Regev A, Suva ML, Hwang TH, Locasale JW, Bao S, Rich JN (2017) Purine synthesis promotes maintenance of brain tumor initiating cells in glioma. Nat Neurosci 20:661–673. https://doi.org/10.1038/nn.4537
Zukotynski KA, Vajapeyam S, Fahey FH, Kocak M, Brown D, Ricci KI, Onar-Thomas A, Fouladi M, Poussaint TY (2017) Correlation of (18)F-FDG PET and MRI apparent diffusion coefficient histogram metrics with survival in diffuse intrinsic pontine glioma: a report from the Pediatric Brain Tumor Consortium. J Nucl Med 58:1264–1269. https://doi.org/10.2967/jnumed.116.185389
Barone F, Alberio N, Iacopino DG, Giammalva GR, D'Arrigo C, Tagnese W, Graziano F, Cicero S, Maugeri R (2018) Brain mapping as helpful tool in brain glioma surgical treatment-toward the "perfect surgery"? Brain Sci 8:192. https://doi.org/10.3390/brainsci8110192
Di K, Lomeli N, Bota DA, Das BC (2019) Magmas inhibition as a potential treatment strategy in malignant glioma. J Neurooncol 141:267–276. https://doi.org/10.1007/s11060-018-03040-8
Kline C, Liu SJ, Duriseti S, Banerjee A, Nicolaides T, Raber S, Gupta N, Haas-Kogan D, Braunstein S, Mueller S (2018) Reirradiation and PD-1 inhibition with nivolumab for the treatment of recurrent diffuse intrinsic pontine glioma: a single-institution experience. J Neurooncol 140:629–638. https://doi.org/10.1007/s11060-018-2991-5
Luo J, Qu J, Wu DK, Lu ZL, Sun YS, Qu Q (2017) Long non-coding RNAs: a rising biotarget in colorectal cancer. Oncotarget 8:22187–22202. https://doi.org/10.18632/oncotarget.14728
Zeng J, Du T, Song Y, Gao Y, Li F, Wu R, Chen Y, Li W, Zhou H, Yang Y, Pei Z (2017) Knockdown of long noncoding RNA CCAT2 inhibits cellular proliferation, invasion, and epithelial-mesenchymal transition in glioma cells. Oncol Res 25:913–921. https://doi.org/10.3727/096504016X14792098307036
Guo H, Hu G, Yang Q, Zhang P, Kuang W, Zhu X, Wu L (2016) Knockdown of long non-coding RNA CCAT2 suppressed proliferation and migration of glioma cells. Oncotarget 7:81806–81814. https://doi.org/10.18632/oncotarget.13242
Yu Y, Nangia-Makker P, Farhana L, Majumdar APN (2017) A novel mechanism of lncRNA and miRNA interaction: CCAT2 regulates miR-145 expression by suppressing its maturation process in colon cancer cells. Mol Cancer 16:155. https://doi.org/10.1186/s12943-017-0725-5
Cai Y, Li X, Shen P, Zhang D (2018) CCAT2 is an oncogenic long non-coding RNA in pancreatic ductal adenocarcinoma. Biol Res 51:1. https://doi.org/10.1186/s40659-017-0149-0
Dong P, Xiong Y, Yue J, Hanley JB, Kobayashi N, Todo Y, Watari H (2019) Exploring lncRNA-mediated regulatory networks in endometrial cancer cells and the tumor microenvironment: advances and challenges. Cancers (Basel) 11:234. https://doi.org/10.3390/cancers11020234
Ruan R, Zhao XL (2018) LncRNA CCAT2 enhances cell proliferation via GSK3beta/beta-catenin signaling pathway in human osteosarcoma. Eur Rev Med Pharmacol Sci 22:2978–2984. https://doi.org/10.26355/eurrev_201805_15053
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sun, SL., Shu, YG. & Tao, MY. LncRNA CCAT2 promotes angiogenesis in glioma through activation of VEGFA signalling by sponging miR-424. Mol Cell Biochem 468, 69–82 (2020). https://doi.org/10.1007/s11010-020-03712-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-020-03712-y