Abstract
The first member of the velvet family of proteins, VeA, regulates sexual development and secondary metabolism in the filamentous fungus Aspergillus nidulans. In our study, through comparative proteome analysis using wild type and veA-deletion strains, new putative regulators of sexual development were identified and functionally analyzed. Among these, SvfA, containing a yeast survival factor 1 domain, plays multiple roles in the growth and differentiation of A. nidulans. Deletion of the svfA gene resulted in increased sensitivity to oxidative and cold stress as in yeast. The svfA-deletion strain showed an increase in bi-polar germination and a decrease in radial growth rate. The deletion strain formed structurally abnormal conidiophores and thus produced lower amounts of conidiospores during asexual development. The svfA-deletion strain produced few Hülle cells and small cleistothecia with no ascospores, indicating the requirement of svfA for the completion of sexual development. Transcription and genetic analyses indicated that SvfA modulates the expression of key development regulatory genes. Western blot analysis revealed two forms of SvfA. The larger form showed sexual-specific and VeA-dependent production. Also, the deletion of svfA caused decreased ST (sterigmatocystin) production. We propose that SvfA is a novel central regulator of growth, differentiation and secondary metabolism in A. nidulans.
Similar content being viewed by others
Introduction
Aspergillus nidulans is a model filamentous fungus that belongs to the phylum Ascomycota. Both asexual and sexual cycles allow the study of various cellular events such as development, stress responses, and secondary metabolism1,2,3,4. Asexual development proceeds in multiple steps with special organs: a foot cell, stalk, vesicle, metulae, phialides, and conidiospores5. While the vegetative cell goes through asexual development under light conditions, sexual development is favored under dark and hypoxic conditions6. In the sexual reproductive organ, cleistothecium, numerous asci containing eight red-purple ascospores are developed1,7,8.
The velvet family proteins, including VeA, VelB, VelC, and VosA, comprise highly conserved fungal specific regulators in ascomycetes and basidiomycetes9. This superfamily plays critical roles in development and secondary metabolism by forming complexes with multiple interacting partners4,10,11,12. VeA, the first identified protein in this family, controls the asexual or sexual development in response to external signals such as light and air. This regulatory function of VeA is dependent on its localization. Under light conditions, VeA disperses in both the cytoplasm and the nuclei. Under dark conditions, the VelB-VeA dimer enters the nucleus with the help of importin α (KapA), and forms a trimeric complex with the global regulator of secondary metabolism, LaeA4,11,13,14,15. Sexual development and secondary metabolism are induced by the VelB-VeA-LaeA complex and VelC12. Asexual development is inhibited by the VosA homodimer9. However, the presence of light blocks the entry of VelB-VeA dimer into the nucleus. One of the LaeA-like methyltransferases, LlmF, interacts with VeA in the cytoplasm, which causes VelB to interact with another VelB or VosA but not VeA. The accumulation of VelB-VelB and VelB-VosA complexes in the nucleus positively regulates asexual development4,16.
While strains with veA1 point mutation produce more conidia and fewer cleistothecia than the wild type (WT), the veA-deletion strain (ΔveA) fails to produce cleistothecia even under dark conditions17,18. Expression of genes such as aflR and stcU, which are required for the biosynthesis of sterigmatocystin (ST, a precursor of aflatoxin), is decreased in the ΔveA strain19.
Other direct interacting partners of VeA are velvet interacting proteins (Vips). VipC, a methyltransferase, is required for repression of sexual development and can interact with a membrane protein, VapA, and another methyltransferase, VapB. VipC-VapB dimers are detached from the VapA-VipC-VapB complex at the plasma membrane by external signals and prevent VeA from entering the nucleus20.
Although the velvet family and interactors are known to be involved in sexual development, further studies are needed to find novel factors and to understand the molecular process of sexual development in A. nidulans. Here, we report the results from comparative proteome analyses of WT and ΔveA strains to identify novel VeA-dependent proteins (Vdps), the expression of which could be affected by the absence of VeA during development. Among the 144 proteins identified, four Vdps showing significant reductions in their intensity during the sexual stage in the ΔveA strain were analyzed by gene-deletion experiments and functional assays. In this report, we show that SvfA, a homolog of yeast survival factor 1, is required for response to oxidative- and cold-stress, and is a novel regulator that plays multiple roles in the regulation of growth and differentiation, which is essential for completion of sexual development in A. nidulans.
Results
Detection and identification of proteins affected by VeA
VeA regulates development and secondary metabolism in A. nidulans through its interactions with other regulatory proteins, including VelB, VosA, LaeA, and Vips, and through feedback control on the expression of various genes15,20. For screening the VeA-target proteins, mycelial balls of veA-deletion strain (ΔveA) and wild type strain (veA+), grown in liquid complete medium, were transferred to agar plates to induce sexual development21. Whole-cell lysates from these samples, harvested at different developmental stages, were analyzed on 2-DE22,23 in triplicates, with independently harvested samples (Supplementary Fig. S1). About 2,400 protein spots were detected on the 2-DE gels (Fig. 1A). By comparing the gel profiles of veA+ and ΔveA strains at the same stage, spots showing significant changes in their intensity were selected and subjected to in-gel tryptic digestion and MALDI-TOF. Among the 200 spots analyzed, only 144 proteins were identified. Out of the 76 protein that were down-regulated during the sexual stage in the ΔveA strain, 56 were significantly enriched in the FunCat categories (P ≤ 0.05). The top 10 enriched FunCat categories were as follows: cellular response to farnesol, ethanol biosynthetic process, cellular response to osmotic stress, acetate metabolic process, oxalate metabolic process, cellular response to oxidative stress, establishment or maintenance of cell polarity, UDP-glucose metabolic process, proteasomal ubiquitin-independent protein catabolic process, and galactose metabolic process (Fig. 1B and Supplementary Table S3).
Proteome analysis of WT and ΔveA strains. (A) A representative map of 2-DE analysis from three independent replicates. Equal amounts of total proteins at the vegetative stage (V9; 9 h) and sexual stage (S6; 6 h) were separated by 2-DE and visualized with silver staining. (B) Gene ontology (GO) enrichment analysis of down-regulated proteins in the ΔveA strains. In each case, the top 10 significantly enriched biological processes are shown along with p-value computed using the Benjamini-Hochberg procedure54. The GO enrichment analysis was performed using FungiFun2 webserver55.
To screen for the Vdps specific to sexual development, the protein spots showing more than 2-fold reduction in the ΔveA strain during sexual development were selected for functional analysis. Cellular functions of most Vdps, except CatB and FbpA, have not been previously characterized and their predicted roles, based on the GO annotation of AspGD (www.AspGD.org), are summarized in Table 1. To investigate the function of Vdps during development of A. nidulans, their corresponding genes (svfA, vdpC, vdpF, and vdpJ) were deleted and these deletions were confirmed by PCR and Southern blotting (Supplementary Fig. S2 for the svfA-deletion). When parameters like vegetative growth, responses to osmotic-, temperature-, oxidative-, cell wall- and cell membrane-stresses, and asexual and sexual development were examined, no significant abnormalities were seen in these deletion strains except svfA-deletion strain (ΔsvfA) (Supplementary Fig. S3). Thereafter, ΔsvfA strain was used for further investigations.
SvfA regulates vegetative growth and functions in oxidative- and cold-stress responses
The ΔsvfA mutant showed retarded radial growth on solid medium (Fig. 2A,B) and reduction in biomass production (Fig. 2C) with smaller mycelial balls (data not shown) in liquid culture. These growth defects were reversed by the re-introduction of the svfA gene (Fig. 2A–C for C’svfA; complementation strain). In S. cerevisiae, the Svf1 protein is required for survival under conditions of oxidative stress and cold stress24. When the sensitivity of the ΔsvfA mutant to menadione, H2O2, and low temperature (20 °C) was tested, the ΔsvfA mutant was sensitive to chemical induction of reactive oxygen species (ROS) (Fig. 2D) and cold stress (Fig. 2E). These data suggested that, as in S. cerevisiae, SvfA function was required for survival of A. nidulans during oxidative stress and cold stress.
Growth patterns of different strains. (A) Colony morphology. Spores of WT, ΔsvfA, and C’svfA strains were inoculated on YCMM plate and incubated for 2 days at 37 °C. (B) Radial growth. Over 9 days, the colony diameter of each culture from the point of inoculation was measured daily. (C) Mycelial production. Over 48 h, dry weight of each culture from the point of inoculation in liquid medium was measured. (D) Sensitivity to oxidative stress. Spores with 10-fold serial dilutions were spotted on YCMM containing menadione and hydrogen peroxide (H2O2) and incubated at 37 °C for 2 days. (E) Sensitivity to cold stress. Spores with 10-fold serial dilutions were spotted on YCMM at 20 °C for 6 days.
SvfA affects conidial germination
When germination was observed in a time-dependent manner in GMM medium (minimal medium containing 1% glucose), WT conidia formed an unipolar germ tube after 4 h incubation (8.7% at 4 h, 25.3% at 5 h, and 80% at 6 h), and a few (2%) bipolar germ tubes in about 6 h (Fig. 3A,B). In contrast, ΔsvfA conidia formed abnormally long, bipolar germ tubes from the beginning (6% at 4 h, 6.7% at 5 h, and 9.3% at 6 h) (Fig. 3A,C). When germination was observed in MM broth without glucose, unipolar germination was observed in both WT and ΔsvfA, after 7 h incubation (Supplementary Fig S4). These results suggest that SvfA influences the establishment of polarity, but not the initiation of conidia germination.
Roles of SvfA in conidial germination. (A) Morphology of germ tube. Conidia of the WT and ΔsvfA strains were incubated in liquid GMM for the indicated time. The proportion of uni-polar and bi-polar germination of conidia in the WT (B) and ΔsvfA strains (C). A total of 50 cells counted at each time point in triplicates, and results are shown as percentages.
SvfA modulates conidiophore development and conidia production
As indicated, the ΔsvfA mutant grew slower than the WT strain, taking 7 days to reach the size of a 5-day WT colony. In addition, colony margins were irregular and colony color changed from yellow to faded-brownish yellow in the ΔsvfA strain (Fig. 4A). Under the stereomicroscope, conidiophore heads in the ΔsvfA strain showed reduction in number and size compared to the WT (Fig. 4B). Microscopic observation revealed a wide variety of abnormalities in conidiophore formation due to lack of SvfA, such as short stalks, branched stalks with abnormal head, and unstructured sterigmata layers (Fig. 4C); all these led to a reduction in conidiospore production. Statistical analysis revealed an approximately 50% decrease in stalk length (Fig. 4D) and 95% reduction in conidiospore production in the ΔsvfA strain (Fig. 4E). In A. nidulans, a central regulatory pathway that controls asexual development is composed of three transcription factors, BrlA, AbaA, and WetA25. The transcript levels of these genes, along with that of vosA, which codes for one of the velvet family proteins involved in spore viability, were evaluated by RT-qPCR. Mycelial balls, produced in liquid YCMM, were shifted to solid MM to induce synchronized asexual development, and RNA was isolated from the cultures at the indicated time, post-induction. The WT strain expressed brlA and vosA, with peaks at 24 and 48 h, and abaA with a peak at 24 h, while the ΔsvfA strain showed significant reduction in the transcript levels of brlA, abaA, and vosA (Fig. 4F). As VosA is reported to affect conidiospore viability9, conidia were collected from WT, ΔsvfA, and C’svfA, grown for 2, 5, and 7 days, and their viability was tested. However, no significant difference among strains was observed (Supplementary Fig. S5). Taken together, these results indicate that SvfA regulates asexual development by regulating the induction of genes that are critical for this process.
Effect of svfA-deletion on asexual development. (A) Colony morphology. Conidia of the WT and ΔsvfA strains were inoculated on solid YCMM and incubated at 37 °C for 5 days (WT) and 7 days (ΔsvfA) until the sizes of the colonies became similar. (B) Conidiophore heads on solid medium (A) as seen under a stereomicroscope. (C) Asexual reproductive organs. Conidia were inoculated on solid agar blocks, and the WT strain was incubated for 24 h whereas the ΔsvfA strain was incubated for 24 h and 28 h, separately. (D) Length of stalks. Mean value of about 50 conidiophores. ∗∗∗P < 0.001. (E) Number of conidia produced. Average of triplicate readings is shown. ∗∗∗P < 0.001. (F) Expression patterns of genes brlA, abaA, and vosA, which are associated with asexual development. Mycelial balls of strains obtained from YCMM liquid culture were shifted to solid GMM to induce asexual development. Total RNA was extracted and RT-qPCR analysis was performed using 18 S rRNA gene as an internal control. ∗∗∗P < 0.001.
SvfA is essential for the completion of sexual development
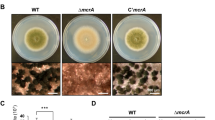
To investigate the effect of svfA-deletion on sexual differentiation, strains were induced to undergo sexual development under dark and hypoxic conditions. The ΔsvfA mutant produced small cleistothecia surrounded by few Hülle cells; this phenotype was rescued by the restoration of a functional svfA gene (Fig. 5A). When cleistothecia were ruptured to observe the formation of asci and ascospores, no ascospore production was observed in the ΔsvfA strain (Fig. 5B). Unlike asexual development, which is controlled by a relatively simple central regulatory pathway25, the mechanism that regulates sexual development is much more complicated in A. nidulans26. For example, esdC and steA play roles in the early sexual stage, and vosA and mutA in the late sexual stage. EsdC has a glycogen binding domain, which is conserved in the beta subunit of the AMPK complex, and plays critical roles in promoting sexual development and regulating conidiation27. SteA is a transcription factor required for sexual development28, while VosA is required for the integrity of both asexual and sexual spores. This probably is the cause of defective cleistothecia, containing very few (~1%) viable ascospores in the ΔvosA mutant9. Alpha-1,3 glucanase MutA (mutanase) is expressed in the Hülle cells to nourish cleistothecia29. When the transcription of early sexual genes, esdC and steA, were analyzed at two different post-induction time-points, transcript levels of both genes were found to be decreased at 24 h in the ΔsvfA strain (Fig. 5C,D). Transcription of late sexual gene vosA reduced by 7-fold at 48 h and 3-fold at 72 h (Fig. 5E), and that of the mutA, another late gene, decreased at 72 h (Fig. 5F). These results indicate that SvfA is a novel regulator that is essential for the completion of sexual development in A. nidulans, and is involved in the transcription of genes associated with early and late sexual development. To examine the possibility of sexual defects caused by insufficient arginine, we studied the phenotypes of the ΔsvfA strain in medium supplemented with arginine. Addition of arginine did not alleviate the defects in colony morphology, asexual development, or sexual development (Supplementary Fig. S6).
Effect of svfA-deletion on sexual development. (A) Sexual reproductive organs produced on MMCA solid medium. Mycelial balls of WT, ΔsvfA, and C’svfA strains were shifted to solid MMCA medium and incubated for 3 days under conditions which induce sexual development. The images were captured under a stereomicroscope. (B) Cleistothecia and ascospores. A cleistothecium was ruptured on a glass slide to observe ascospores (enlarged image). (C–H) RT-qPCR analyses of genes during sexual development using 18 S rRNA gene as internal control. Expression patterns of esdC (C) and steA (D), which are genes involved in early sexual development. Expression patterns of the vosA gene for spore viability (E) and the mutA gene for the mutanase expressed mainly in Hülle cells (F). Expression patterns of veA (G) and nsdD (H), which are developmental regulators that activate sexual development. ∗P < 0.05; ∗∗∗P < 0.001.
SvfA is linked to other developmental processes
Next, we examined the expression of genes for upstream sexual regulators, veA and nsdD. VeA, the founding member of the velvet family, is highly conserved in dimorphic and filamentous fungi11 and activates sexual development and secondary metabolism4,18. NsdD is a GATA type transcription factor required for sexual development, and the gene nsdD is highly expressed in the vegetative and asexual stages of A. nidulans, when compared to the sexual stages30. In the ΔsvfA mutant, both veA and nsdD genes showed very high expression, unlike in the WT (Fig. 5G,H), suggesting that SvfA is required for down-regulation of veA and nsdD to regulate sexual development temporally. Development in A. nidulans is also closely connected to secondary metabolism16. To confirm this, the effect of svfA-deletion on sterigmatocystin (ST) production was tested by thin-layer chromatography (TLC) analysis. The ΔsvfA strain produced a lower amount of ST compared to WT as well as C’svfA strains grown on solid GMM for 4 days (Supplementary Fig. S7A). To monitor the time-course profile of ST production, mycelial balls from liquid GMM were transferred to solid GMM and incubated for 6, 18, 24, and 48 h. The ΔsvfA mutant showed a significantly lower amount of ST production during asexual development, compared to the WT strain (Supplementary Fig. S7B,C). These results suggested that SvfA is required for ST production.
Cytoplasmic localization of SvfA is light-independent
To localize SvfA in the cell, an AYA strain expressing the SvfA fused to a 3xYFP C-terminal tag in the ΔsvfA background was constructed. The AYA strain complemented the svfA-deletion phenotype (Supplementary Fig. S8A). The svfA expression pattern of the AYA strain was similar to that of the WT (Supplementary Fig. S8B). Strains were cultured on coverslips on solid MM at 30 °C for 24 h under light and dark conditions separately. Under both these conditions, SvfA-YFP fusion protein was localized to the cytoplasm and homogenously distributed in the hyphae (Fig. 6).
VeA affects production of a larger form of SvfA protein during sexual development
To investigate the effect of VeA on SvfA protein production during sexual development of A. nidulans, recombinant strains expressing SvfA fused to a FLAG C-terminal tag in the WT and ΔveA backgrounds were generated. Western blot using anti-FLAG antibody revealed that the larger form was expressed only in the WT strain during sexual development (S6) and that the smaller form was expressed in the WT and ΔveA strains during both vegetative and sexual development (Fig. 7A), suggesting that VeA positively modulates production of the sexual-specific larger form of SvfA. In 2-DE analysis, two different protein spots, both identified as SvfA, showed the same isoelectric point (pI 4.8) but different molecular weights (MW 53.35 and 51.93 kDa) and an opposite pattern in intensity changes at the sexual stage (Table 1). These results suggest possible VeA-dependent post-translational modifications, such as glycosylation, which affects the MW but not the pI of the SvfA protein during early sexual development. To study the functional relationship between svfA and veA, svfA was overexpressed in the ΔveA strain to investigate whether the defects in sexual development caused by veA-deletion were reversed. Under the conditions that induced sexual development, the ΔveA;OEsvfA strain did not alleviate the sexual defects, i.e., failed to produce cleistothecia (Fig. 7B). Furthermore, the gene expression pattern of svfA in the ΔveA strain showed no change at 6, 24, and 48 h, and 2-fold increase at 72 h, compared to that in the WT, as indicated by RT-qPCR (Supplementary Fig. S9). Next, the effect of svfA overexpression on sexual development was determined. Unlike the complementation strain (C’svfA), which showed normal sexual development (Fig. 5A), the svfA-overproducing strain (OEsvfA) showed defects similar to those seen in the ΔsvfA strain, which produced small cleistothecia (Fig. 7C). Although the ΔsvfA strain failed to produce ascospores, cleistothecia of the OEsvfA strain contained a few ascospores (Supplementary Fig. S10). Taken together, these data indicate that SvfA functions downstream to VeA together with other VeA-regulated proteins and that normal level of svfA gene expression is crucial for progression and completion of sexual development in A. nidulans.
Effect of SvfA overexpression on sexual development and expression pattern of svfA during sexual development. (A) Upper panel: photograph of Western blotting of WT and ΔveA strains. The SvfA::FLAG fusion proteins were detected using an anti-FLAG antibody at the predicted size of approximately 54.3 kDa and 52.9 kDa. The capital letters V and S indicate the vegetative stage (V) and sexual development (S). Numbers indicate incubation time (h). Lower panel: photograph of SDS-polyacrylamide (8%) gel of total proteins visualized by silver staining. (B) Micrographs of cultured mycelia. Mycelial balls were transferred onto non-inducing medium (GMM containing 0.2% ammonium tartrate as a nitrogen source) or inducing medium (GMM containing 0.6% sodium nitrate as a nitrogen source) and incubated at 37 °C for 6 days under conditions which induce sexual development. Images were captured under a stereomicroscope. (C) Colonies of the WT and ΔsvfA;OEsvfA strains grown for 4 days with sealing and 2 additional days without sealing on non-inducing medium (GMM containing 0.2% ammonium tartrate as a nitrogen source) or inducing medium (GMM containing 0.6% sodium nitrate as a nitrogen source). (D) Proposed model for the involvement of SvfA in asexual and sexual development in A. nidulans. SvfA activates brlA gene expression and thus affects expression of downstream effectors abaA and vosA during asexual development. During sexual development, VeA-dependent post-translational modification activates SvfA, which in turn down-regulates the expression of veA and nsdD, and up-regulates the expression of esdC, steA, vosA, and mutA. SvfA is possibly regulated through the StuA for temporal regulation of both asexual and sexual development. Arrowheads denote positive regulation and flat arrows denote negative regulation. Solid lines denote transcriptional regulation and dash-dotted line denotes translational regulation. Genes, of which transcription were affected by SvfA, are emboldened.
Discussion
In the present study, we identified SvfA, a yeast survival factor 1 (Svf1) homologous protein, as a candidate for novel VeA-dependent protein associated with sexual development in A. nidulans, and provided evidence of its multiple roles in growth and developmental processes.
The survival factor 1 protein was first identified in S. cerevisiae and is required for survival under conditions of oxidative stress and cold stress24. Expression of the svf1 gene in mammalian cells protects them from oxidative stress, which suggests that Svf1 can play a role in protecting yeast cells from ROS, preventing possible cell death24. A recent report also revealed that survival factor SsSvf1 is required for oxidative stress response and full virulence in plant pathogenic fungus Sclerotinia sclerotiorum; SsSvf1-gene silenced strains showed overproduction of ROS, impaired cell wall integrity, and reduced virulence31. The A. nidulans svfA-deletion (ΔsvfA) strain showed sensitivity to chemical induction of reactive oxygen species (Fig. 2D). Although the molecular mechanisms underlying survival factor functions in fungi remain unclear24,31, the results presented in our study showed that SvfA is required for survival under oxidative stress in A. nidulans, as in S. cerevisiae and S. sclerotiorum. It is also noteworthy that amino acid homology searches indicated the presence of Svf1 homologous sequences in plant pathogenic fungi40 and other fungi including human pathogenic fungi (Supplementary Fig. S11).
Normal conidiophore morphogenesis requires functional interactions between transcription factors such as BrlA, AbaA, and StuA1,27. The ΔsvfA strain formed structurally abnormal conidiophores and thus produced lower amounts of conidiospores due to the reduction in the transcript levels of brlA, abaA, and vosA (Fig. 4F), indicating that SvfA regulates the induction of genes for transcription factors that are critical for asexual development. A rather complicated mechanism is involved in regulation of sexual development26, with esdC and steA genes being expressed at early sexual stage, while vosA and mutA are expressed at later sexual stages. The svfA-deletion strain produced few Hülle cells and small cleistothecia with no ascospores due to the decreased transcription of esdC, steA, vosA, and mutA, indicating that SvfA is a novel regulator, essential for the completion of sexual development in A. nidulans. When we analyzed the expression of genes for upstream sexual regulators VeA and NsdD, expression of both veA and nsdD genes was highly increased by the svfA-deletion (Fig. 5G,H), suggesting that SvfA is required for negative feedback-regulation of veA and nsdD transcription to down-regulate sexual development temporally.
In consistency with the 2-DE-identification of Vdps, which revealed two protein spots of SvfA (Table 1), production of two forms of SvfA protein was revealed by Western blot analysis; production of the larger form is VeA-dependent and sexual-specific, but that of smaller one is VeA-independent and sexual-non-specific (Fig. 7A). Therefore, the SvfA-YFP fusion protein localized in the cytoplasm (Fig. 6) may be the smaller protein. In addition, overexpression of svfA in the ΔveA strain did not alleviate the sexual defects (i.e. failed to produce cleistothecia) (Fig. 7B) and overexpression of svfA in the WT showed defects similar to those seen in the ΔsvfA strain, which produced small cleistothecia (Fig. 7C). These results indicate the following: post-translational modifications can occur to activate SvfA during sexual development, VeA is involved in this modification of SvfA, SvfA functions downstream to VeA together with other VeA-regulated proteins, and normal expression level of the svfA gene is crucial for progression and completion of sexual development in A. nidulans.
SvfA is involved in ST biosynthesis and development, both of which are known to be functionally interconnected4. CpcB, a Gβ-like protein, governs diverse cellular events such as germination, growth, development, and ST synthesis32. Similar to the ΔsvfA strain, the cpcB deletion strain produced small and fragile cleistothecia with no ascospores and showed upregulation of veA and nsdD expression during sexual development32, suggesting a possible interaction between SvfA and CpcB. When levels of cpcB gene expression in the ΔsvfA strain and svfA gene expression in the ΔcpcB strain were studied during sexual development, cpcB expression was not affected by the lack of SvfA and in turn svfA expression was not affected in ΔcpcB mutants (data not shown).
StuA, an APSES domain transcription factor, affects spatial organization of the conidiophores and is also required for the formation of Hülle cells and cleistothecia during early sexual development1,33. The stuA mutant has greatly shortened conidiophores lacking normal metulae and phialides34,35, which are quite similar to those from the ΔsvfA strain. Interestingly, sequence analysis of the 5′-upstream region of svfA revealed 5′-(A/T)CGCG(T/A)N(A/C)-3′ for putative StuA Response Element (StRE)36 at positions −917, −744, −703, −701, −100, relative to the ATG codon. Although further experiments are required to reveal the relationship between StuA and SvfA, these results hint the possibility of SvfA being necessary for the temporal regulation of both asexual and sexual development through functional interaction with StuA (Fig. 7D).
Although mechanism for the multiple functions of the oxidative stress protein SvfA in the developmental processes of A. nidulans remains unclear, several data support that oxidative stress or an imbalanced intracellular redox environment affects development. The cellular oxidation state is one of the physiological changes during early sexual development. For example, NoxA, a NADPH oxidase in A. nidulans, is involved in the production of ROS and cleistothecial development at the early stage37. Transcription of noxA gene is suppressed by SakA MAP kinase37, and deletion of SakA shows an increased number of prematurely developed cleistothecia38. A trxA deletion strain fails to produce cleistothecia under standard conditions. However, low GSH levels leads to the development of cleistothecia, whereas high GSH levels results in the formation of asexual conidiophores39. Activation of the expression of cpeA, a catalase-peroxidase gene, by StuA is also required for the formation of Hülle cells and cleistothecia during early sexual development1,33. It is also noted that Svf1 of S. cerevisiae is involved in cell survival by affecting the sphingolipid metabolism40 and is a substrate of serine/threonine protein kinase CK2, which is essential for life in all eukaryotes by regulating the cell cycle, tumorigenesis, and apoptosis41. Interestingly, exogenous expression of the human anti-apoptotic gene Bcl-xL, which regulates apoptosis and the cell cycle42, could functionally complement the defect of Svf1 in S. cerevisiae24. Mammalian Bcl-xL regulates the intrinsic pathway of apoptotic cell death activated in β-cells under prolonged oxidative and endoplasmic reticulum stress43.
Taken together, we can propose a model for the involvement of SvfA in asexual and sexual development in A. nidulans (Fig. 7D). Further studies should be performed to provide insight into the molecular mechanisms of SvfA-mediated developmental processes; however, our data presented in the present study indicate that the oxidative stress protein SvfA is a novel central regulator of growth, differentiation, and secondary metabolism in A. nidulans.
Methods
Proteome analysis
A. nidulans strains FGSC A4 (veA+) and KVE9 (ΔveA::argB) were used for proteome analysis. Fungal techniques for culture condition, observation, transformation, genetic analyses, and phenotypic analyses were performed according to the previous report2. For protein extraction, the A. nidulans cultures were harvested by filtration through a Miracloth and ground to powder using liquid nitrogen. The grinded samples were homogenized directly by motor-driven homogenizer (PowerGen125, Fisher Scientific) in sample lysis buffer with 7 M urea, 2 M thiourea containing 4% (w/v) CHAPS, 1% (w/v) DTT and 2% (v/v) Pharmalyte (pH 3.5–10, Amersham Biosciences), and 1 mM benzamidine44. Proteins were extracted for 1 h at room temperature with vortexing. After centrifugation at 15,000 × g for 1 h at 15 °C, the supernatant was used for further experiment.
Two-dimensional gel electrophoresis (2-DE) analysis was carried out essentially as described previously44,45. In brief, 200 μg of samples was loaded to rehydrated IPG strips with a nonlinear pH gradient from 4 to 10. Isoelectric focusing (IEF) was performed at 20 °C. The second dimensional SDS-PAGE (20 × 24 cm, 10–16%) was performed using Höefer DALT 2D system (Amersham Biosciences). 2D gels were silver stained as described by Oakley et al.46 but the fixing and sensitization step with glutaraldehyde was omitted. Images were analyzed by the PDQuest (version 7.0, BioRad) software45. Protein spots were selected for the significant expression variation deviated over two-fold in its expression level compared with the control or normal sample. Protein spots were enzymatically digested in-gel in a manner similar to that previously described by Shevchenko et al.47 using modified porcine trypsin (Promega) and analyzed using an Ettan MALDI-TOF mass spectrometer (Amersham Biosciences). The search program ProFound, developed by The Rockefeller University (https://129.85.19.192/profound_bin/ WebProFound.exe), was used for protein identification by peptide mass fingerprinting. Spectra were calibrated with trypsin auto-digestion ion peaks m/z (842.510, 2211.1046) as internal standards48.
Generation of recombinant strains
The A. nidulans strains employed in the present study are listed in Supplementary Table S1. To construct the disruption cassette, amplified argB gene product was inserted between 5′-and 3′ flanking regions of each svfA, vpdC, vpdF, and vdpJ genes as a selective marker using double-joint PCR49. The information of primers for fusion products are listed in Supplementary Table S2. Disruption cassettes were amplified with each set of nested primers and introduced into the TJ1–1 strain. The correct recombination in genomic DNA was confirmed by PCR and Southern blotting. To complement ΔsvfA, the svfA gene region, including its predicted promoter, was amplified and cloned into pHS13, which contains 3/4 of the pyroA gene, a FLAG tag, and the trpC terminator11. For construction of the SvfA-YFP strain, the svfA gene region, including its predicted promoter, was amplified and cloned into pHS-YFP50. To generate svfA-overexpressing strains, the svfA ORF was cloned into pHS11 containing the niiA promoter. The resulting plasmid was introduced into the recipient strain.
Media and culture conditions
Strains were maintained in Aspergillus minimal medium with glucose (GMM)32. The minimal medium supplemented with 0.15% yeast extract and 0.15% casamino acid (YCMM) was used as a complete medium for vegetative mycelial ball production. To observe germination of conidia, conidia were inoculated in liquid GMM and MM (without glucose) and observed every 1 h after incubation at 37 °C using a microscope. For phenotypic analysis during development, vegetative mycelial balls incubated in liquid YCMM for 16–18 h were transferred to GMM solid or MMCA (MM with 0.15% casamino acid) solid media to induce asexual or sexual development, respectively. For sexual development, the media were sealed with parafilm for 24 h and incubated further without sealing under dark conditions. To control the expression by niiA promoter, 0.2% ammonium tartrate or 0.6% sodium nitrate (as a nitrogen source) was added as non-inducing medium or inducing medium, respectively.
Sensitivity test to oxidative and cold stresses
Spotting susceptibility assays was performed as described previously51,52 but modified to some extent. Conidia were resuspended to 2 × 106 cells per ml in distilled water and prepared at 10-fold dilutions. For each dilution, 5 µl was spotted onto a YCMM agar plate (control) or plates containing 50 μM menadione and 4 mM H2O2 for oxidative stress, and the plates were incubated for 2 days at 37 °C. To test the response to cold stress, conidia were spotted on YCMM and incubated for 6 days at 20 °C.
RNA preparation, cDNA synthesis, and quantitative real-time PCR
Cells at each of the developmental stages were ground using liquid nitrogen with a pestle and mortar52. Total RNA was extracted using Trizol according to the manufacturer’s protocols (Invitrogen). cDNA was synthesized using 4 μg extracted RNA, hexamer primer, and M-MLV reverse transcriptase (Enzynomics) as described in the manufacturer’s instructions. RT-qPCR was performed using a Bio-Rad CFX96 Real-Time PCR System (Bio-Rad) and a TOPrealTM qPCR 2X PreMIX Kit (Enzynomics). Transcript levels of target genes were normalized against those of 18 S rRNA using 2−ΔCt method described previously53. The information of primers for RT-qPCR are listed in Supplementary Table S2.
Western blot analysis
Protein extraction and western blot analysis were performed as previously described2. The A. nidulans cultures were ground using liquid nitrogen, and cells were resuspended in protein extraction buffer (50 mM Tris-HCl, pH 8, 150 mM NaCl, 1 mM EDTA, and 1% NP-40) with 2 mM phenylmethylsulfonyl fluoride (PMSF), 10 mM sodium fluoride, and 1 mM sodium vanadate. The supernatant was obtained after centrifugation at 15,000 × g at 4 °C for 30 min. Total protein samples were electrophoresed on 8% SDS-PAGE and subsequently electroblotted onto Hybond-P polyvinylidene difluoride (PVDF) membranes (GE Healthcare). The membrane was blocked with 5% skimmed milk, and protein detection was carried out using anti-FLAG (Sigma-Aldrich) and goat anti-mouse IgG-HRP (Santa Cruz Biotechnology) secondary antibody following the manufacturers’ protocols (ELPISBIO). SDS-PAGE and silver staining kit (ELPISBIO) were used for silver staining.
Microscopy
For microcopy, an Olympus System microscope Model BX51 (Olympus) equipped with UPlanSApo 60X and UPlanFL 100X objective lenses (Olympus) and stereomicroscope Model SMZ800 (Nikon) were used. Images were captured with a DP71 digital camera (Olympus) and processed using the DP manager imaging software (Olympus). For microscopic observation of the fungal hyphae, each strain was coverslip-cultured on a block of appropriate agar medium or incubated in liquid GMM medium. The coverslips were stained with 1 mg/ml Hoechst 33342 (Sigma-Aldrich) for labeling DNA2. DAPI (high brightness) filter cubes (excitation filter: center wavelength 377 nm, emission filter: center wavelength 447 nm, Olympus) and FITC filter cubes (excitation filter: center wavelength 483 nm, emission filter: center wavelength 535 nm, Olympus) were used to observe the fluorescence of Hoechst and YFP, respectively50.
References
Wu, J. & Miller, B. L. Aspergillus asexual reproduction and sexual reproduction are differentially affected by transcriptional and translational mechanisms regulating stunted gene expression. Mol. Cell. Biol. 17, 6191–6201 (1997).
Kang, E. H., Kim, J., Oh, H. W. & Park, H. M. LAMMER kinase LkhA plays multiple roles in the vegetative growth and asexual and sexual development of Aspergillus nidulans. PLoS One 8, e58762, https://doi.org/10.1093/mmy/myz049 (2013).
Bok, J. W. & Keller, N. P. LaeA, a regulator of secondary metabolism in Aspergillus spp. Eukaryot. Cell 3, 527–535, https://doi.org/10.1128/EC.3.2.527-535.2004 (2004).
Bayram, Ö. & Braus, G. H. Coordination of secondary metabolism and development in fungi: The velvet family of regulatory proteins. FEMS Microbiol. Rev. 36, 1–24, https://doi.org/10.1111/j.1574-6976.2011.00285.x (2012).
Adams, T. H., Wieser, J. K. & Yu, J. H. Asexual sporulation in Aspergillus nidulans. Microbiol. Mol. Biol. Rev 62, 35–54 (1998).
Han, K. et al. Environmental factors affecting development of Aspergillus nidulans. J. Microbiol. 41, 34–40 (2003).
Han, K. H. Molecular genetics of Emericella nidulans sexual development. Mycobiology 37, 171–182, https://doi.org/10.4489/MYCO.2009.37.3.171 (2009).
Bayram, Ö. et al. The Aspergillus nidulans MAPK module AnSte11-Ste50-Ste7-Fus3 controls development and secondary metabolism. PLoS Genet. 8, e1002816, https://doi.org/10.1371/journal.pgen.1002816 (2012).
Ni, M. & Yu, J. H. A novel regulator couples sporogenesis and trehalose biogenesis in Aspergillus nidulans. PLoS One 2, e970, https://doi.org/10.1371/journal.pone.0000970 (2007).
Dhingra, S., Andes, D. & Calvoa, A. M. VeA regulates conidiation, gliotoxin production, and protease activity in the opportunistic human pathogen Aspergillus fumigatus. Eukaryot. Cell 11, 1531–1543, https://doi.org/10.1128/EC.00222-12 (2012).
Park, H. S., Ni, M., Jeong, K. C., Kim, Y. H. & Yu, J. H. The role, interaction and regulation of the velvet regulator VelB in Aspergillus nidulans. PLoS One 7, e45935, https://doi.org/10.1371/journal.pone.0045935 (2012).
Park, H. S., Nam, T. Y., Han, K. H., Kim, S. C. & Yu, J. H. VelC positively controls sexual development in Aspergillus nidulans. PLoS One 9, e89883, https://doi.org/10.1371/journal.pone.0089883 (2014).
Stinnett, S. M., Espeso, E. A., Cobeño, L., Araújo-Bazán, L. & Calvo, A. M. Aspergillus nidulans VeA subcellular localization is dependent on the importin α carrier and on light. Mol. Microbiol. 63, 242–255, https://doi.org/10.1111/j.1365-2958.2006.05506.x (2007).
Bayram, Ö. et al. VelB/VeA/LaeA complex coordinates light signal with fungal development and secondary metabolism. Science. 320, 1504–1506, https://doi.org/10.1126/science.1155888 (2008).
Sarikaya Bayram, Ö. et al. LaeA control of velvet family regulatory proteins for light-dependent development and fungal cell-type specificity. PLoS Genet. 6, 1–17, https://doi.org/10.1371/journal.pgen.1001226 (2010).
Palmer, J. M. et al. Secondary metabolism and development is mediated by LlmF control of VeA subcellular localization in Aspergillus nidulans. PLoS Genet. 9, e1003193, https://doi.org/10.1371/journal.pgen.1003193 (2013).
Han, K. H., Park, J. S., Chae, K. S. & Han, D. M. Simple identification of veA1 mutation in Aspergillus nidulans. J. Microbiol. 48, 885–887, https://doi.org/10.1007/s12275-010-0506-y (2010).
Kim, H. S. et al. The veA gene activates sexual development in Aspergillus nidulans. Fungal Genet. Biol. 37, 72–80 (2002).
Kato, N., Brooks, W. & Calvo, A. M. The expression of sterigmatocystin and penicillin genes in Aspergillus nidulans is controlled by veA, a gene required for sexual development. Eukaryot. Cell 2, 1178–86, https://doi.org/10.1128/ec.2.6.1178-1186.2003 (2003).
Sarikaya-Bayram, Ö. et al. Membrane-bound methyltransferase complex VapA-VipC-VapB guides epigenetic control of fungal development. Dev. Cell 29, 406–420, https://doi.org/10.1016/j.devcel.2014.03.020 (2014).
Kim, H. R., Chae, K. S., Han, K. H. & Han, D. M. The nsdC gene encoding a putative C2H2 -type transcription factor is a key activator of sexual development in Aspergillus nidulans. Genetics 182, 771–783, https://doi.org/10.1534/genetics.109.101667 (2009).
Kim, Y., Nandakumar, M. P. & Marten, M. R. Proteome map of Aspergillus nidulans during osmoadaptation. Fungal Genet. Biol. 44, 886–895, https://doi.org/10.1016/j.fgb.2006.12.001 (2007).
Wartenberg, D. et al. Proteome analysis of the farnesol-induced stress response in Aspergillus nidulans-The role of a putative dehydrin. J. Proteomics 75, 4038–4049, https://doi.org/10.1016/j.jprot.2012.05.023 (2012).
Brace, J. L., VanderWeele, D. J. & Rudin, C. M. Svf1 inhibits reactive oxygen species generation and promotes survival under conditions of oxidative stress in Saccharomyces cerevisiae. Yeast 22, 641–652, https://doi.org/10.1002/yea.1235 (2005).
Boylan, M. T., Mirabito, P. M., Willett, C. E., Zimmerman, C. R. & Timberlake, W. E. Isolation and physical characterization of three essential conidiation genes from Aspergillus nidulans. Mol. Cell. Biol. 7, 3113–8, https://doi.org/10.1128/mcb.7.9.3113 (1987).
Dyer, P. S. & O’Gorman, C. M. Sexual development and cryptic sexuality in fungi: insights from Aspergillus species. FEMS Microbiol. Rev. 36, 165–192, https://doi.org/10.1111/j.1574-6976.2011.00308.x (2012).
Han, K. H. et al. The Aspergillus nidulans esdC (early sexual development) gene is necessary for sexual development and is controlled by veA and a heterotrimeric G protein. Fungal Genet. Biol. 45, 310–318, https://doi.org/10.1016/j.fgb.2007.09.008 (2008).
Vallim, M. A., Miller, K. Y. & Miller, B. L. Aspergillus SteA (sterile12-like) is a homeodomain-C2/H2-Zn+2 finger transcription factor required for sexual reproduction. Mol. Microbiol. 36, 290–301, https://doi.org/10.1046/j.1365-2958.2000.01874.x (2000).
Wei, H., Scherer, M., Singh, A., Liese, R. & Fischer, R. Aspergillus nidulans α-1,3 glucanase (mutanase), mutA, is expressed during sexual development and mobilizes mutan. Fungal Genet. Biol. 34, 217–227, https://doi.org/10.1006/fgbi.2001.1303 (2001).
Han, K. H. et al. The nsdD gene encodes a putative GATA-type transcription factor necessary for sexual development of Aspergillus nidulans. Mol. Microbiol. 41, 299–309, https://doi.org/10.1046/j.1365-2958.2001.02472.x (2001).
Yu, Y. et al. Survival factor 1 contributes to the oxidative stress response and is required for full virulence of Sclerotinia sclerotiorum. Mol. Plant Pathol. 20, 895–906, https://doi.org/10.1111/mpp.12801 (2019).
Kong, Q. et al. Gβ-Like CpcB plays a crucial role for growth and development of Aspergillus nidulans and Aspergillus fumigatus. PLoS One 8, e70355, https://doi.org/10.1371/journal.pone.0070355 (2013).
Scherer, M., Wei, H., Liese, R. & Fischer, R. Aspergillus nidulans catalase-peroxidase gene (cpeA) is transcriptionally induced during sexual development through the transcription factor StuA. Eukaryot. Cell 1, 725–735, https://doi.org/10.1128/ec.1.5.725-735.2002 (2002).
Miller, K. Y., Wu, J. G. & Miller, B. L. StuA is required for cell pattern-formation in Aspergillus. Genes Dev 6, 1770–1782, https://doi.org/10.1101/gad.6.9.1770 (1992).
Miller, K. Y., Toennis, T. M., Adams, T. H. & Miller, B. L. Isolation and transcriptional characterization of a morphological modifier: the Aspergillus nidulans stunted (stuA) gene. Mol. Gen. Gene. 227, 285–292, https://doi.org/10.1007/bf00259682 (1991).
Park, B. C. et al. Transcriptional regulation of fksA, a β-1,3-glucan synthase gene, by the APSES protein StuA during Aspergillus nidulans development. J. Microbiol. 52, 940–7, https://doi.org/10.1007/s12275-014-4517-y (2014).
Lara-Ortíz, T., Riveros-Rosas, H. & Aguirre, J. Reactive oxygen species generated by microbial NADPH oxidase NoxA regulate sexual development in Aspergillus nidulans. Mol. Microbiol 50, 1241–1255, https://doi.org/10.1046/j.1365-2958.2003.03800.x (2003).
Kawasaki, L., Sánchez, O., Shiozaki, K. & Aguirre, J. SakA MAP kinase is involved in stress signal transduction, sexual development and spore viability in Aspergillus nidulans. Mol. Microbiol. 45, 1153–1163, https://doi.org/10.1046/j.1365-2958.2002.03087.x (2002).
Sato, I., Shimizu, M., Hoshino, T. & Takaya, N. The glutathione system of Aspergillus nidulans involves a fungus-specific glutathione S-transferase. J. Biol. Chem. 284, 8042–8053, https://doi.org/10.1074/jbc.M807771200 (2009).
Brace, J. L., Lester, R. L., Dickson, R. C. & Rudin, C. M. SVF1 regulates cell survival by affecting sphingolipid metabolism in Saccharomyces cerevisiae. Genetics 175, 65–76, https://doi.org/10.1534/genetics.106.064527 (2007).
Masłyk, M. et al. Yeast surviving factor Svf1 as a new interacting partner, regulator and in vitro substrate of protein kinase CK2. Mol. Cell. Biochem. 312, 61–69, https://doi.org/10.1007/s11010-008-9721-9 (2008).
Janumyan, Y. M. et al. Bcl-x /Bcl-2 coordinately regulates apoptosis, cell cycle arrest and cell cycle entry. EMBO J. 22, 5459–5470, https://doi.org/10.1093/emboj/cdg533 (2003).
Aharoni-Simon, M. et al. Bcl-2 regulates reactive oxygen species signaling and a redox-sensitive mitochondrial proton leak in mouse pancreatic β-cells. Endocrinology 157, 2270–2281, https://doi.org/10.1210/en.2015-1964 (2016).
Uhm, Y. K. et al. Effects of Machilus thunbergii Sieb et Zucc on UV-induced photoaging in hairless mice. Phyther. Res 24, 1339–1346, https://doi.org/10.1002/ptr.3117 (2010).
Deng, W. W., Sasamoto, H. & Ashihara, H. Effect of caffeine on the expression pattern of water-soluble proteins in rice (Oryza sativa) seedlings. Nat. Prod. Commun 10, 733–6, https://doi.org/10.1177/1934578X1501000509 (2015).
Oakley, B. R., Kirsch, D. R. & Morris, N. R. A simplified ultrasensitive silver stain for detecting proteins in polyacrylamide gels. Anal. Biochem. 105, 361–363, https://doi.org/10.1016/0003-2697(80)90470-4 (1980).
Shevchenko, A., Wilm, M., Vorm, O. & Mann, M. Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal. Chem. 68, 850–8, https://doi.org/10.1021/ac950914h (1996).
Lee, S. Y. & Lee, K. O. Proteomic analysis of RNA interference induced knockdown plant. Methods Mol. Biol 744, 211–24, https://doi.org/10.1007/978-1-61779-123-9_15 (2011).
Yu, J. H. et al. Double-joint PCR: a PCR-based molecular tool for gene manipulations in filamentous fungi. Fungal Genet. Biol. 41, 973–981, https://doi.org/10.1016/j.fgb.2004.08.001 (2004).
Kim, Y. J., Yeong Man, Y. & Maeng, P. J. Differential control of asexual development and sterigmatocystin biosynthesis by a novel regulator in Aspergillus nidulans. Sci. Rep 7, 46340, https://doi.org/10.1038/srep46340 (2017).
Rocha, M. C. et al. Aspergillus fumigatus MADS-Box transcription factor rlmA is required for regulation of the cell wall integrity and virulence. G3 (Bethesda) 6, 2983–3002, https://doi.org/10.1534/g3.116.031112 (2016).
Rocha, M. C. et al. The Aspergillus fumigatus pkcA mutant is defective in the activation of the cell wall integrity pathway but is dispensable for virulence in a neutropenic mouse infection model. PLoS One 10, e0135195, https://doi.org/10.1371/journal.pone.0135195 (2015).
Park, D. S., Yu, Y. M., Kim, Y. J. & Maeng, P. J. Negative regulation of the vacuole-mediated resistance to K+ stress by a novel C2H2 zinc finger transcription factor encoded by aslA in Aspergillus nidulans. J. Microbiol. 53, 100–110, https://doi.org/10.1007/s12275-015-4701-8 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Priebe, S., Kreisel, C., Horn, F., Guthke, R. & Linde, J. FungiFun2: a comprehensive online resource for systematic analysis of gene lists from fungal species. Bioinformatics 31, 445–446, https://doi.org/10.1093/bioinformatics/btu627 (2015).
Acknowledgements
This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) by the Ministry of Education (Grant No. 2016R1D1A1B03936038). We would like to thank Editage (www.editage.co.kr) for English language editing.
Author information
Authors and Affiliations
Contributions
J.Y.L., E.H.K., Y.H.P., and J.H.K. performed the experiments; J.Y.L., E.H.K., and H.M.P. designed the experiments, analyzed the data, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lim, JY., Kang, EH., Park, YH. et al. Survival factor SvfA plays multiple roles in differentiation and is essential for completion of sexual development in Aspergillus nidulans. Sci Rep 10, 5586 (2020). https://doi.org/10.1038/s41598-020-62455-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-62455-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.