Abstract
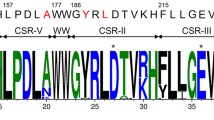
This study reported simultaneously improved thermostability and hydrolytic pattern of α-amylase from Bacillus subtilis CN7 by rationally engineering the mostly conserved central beta strands in TIM barrel fold. Nine single point mutations and a double mutation were introduced at the 2nd site of the β7 strand and 3rd site of the β5 strand to rationalize the weak interactions in the beta strands of the TIM barrel of α-amylase. All the five active mutants changed the compositions and percentages of maltooligosaccharides in final hydrolytic products compared to the product spectrum of the wild-type. A mutant Y204V produced only maltose, maltotriose, and maltopentaose without any glucose and maltotetraose, indicating a conversion from typical endo-amylase to novel maltooligosaccharide-producing amylase. A mutant V260I enhanced the thermal stability by 7.1 °C. To our best knowledge, this is the first report on the simultaneous improvement of thermostability and hydrolytic pattern of α-amylase by engineering central beta strands of TIM barrel and the novel “beta strands” strategy proposed here may be useful for the protein engineering of other TIM barrel proteins.


Similar content being viewed by others
References
Farber, G. K., & Petsko, G. A. (1990). The evolution of [alpha]/[beta] barrel enzymes. Trends in Biochemical Sciences, 15(6), 228–234.
Brändén, R. I. (1991). The TIM barrel—The most frequently occurring folding motif in proteis. Current Opinion in Structural Biology, 1, 978–983.
Petsko, G. A. (2000). Enzyme evolution. Design by necessity. Nature, 403(6770), 606–607.
Wierenga, R. K. (2001). The TIM-barrel fold: A versatile framework for efficient enzymes. FEBS Letters, 492(3), 193–198.
Hocker, B., Jurgens, C., Wilmanns, M., & Sterner, R. (2001). Stability, catalytic versatility and evolution of the (beta alpha)(8)-barrel fold. Current Opinion in Biotechnology, 12(4), 376–381.
Sterner, R., & Hocker, B. (2005). Catalytic versatility, stability, and evolution of the (betaalpha)8-barrel enzyme fold. Chemical Reviews, 105(11), 4038–4055.
Jaenicke, R., & Bohm, G. (1998). The stability of proteins in extreme environments. Current Opinion in Structural Biology, 8(6), 738–748.
Vieille, C., & Zeikus, G. J. (2001). Hyperthermophilic enzymes: Sources, uses, and molecular mechanisms for thermostability. Microbiology and Molecular Biology Reviews, 65(1), 1–43.
Lesk, A. M., Branden, C. I., & Chothia, C. (1989). Structural principles of alpha/beta barrel proteins: The packing of the interior of the sheet. Proteins, 5(2), 139–148.
Selvaraj, S., & Gromiha, M. M. (1998). An analysis of the amino acid clustering pattern in (alpha/beta)8 barrel proteins. Journal of Protein Chemistry, 17(5), 407–415.
Selvaraj, S., & Gromiha, M. M. (1998). Importance of long-range interactions in (alpha/beta)8 barrel fold. Journal of Protein Chemistry, 17(7), 691–697.
Chakkaravarthi, S., Babu, M. M., Gromiha, M. M., Jayaraman, G., & Sethumadhavan, R. (2006). Exploring the environmental preference of weak interactions in (alpha/beta)8 barrel proteins. Proteins, 65(1), 75–86.
Vijayabaskar, M. S., & Vishveshwara, S. (2012). Insights into the fold organization of TIM barrel from interaction energy based structure networks. PLoS Computational Biology, 8(5), e1002505.
Gromiha, M. M., Pujadas, G., Magyar, C., Selvaraj, S., & Simon, I. (2004). Locating the stabilizing residues in (alpha/beta)8 barrel proteins based on hydrophobicity, long-range interactions, and sequence conservation. Proteins, 55(2), 316–329.
Ponder, J. W., & Richards, F. M. (1987). Tertiary templates for proteins. Use of packing criteria in the enumeration of allowed sequences for different structural classes. Journal of Molecular Biology, 193(4), 775–791.
Silverman, J. A., Balakrishnan, R., & Harbury, P. B. (2001). Reverse engineering the (beta/alpha)8 barrel fold. Proceedings of the National Academy of Sciences of the United States of America, 98(6), 3092–3097.
Chothia, C. (1976). The nature of the accessible and buried surfaces in proteins. Journal of Molecular Biology, 105(1), 1–12.
MacGregor, E. A. (1988). Alpha-amylase structure and activity. Journal of Protein Chemistry, 7(4), 399–415.
Buisson, G., Duee, E., Haser, R., & Payan, F. (1987). Three dimensional structure of porcine pancreatic alpha-amylase at 2.9 A resolution. Role of calcium in structure and activity. The EMBO Journal, 6(13), 3909–3916.
Nielsen, J. E., & Borchert, T. V. (2000). Protein engineering of bacterial alpha-amylases. Biochimica et Biophysica Acta, 1543(2), 253–274.
Bessler, C., Schmitt, J., Maurer, K. H., & Schmid, R. D. (2003). Directed evolution of a bacterial alpha-amylase: Toward enhanced pH-performance and higher specific activity. Protein Science, 12(10), 2141–2149.
Yang, H., Liu, L., Shin, H. D., Chen, R. R., Li, J., Du, G., & Chen, J. (2013). Structure-based engineering of histidine residues in the catalytic domain of alpha-amylase from Bacillus subtilis for improved protein stability and catalytic efficiency under acidic conditions. Journal of Biotechnology, 164(1), 59–66.
Liu, Y., Fan, S., Liu, X., Zhang, Z., Wang, J., Wang, Z., & Lu, F. (2014). A highly active alpha amylase from Bacillus licheniformis: Directed evolution, enzyme characterization and structural analysis. Journal of Microbiology and Biotechnology, 24(7), 898–904.
Wang, C. H., Liu, X. L., Huang, R. B., He, B. F., & Zhao, M. M. (2018). Enhanced acidic adaptation of Bacillus subtilis Ca-independent alpha-amylase by rational engineering of pKa values. Biochemical Engineering Journal, 139, 146–153.
Wang, C., Huang, R., He, B., & Du, Q. (2012). Improving the thermostability of alpha-amylase by combinatorial coevolving-site saturation mutagenesis. BMC Bioinformatics, 13, 263.
Peitsch, M. C. (1995). Protein modeling by E-mail. Nature Biotechnology, 13(7), 658–660.
Berman, H. M., Westbrook, J., Feng, Z., Gilliland, G., Bhat, T. N., Weissig, H., Shindyalov, I. N., & Bourne, P. E. (2000). The protein data bank. Nucleic Acids Research, 28(1), 235–242.
Guex, N., & Peitsch, M. C. (1997). SWISS-MODEL and the Swiss-PdbViewer: An environment for comparative protein modeling. Electrophoresis, 18(15), 2714–2723.
Tiwari, A., & Panigrahi, S. K. (2007). HBAT: A complete package for analysing strong and weak hydrogen bonds in macromolecular crystal structures. In Silico Biology, 7(6), 651–661.
Gallivan, J. P., & Dougherty, D. A. (1999). Cation-pi interactions in structural biology. Proceedings of the National Academy of Sciences of the United States of America, 96(17), 9459–9464.
Sweet, R. M., & Eisenberg, D. (1983). Correlation of sequence hydrophobicities measures similarity in three-dimensional protein structure. Journal of Molecular Biology, 171(4), 479–488.
Sambrook, J., & Russell, D. (2001). Molecular cloning: A laboratory manual (third ed.). New York: Cold Spring Harbor Laboratory Press.
Bradford, M. M. (1976). A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Analytical Biochemistry, 72, 248–254.
Miller, G. L. (1959). Use of dinitrosalicylic acid reagent for determination of reducing sugar. Analytical Chemistry, 31(3), 426–428.
Lineweaver, H., & Burk, D. (1934). The determination of enzyme dissociation constants. Journal of the American Chemical Society, 56(3), 658–666.
Wang, C., Wang, Q., Liao, S., He, B., & Huang, R. (2014). Thermal stability and activity improvements of a Ca-independent alpha-amylase from Bacillus subtilis CN7 by C-terminal truncation and hexahistidine-tag fusion. Biotechnology and Applied Biochemistry, 61(2), 93–100.
van der Maarel, M. J., van der Veen, B., Uitdehaag, J. C., Leemhuis, H., & Dijkhuizen, L. (2002). Properties and applications of starch-converting enzymes of the alpha-amylase family. Journal of Biotechnology, 94(2), 137–155.
Pan, S., Ding, N., Ren, J., Gu, Z., Li, C., Hong, Y., Cheng, L., Holler, T. P., & Li, Z. (2017). Maltooligosaccharide-forming amylase: Characteristics, preparation, and application. Biotechnology Advances, 35(5), 619–632.
Bowie, J. U., Reidhaar-Olson, J. F., Lim, W. A., & Sauer, R. T. (1990). Deciphering the message in protein sequences: Tolerance to amino acid substitutions. Science, 247(4948), 1306–1310.
Skolnick, J., Kolinski, A., & Yaris, R. (1989). Monte Carlo studies on equilibrium globular protein folding. II. Beta-barrel globular protein models. Biopolymers, 28(6), 1059–1095.
Yue, K., & Dill, K. A. (1992). Inverse protein folding problem: Designing polymer sequences. Proceedings of the National Academy of Sciences of the United States of America, 89(9), 4163–4167.
Hocker, B., Beismann-Driemeyer, S., Hettwer, S., Lustig, A., & Sterner, R. (2001). Dissection of a (betaalpha)8-barrel enzyme into two folded halves. Nature Structural Biology, 8(1), 32–36.
Declerck, N., Machius, M., Joyet, P., Wiegand, G., Huber, R., & Gaillardin, C. (2002). Engineering the thermostability of Bacillus licheniformis a-amylase. Biologia, Bratislava, 57(Suppl. 11), 9.
Suzuki, Y., Ito, N., Yuuki, T., Yamagata, H., & Udaka, S. (1989). Amino acid residues stabilizing a Bacillus alpha-amylase against irreversible thermoinactivation. The Journal of Biological Chemistry, 264(32), 18933–18938.
Igarashi, K., Hatada, Y., Ikawa, K., Araki, H., Ozawa, T., Kobayashi, T., Ozaki, K., & Ito, S. (1998). Improved thermostability of a Bacillus alpha-amylase by deletion of an arginine-glycine residue is caused by enhanced calcium binding. Biochemical and Biophysical Research Communications, 248(2), 372–377.
Gai, Y., Cheng, J., Zhang, S., Zhu, B., & Zhang, D. (2018). Property improvement of α-amylase from Bacillus stearothermophilus by deletion of amino acid residues arginine 179 and glycine 180. Food Technology and Biotechnology, 56(1), 58–64.
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China [grant number 21868003]; the Chinese National Basic Research Program (“973”) [grant number 2009CB724703]; the Major Program of Natural Science Foundation of Guangxi [grant number 2016GXNSFEA380003]; and the Guangxi BaGui Scholars of Guangxi Zhuang Autonomous Region of China.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
ESM 1
(DOCX 738 kb)
Rights and permissions
About this article
Cite this article
Wang, CH., Lu, LH., Huang, C. et al. Simultaneously Improved Thermostability and Hydrolytic Pattern of Alpha-Amylase by Engineering Central Beta Strands of TIM Barrel. Appl Biochem Biotechnol 192, 57–70 (2020). https://doi.org/10.1007/s12010-020-03308-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-020-03308-8