Abstract
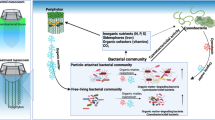
Phytoplankton growth and degradation are associated with bacterial communities in phycosphere. We compared the bacterial communities in two algal blooms (Microcystis aeruginosa and Euglena sanguinea). In particular, large particle-associated, small particle-associated, and free-living bacteria were collected in succession using membranes with three different pore sizes (10 µm, 1 µm, and 0.22 µm) in this study. The results showed that Proteobacteria and Bacteroidetes were the dominant phyla in both blooms. Significant differences in the structure, function, and assembly were observed between large particle-associated and free-living bacterial community in both blooms. Specifically, relative abundance of Roseomonas was higher in the large-size (> 10 µm) aggregates, while Candidate_division_TM7 and Candidate_division_SR1 were more abundant in the free-living bacterial community (0.22–1 µm). The large particle-associated bacterial community was mainly driven by heterogeneous selection and dispersal limitation, whereas the small particle-associated and free-living bacterial communities were mostly determined by dispersal limitation. Besides, some specific bacterial subgroups were found between M. aeruginosa and E. sanguinea blooms. Chlorobi, Chloroflexi, and Fusobacteria were abundant in the M. aeruginosa blooms, whereas Deinococcus–Thermus was abundant in the E. sanguinea blooms. Those specific bacteria provide competitive advantages for the dominated algae. Altogether, these findings indicate that the phycosphere of bloom-forming algae provides different ecological niches where different functional bacterial groups can adapt.




Similar content being viewed by others

References
APHA (2005) Standard methods for the examination of water and wastewater. American Public Health Association, Washington, DC, USA
Baik KS, Park SC, Choe HN, Kim SN, Moon J-H, Seong CN (2012) Roseomonas riguiloci sp. nov., isolated from wetland freshwater. Int J Syst Evol Micr 62:3024–3029. https://doi.org/10.1099/ijs.0.036186-0
Bittencourt-Oliveira MD, de Oliveira MC, Bolch CJS (2001) Genetic variability of Brazilian strains of the Microcystis aeruginosa complex (Cyanobacteria/Cyanophyceae) using the phycocyanin intergenic spacer and flanking regions (cpcBA). J Phycol 37:810–818. https://doi.org/10.1046/j.1529-8817.2001.00102.x
Cai HY, Jiang HL, Krumholz LR, Yang Z (2014) Bacterial community composition of size-fractioned aggregates within the phycosphere of cyanobacterial blooms in a eutrophic freshwater lake. PLoS ONE. https://doi.org/10.1371/journal.pone.0102879
Cao X, Zhou Y, Wang Z, Song C (2016) The contribution of attached bacteria to Microcystis bloom: evidence from field investigation and microcosm experiment. Geomicrobiol J 33:1–11. https://doi.org/10.1080/01490451.2015.1074319
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Tumbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Cho DH, Ramanan R, Heo J, Lee J, Kim BH, Oh HM, Kim HS (2015) Enhancing microalgal biomass productivity by engineering a microalgal-bacterial community. Bioresour Technol 175:578–585. https://doi.org/10.1016/j.biortech.2014.10.159
Cloern JE (1996) Phytoplankton bloom dynamics in coastal ecosystems: a review with some general lessons from sustained investigation of San Francisco Bay, California. Rev Geophys 34:127–168. https://doi.org/10.1029/96RG00986
Dang H, Lovell CR (2015) Microbial surface colonization and biofilm development in marine environments. Microbiol Mol Biol Rev 80:91–138. https://doi.org/10.1128/MMBR.00037-15
Datta MS, Sliwerska E, Gore J, Polz MF, Cordero OX (2016) Microbial interactions lead to rapid micro-scale successions on model marine particles. Nat Commun 7:11965. https://doi.org/10.1038/ncomms11965
Dixon P (2003) VEGAN, a package of R functions for community ecology. J Veg Sci 14:927–930. https://doi.org/10.1111/j.1654-1103.2003.tb02228.x
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Enke TN, Leventhal GE, Metzger M, Saavedra JT, Cordero OX (2018) Microscale ecology regulates particulate organic matter turnover in model marine microbial communities. Nat Commun 9:2743. https://doi.org/10.1038/s41467-018-05159-8
Grossart HP, Levold F, Allgaier M, Simon M, Brinkhoff T (2005) Marine diatom species harbour distinct bacterial communities. Environ Microbiol 7:860–873. https://doi.org/10.1111/j.1462-2920.2005.00759.x
Hillebrand H, Durselen CD, Kirschtel D, Pollingher U, Zohary T (1999) Biovolume calculation for pelagic and benthic microalgae. J Phycol 35:403–424. https://doi.org/10.1046/j.1529-8817.1999.3520403.x
Hunken M, Harder J, Kirst GO (2008) Epiphytic bacteria on the Antarctic ice diatom Amphiprora kufferathii Manguin cleave hydrogen peroxide produced during algal photosynthesis. Plant Biol 10:519–526. https://doi.org/10.1111/j.1438-8677.2008.00040.x
Huws SA, Edwards JE, Kim EJ, Scollan ND (2007) Specificity and sensitivity of eubacterial primers utilized for molecular profiling of bacteria within complex microbial ecosystems. J Microbiol Meth 70:565–569. https://doi.org/10.1016/j.mimet.2007.06.013
Iyengar MOP (1962) Euglena studies from Madras. Arch Mikrobiol 42:322–332. https://doi.org/10.1007/Bf00422049
Kimbrel JA, Samo TJ, Ward C, Nilson D, Thelen MP, Siccardi A, Zimba P, Lane TW, Mayali X (2019) Host selection and stochastic effects influence bacterial community assembly on the microalgal phycosphere. Algal Res 40:101489. https://doi.org/10.1016/j.algal.2019.101489
Kosten S, Huszar VLM, Becares E, Costa LS, van Donk E, Hansson LA, Jeppesenk E, Kruk C, Lacerot G, Mazzeo N, De Meester L, Moss B, Lurling M, Noges T, Romo S, Scheffer M (2012) Warmer climates boost cyanobacterial dominance in shallow lakes. Glob Change Biol 18:118–126. https://doi.org/10.1111/j.1365-2486.2011.02488.x
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Clemente JC, Burkepile DE, Vega Thurber RL, Knight R, Beiko RG, Huttenhower C (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821. https://doi.org/10.1038/nbt.2676
Liu H, Guo X, Gooneratne R, Lai R, Zeng C, Zhan F, Wang W (2016) The gut microbiome and degradation enzyme activity of wild freshwater fishes influenced by their trophic levels. Sci Rep 6:24340. https://doi.org/10.1038/srep24340
Liu M, Liu L, Chen H, Yu Z, Yang JR, Xue Y, Huang B, Yang J (2019) Community dynamics of free-living and particle-attached bacteria following a reservoir Microcystis bloom. Sci Total Environ 660:501–511. https://doi.org/10.1016/j.scitotenv.2018.12.414
Louati I, Pascault N, Debroas D, Bernard C, Humbert JF, Leloup J (2015) Structural diversity of bacterial communities associated with bloom-forming freshwater cyanobacteria differs according to the cyanobacterial genus. PLoS ONE 10:e0146866. https://doi.org/10.1371/journal.pone.0140614
Main CR, Salvitti LR, Whereat EB, Coyne KJ (2015) Community level and species specific associations between phytoplankton and particle associated vibrio species in Delaware’s inland bays. Appl Environ Microbiol 81:5703–5713. https://doi.org/10.1128/Aem.00580-15
Mittler R, Vanderauwera S, Suzuki N, Miller G, Tognetti VB, Vandepoele K, Gollery M, Shulaev V, Van Breusegem F (2011) ROS signaling: the new wave? Trends Plant Sci 16:300–309. https://doi.org/10.1016/j.tplants.2011.03.007
Mulkidjanian AY, Koonin EV, Makarova KS, Mekhedov SL, Sorokin A, Wolf YI, Dufresne A, Partensky F, Burd H, Kaznadzey D, Haselkorn R, Galperin MY (2006) The cyanobacterial genome core and the origin of photosynthesis. Proc Natl Acad Sci USA 103:13126–13131. https://doi.org/10.1073/pnas.0605709103
O’Neil JM, Davis TW, Burford MA, Gobler CJ (2012) The rise of harmful cyanobacteria blooms: the potential roles of eutrophication and climate change. Harmful Algae 14:313–334. https://doi.org/10.1016/j.hal.2011.10.027
Paerl HW, Tucker CS (1995) Ecology of blue-green algae in aquaculture ponds. J World Aquacult Soc 26:109–131. https://doi.org/10.1111/j.1749-7345.1995.tb00235.x
Paradis E, Claude J, Strimmer K (2004) APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20:289–290. https://doi.org/10.1093/bioinformatics/btg412
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30:3123–3124. https://doi.org/10.1093/bioinformatics/btu494
Parveen B, Ravet V, Djediat C, Mary I, Quiblier C, Debroas D, Humbert JF (2013) Bacterial communities associated with Microcystis colonies differ from free-living communities living in the same ecosystem. Env Microbiol Rep 5:716–724. https://doi.org/10.1111/1758-2229.12071
Pernthaler J (2017) Competition and niche separation of pelagic bacteria in freshwater habitats. Environ Microbiol 19:2133–2150. https://doi.org/10.1111/1462-2920.13742
R Core Team (2010) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Rodgers JH (2008) Algal toxins in pond aquaculture. Southern Regional Aquaculture Center USA
Salazar G, Cornejo-Castillo FM, Borrull E, Díez-Vives C, Lara E, Vaqué D, Arrieta JM, Duarte CM, Gasol JM, Acinas SG (2015) Particle-association lifestyle is a phylogenetically conserved trait in bathypelagic prokaryotes. Mol Ecol 24:5692–5706. https://doi.org/10.1111/mec.13419
Sapp M, Schwaderer AS, Wiltshire KH, Hoppe H-G, Gerdts G, Wichels A (2007) Species-specific bacterial communities in the phycosphere of microalgae? Microb Ecol 53:683–699. https://doi.org/10.1007/s00248-006-9162-5
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:R60. https://doi.org/10.1186/Gb-2011-12-6-R60
Seymour JR, Amin SA, Raina JB, Stocker R (2017) Zooming in on the phycosphere: the ecological interface for phytoplankton-bacteria relationships. Nat Microbiol 2:17065. https://doi.org/10.1038/nmicrobiol.2017.65
Shen H, Niu Y, Xie P, Tao M, Yang X (2011) Morphological and physiological changes in Microcystis aeruginosa as a result of interactions with heterotrophic bacteria. Freshw Biol 56:1065–1080. https://doi.org/10.1111/j.1365-2427.2010.02551.x
Simon HM, Smith MW, Herfort L (2014) Metagenomic insights into particles and their associated microbiota in a coastal margin ecosystem. Front Microbiol 5:466. https://doi.org/10.3389/fmicb.2014.00466
Smriga S, Fernandez VI, Mitchell JG, Stocker R (2016) Chemotaxis toward phytoplankton drives organic matter partitioning among marine bacteria. Proc Natl Acad Sci USA 113:1576–1581. https://doi.org/10.1073/pnas.1512307113
Stegen JC, Lin XJ, Fredrickson JK, Chen XY, Kennedy DW, Murray CJ, Rockhold ML, Konopka A (2013) Quantifying community assembly processes and identifying features that impose them. ISME J 7:2069–2079. https://doi.org/10.1038/ismej.2013.93
Tang JY, Dai YX, Wang Y, Qin JG, Li YM (2015) Improvement of fish and pearl yields and nutrient utilization efficiency through fish-mussel integration and feed supplementation. Aquaculture 448:321–326. https://doi.org/10.1016/j.aquaculture.2015.05.028
Tian B, Hua YJ (2010) Carotenoid biosynthesis in extremophilic Deinococcus–Thermus bacteria. Trends Microbiol 18:512–520. https://doi.org/10.1016/j.tim.2010.07.007
Tsuchiya C, Sakata T, Sugita H (2008) Novel ecological niche of Cetobacterium somerae, an anaerobic bacterium in the intestinal tracts of freshwater fish. Lett Appl Microbiol 46:43–48. https://doi.org/10.1111/j.1472-765X.2007.02258.x
Tucker S, Pollard P (2005) Identification of cyanophage Ma-LBP and infection of the cyanobacterium Microcystis aeruginosa from an Australian subtropical lake by the virus. Appl Environ Microbiol 71:629–635. https://doi.org/10.1128/Aem.71.2.629-635.2005
Verdugo P, Alldredge AL, Azam F, Kirchman DL, Passow U, Santschi PH (2004) The oceanic gel phase: a bridge in the DOM–POM continuum. Mar Chem 92:67–85. https://doi.org/10.1016/j.marchem.2004.06.017
Wang X, Li ZJ, Su JQ, Tian Y, Ning XR, Hong HS, Zheng TL (2010) Lysis of a red-tide causing alga, Alexandrium tamarense, caused by bacteria from its phycosphere. Biol Control 52:123–130. https://doi.org/10.1016/j.biocontrol.2009.10.004
Williams TJ, Cavicchioli R (2014) Marine metaproteomics: deciphering the microbial metabolic food web. Trends Microbiol 22:248–260. https://doi.org/10.1016/j.tim.2014.03.004
Wood ED, Armstrong FAJ, Richards FA (1967) Determination of nitrate in sea water by cadmium-copper reduction to nitrite. J Mar Biol Assoc UK 47:23–31. https://doi.org/10.1017/S002531540003352X
Worm J, Sondergaard M (1998) Dynamics of heterotrophic bacteria attached to Microcystis spp. (Cyanobacteria). Aquat Microb Ecol 14:19–28. https://doi.org/10.3354/ame014019
Xiong J (2007) Photosynthesis: what color was its origin? Genome Biol 7:245. https://doi.org/10.1186/gb-2006-7-12-245
Xu Z, Woodhouse JN, Te SH, Yew-Hoong Gin K, He Y, Xu C, Chen L (2018) Seasonal variation in the bacterial community composition of a large estuarine reservoir and response to cyanobacterial proliferation. Chemosphere 202:576–585. https://doi.org/10.1016/j.chemosphere.2018.03.037
Zhang H, Jia J, Chen S, Huang T, Wang Y, Zhao Z, Feng J, Hao H, Li S, Ma X (2018) Dynamics of bacterial and fungal communities during the outbreak and decline of an algal bloom in a drinking water reservoir. Int J Environ Res Public Health 15:361. https://doi.org/10.3390/ijerph15020361
Zheng XF, Tang JY, Zhang CF, Qin JG, Wang Y (2017) Bacterial composition, abundance and diversity in fish polyculture and mussel-fish integrated cultured ponds in China. Aquac Res 48:3950–3961. https://doi.org/10.1111/are.13221
Zhou JZ, Ning DL (2017) Stochastic community assembly: does it matter in microbial ecology? Microbiol Mol Biol Rev 81:e00002–e00017. https://doi.org/10.1128/MMBR.00002-17
Zhou J, Chen GF, Ying KZ, Jin H, Song JT, Cai ZH (2019) Phycosphere microbial succession patterns and assembly mechanisms in a marine dinoflagellate bloom. Appl Environ Microbiol. https://doi.org/10.1128/AEM.00349-19
Zimba PV, Moeller PD, Beauchesne K, Lane HE, Triemer RE (2010) Identification of euglenophycin—a toxin found in certain euglenoids. Toxicon 55:100–104. https://doi.org/10.1016/j.toxicon.2009.07.004
Acknowledgements
We thanked Dr. Tang Jinyu for the help of sample collection and phytoplankton counting. This work was supported by the Special Fund for Agro-scientific Research in the Public Interest (201303056-2), National Natural Science Foundation of China (31672262), and the Fundamental Research Funds for the Central Universities (19lgpy164, 19lgzd28).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Handling Editor: Télesphore Sime-Ngando.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zheng, X., Wang, Y., Yang, T. et al. Size-fractioned aggregates within phycosphere define functional bacterial communities related to Microcystis aeruginosa and Euglena sanguinea blooms. Aquat Ecol 54, 609–623 (2020). https://doi.org/10.1007/s10452-020-09762-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10452-020-09762-0