Abstract
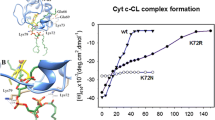
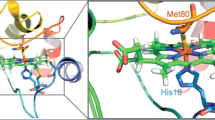
The interaction of cytochrome c with cardiolipin (CL) is a critical step in the initial stages of apoptosis and is mediated by a positively charged region on the protein surface comprising several lysine residues (site A). Here, the interaction of wt S. cerevisiae cytochrome c (ycc) and its K72A/K73A, K72A/K79A, K73A/K79A and K72A/K73A/K79A variants with CL was studied through UV–Vis and MCD spectroscopies at pH 7 and molecular dynamics (MD) simulations, to clarify the role of the mutated lysines. Moreover, the influence of the lipid to protein ratio on the interaction mechanism was investigated using low (0.5–10) and high (5–60) CL/ycc molar ratios, obtained with small and gradual or large and abrupt CL additions, respectively. Although all proteins bind to CL, switching from the native low-spin His/Met-ligated form to a low-spin bis-His conformer and to a high-spin species at larger CL concentrations, the two schemes of CL addition show relevant differences in the CL/ycc molar ratios at which the various conformers appear, due to differences in the interaction mechanism. Extended lipid anchorage and peripheral binding appear to prevail at low and high CL/ycc molar ratios, respectively. Simultaneous deletion of two or three surface positive charges from Site A does not abolish CL binding, but instead increases protein affinity for CL. MD calculations suggest this unexpected behavior results from the mutation-induced severe weakening of the H-bond connecting the Nε of His26 with the backbone oxygen of Glu44, which lowers the conformational stability compared to the wt species, overcoming the decreased surface electrostatic interaction.









Similar content being viewed by others
References
Alvarez-Paggi D, Hannibal L, Castro MA et al (2017) Multifunctional cytochrome c: learning new tricks from an old dog. Chem Rev 117:13382–13460. https://doi.org/10.1021/acs.chemrev.7b00257
Bertini I, Cavallaro G, Rosato A (2006) Cytochrome c: occurrence and functions. Chem Rev 106:90–115. https://doi.org/10.1021/cr050241v
Hüttemann M, Pecina P, Rainbolt M et al (2011) The multiple functions of cytochrome c and their regulation in life and death decisions of the mammalian cell: from respiration to apoptosis. Mitochondrion 11:369–381. https://doi.org/10.1016/j.mito.2011.01.010
Moore GR, Pettigrew GW (1990) Cytochromes c. evolutionary, structural, and physicochemical aspects. Springer, Berlin, Germany
Scott RA, Mauk AG (1996) Cytochrome c—a multidisciplinary approach. University Science Books, Sausalito
Zaidi S, Hassan MI, Islam A, Ahmad F (2014) The role of key residues in structure, function, and stability of cytochrome-c. Cell Mol Life Sci 71:229–255. https://doi.org/10.1007/s00018-013-1341-1
Schweitzer-Stenner R (2018) Relating the multi-functionality of cytochrome c to membrane binding and structural conversion. Biophys Rev 10:1151–1185. https://doi.org/10.1007/s12551-018-0409-4
Battistuzzi G, Borsari M, Cowan JA et al (2002) Control of cytochrome c redox potential: axial ligation and protein environment effects. J Am Chem Soc 124:5315–5324. https://doi.org/10.1021/ja017479v
O’Reilly NJ, Magner E (2005) Electrochemistry of cytochrome c in aqueous and mixed solvent solutions: thermodynamics, kinetics, and the effect of solvent dielectric constant. Langmuir 21:1009–1014. https://doi.org/10.1021/la048796t
Crilly S, Magner E (2009) Reversible increase in the redox potential of cytochrome c in methanol. Chem Commun. https://doi.org/10.1039/b819618d
Ranieri A, Millo D, Di Rocco G et al (2015) Immobilized cytochrome c bound to cardiolipin exhibits peculiar oxidation state-dependent axial heme ligation and catalytically reduces dioxygen. J Biol Inorg Chem 20:1019–1028. https://doi.org/10.1007/s00775-015-1238-6
Ranieri A, Di Rocco G, Millo D et al (2015) Thermodynamics and kinetics of reduction and species conversion at a hydrophobic surface for mitochondrial cytochromes c and their cardiolipin adducts. Electrochim Acta 176:1019–1028. https://doi.org/10.1016/j.electacta.2015.07.065
Battistuzzi G, Borsari M, Sola M (2001) Medium and temperature effects on the redox chemistry of cytochrome c. Eur J Inorg Chem 2001:2989. https://doi.org/10.1002/1099-0682(200112)2001:12%3c2989:AID-EJIC2989%3e3.3.CO;2-5
Cherney MM, Bowler BE (2011) Protein dynamics and function: making new strides with an old warhorse, the alkaline conformational transition of cytochrome c. Coord Chem Rev 255:664–677. https://doi.org/10.1016/j.ccr.2010.09.014
Baddam S, Bowler BE (2005) Thermodynamics and kinetics of formation of the alkaline state of a Lys 79 → Ala/Lys 73 → His variant of iso-1-cytochrome c. Biochemistry 44:14956–14968. https://doi.org/10.1021/bi0515873
Schweitzer-Stenner R (2014) Cytochrome c: a multifunctional protein combining conformational rigidity with flexibility. New J Sci 2014:1–28. https://doi.org/10.1155/2014/484538
Battistuzzi G, Borsari M, Loschi L et al (1999) Thermodynamics of the alkaline transition of cytochrome c. Biochemistry 38:7900–7907. https://doi.org/10.1021/bi983060e
Battistuzzi G, Borsari M, De Rienzo F et al (2007) Free energy of transition for the individual alkaline conformers of yeast iso-1-cytochrome c. Biochemistry 46:1694–1702. https://doi.org/10.1021/bi061961e
Kagan VE, Bayir HA, Belikova NA et al (2009) Cytochrome c/cardiolipin relations in mitochondria: a kiss of death. Free Radic Biol Med 46:1439–1453
Milorey B, Schweitzer-Stenner R, Kurbaj R, Malyshka D (2019) PH-induced switch between different modes of cytochrome c binding to cardiolipin-containing liposomes. ACS Omega 4:1386–1400. https://doi.org/10.1021/acsomega.8b02574
Ascenzi P, Coletta M, Wilson MT et al (2015) Cardiolipin-cytochrome c complex: switching cytochrome c from an electron-transfer shuttle to a myoglobin- and a peroxidase-like heme-protein. IUBMB Life 67:98–109
Kalanxhi E, Wallace CJA (2007) Cytochrome c impaled: investigation of the extended lipid anchorage of a soluble protein to mitochondrial membrane models. Biochem J 187:179–187. https://doi.org/10.1042/BJ20070459
Piro MC, Santucci R, Droghetti E et al (2013) Role of lysines in cytochrome c–cardiolipin interaction. Biochemistry 52:4578–4588. https://doi.org/10.1021/bi400324c
Mohammadyani D, Yanamala N, Samhan-Arias AK et al (2018) Structural characterization of cardiolipin-driven activation of cytochrome c into a peroxidase and membrane perturbation. Biochim Biophys Acta Biomembr 1860:1057–1068. https://doi.org/10.1016/j.bbamem.2018.01.009
Hannibal L, Tomasina F, Capdevila DA et al (2016) Alternative conformations of cytochrome c: structure, function, and detection. Biochemistry 55:407–428. https://doi.org/10.1021/acs.biochem.5b01385
Pandiscia LA, Schweitzer-Stenner R (2015) Coexistence of native-like and non-native cytochrome c on anionic liposomes with different cardiolipin content. J Phys Chem B 119:12846–12859. https://doi.org/10.1021/acs.jpcb.5b07328
Pandiscia L, Schweitzer-Stenner R (2015) The role of salt in mitochondria: returning cytochrome C to its native state after its dissociation from cardiolipin containing membranes. Biophys J 106:517a. https://doi.org/10.1016/j.bpj.2013.11.2887
Li M, Mandal A, Tyurin VA et al (2019) Surface-binding to cardiolipin nanodomains triggers cytochrome c pro-apoptotic peroxidase activity via localized dynamics. Structure. https://doi.org/10.1016/j.str.2019.02.007
Belikova NA, Kurnikov IV, Kagan VE et al (2006) Peroxidase activity and structural transitions of cytochrome c bound to cardiolipin-containing membranes. Biochemistry 45:4998–5009. https://doi.org/10.1021/bi0525573
Milorey B, Malyshka D, Schweitzer-Stenner R (2017) pH Dependence of Ferricytochrome c Conformational transitions during binding to cardiolipin membranes: evidence for histidine as the distal ligand at neutral pH. J Phys Chem Lett 8:1993–1998. https://doi.org/10.1021/acs.jpclett.7b00597
Bradley JM, Silkstone G, Wilson MT et al (2011) Probing a complex of cytochrome c and cardiolipin by magnetic circular dichroism spectroscopy: implications for the initial events in apoptosis. J Am Chem Soc 133:19676–19679. https://doi.org/10.1021/ja209144h
Rajagopal BS, Silkstone GG, Nicholls P et al (2012) An investigation into a cardiolipin acyl chain insertion site in cytochrome c. Biochim Biophys Acta Bioenerg 1817:780–791. https://doi.org/10.1016/j.bbabio.2012.02.010
Hanske J, Toffey JR, Morenz AM et al (2011) Conformational properties of cardiolipin-bound cytochrome c. Proc Natl Acad Sci 109:125–130. https://doi.org/10.1073/pnas.1112312108
Hong Y, Muenzner J, Grimm SK, Pletneva EV (2012) Origin of the conformational heterogeneity of cardiolipin-bound cytochrome c. J Am Chem Soc 134:18713–18723. https://doi.org/10.1021/ja307426k
Muenzner J, Pletneva EV (2014) Structural transformations of cytochrome c upon interaction with cardiolipin. Chem Phys Lipids 179:57–63. https://doi.org/10.1016/j.chemphyslip.2013.11.002
Kawai C, Ferreira JC, Baptista MS, Nantes IL (2014) Not only oxidation of cardiolipin affects the affinity of cytochrome c for lipid bilayers. J Phys Chem B 118:11863–11872. https://doi.org/10.1021/jp504518g
Muenzner J, Toffey JR, Hong Y, Pletneva EV (2013) Becoming a peroxidase: cardiolipin-induced unfolding of cytochrome c. J Phys Chem B 117:12878–12886. https://doi.org/10.1021/jp402104r
Oellerich S, Wackerbarth H, Hildebrandt P (2002) Spectroscopic characterization of nonnative conformational states of cytochrome c. J Phys Chem B 106:6566–6580. https://doi.org/10.1021/jp013841g
Tuominen EKJ, Wallace CJA, Kinnunen PKJ (2002) Phospholipid-cytochrome c interaction. Evidence for the extended lipid anchorage. J Biol Chem 277:8822–8826. https://doi.org/10.1074/jbc.M200056200
Kalanxhi E, Wallace CJA (2007) Cytochrome c impaled: investigation of the extended lipid anchorage of a soluble protein to mitochondrial membrane models. Biochem J 407:179–187. https://doi.org/10.1042/bj20070459
Sinibaldi F, Fiorucci L, Patriarca A et al (2008) Insights into cytochrome c-cardiolipin interaction. Role played by ionic strength. Biochemistry 47:6928–6935. https://doi.org/10.1021/bi800048v
Sinibaldi F, Howes BD, Piro MC et al (2010) Extended cardiolipin anchorage to cytochrome c: a model for protein-mitochondrial membrane binding. J Biol Inorg Chem 15:689–700. https://doi.org/10.1007/s00775-010-0636-z
Sinibaldi F, Droghetti E, Polticelli F et al (2011) The effects of ATP and sodium chloride on the cytochrome c-cardiolipin interaction: the contrasting behavior of the horse heart and yeast proteins. J Inorg Biochem 105:1365–1372. https://doi.org/10.1016/j.jinorgbio.2011.07.022
Milazzo L, Tognaccini L, Howes BD et al (2017) Unravelling the non-native low-spin state of the cytochrome c-cardiolipin complex: evidence of the formation of a his–ligated species only. Biochemistry 56:1887–1898. https://doi.org/10.1021/acs.biochem.6b01281
Silkstone G, Kapetanaki SM, Husu I et al (2012) Nitric oxide binding to the cardiolipin complex of ferric cytochrome. Biochemistry 51:6760–6766. https://doi.org/10.1021/bi300596
Husu I, Kapetanaki S, Silkstone G et al (2010) Interaction of CO/NO with the apoptosis-inducing cytochrome C–cardiolipin complex. Biophys J 98:631a. https://doi.org/10.1016/j.bpj.2009.12.3455
Kapetanaki SM, Silkstone G, Husu I et al (2009) Interaction of carbon monoxide with the apoptosis-inducing cytochrome c-cardiolipin complex. Biochemistry 48:1613–1619. https://doi.org/10.1021/bi801817v
Elmer-Dixon MM, Bowler BE (2017) Site A-mediated partial unfolding of cytochrome c on cardiolipin vesicles Is species-dependent and does not require Lys72. Biochemistry 56:4830–4839. https://doi.org/10.1021/acs.biochem.7b00694
Elmer-Dixon MM, Bowler BE (2018) Electrostatic constituents of the interaction of cardiolipin with site A of cytochrome c. Biochemistry 57:5683–5695. https://doi.org/10.1021/acs.biochem.8b00704
Pandiscia LA, Schweitzer-Stenner R (2014) Salt as a catalyst in the mitochondria: returning cytochrome c to its native state after it misfolds on the surface of cardiolipin containing membranes. Chem Commun 50:3674–3676. https://doi.org/10.1039/c3cc48709a
Sinibaldi F, Milazzo L, Howes BD et al (2017) The key role played by charge in the interaction of cytochrome c with cardiolipin. J Biol Inorg Chem 22:19–29. https://doi.org/10.1007/s00775-016-1404-5
Ascenzi P, Sbardella D, Sinibaldi F et al (2016) The nitrite reductase activity of horse heart carboxymethylated-cytochrome c is modulated by cardiolipin. J Biol Inorg Chem 21:421–432. https://doi.org/10.1007/s00775-016-1351-1
Zeng L, Wu L, Liu L, Jiang X (2016) Analyzing structural properties of heterogeneous cardiolipin-bound cytochrome c and their regulation by surface-enhanced infrared absorption spectroscopy. Anal Chem 88:11727–11733. https://doi.org/10.1021/acs.analchem.6b03360
Paul SS, Sil P, Haldar S et al (2015) Subtle change in the charge distribution of surface residues may affect the secondary functions of cytochrome c. J Biol Chem 290:14476–14490. https://doi.org/10.1074/jbc.M114.607010
Godoy LC, Muñoz-Pinedo C, Castro L et al (2009) Disruption of the M80-Fe ligation stimulates the translocation of cytochrome c to the cytoplasm and nucleus in nonapoptotic cells. Proc Natl Acad Sci 106:2653–2658. https://doi.org/10.1073/pnas.0809279106
Capdevila DA, Oviedo RS, Tomasina F et al (2015) Active site structure and peroxidase activity of oxidatively modified cytochrome c species in complexes with cardiolipin. Biochemistry 54:7491–7504. https://doi.org/10.1021/acs.biochem.5b00922
Abe M, Niibayashi R, Koubori S et al (2011) Molecular mechanisms for the induction of peroxidase activity of the cytochrome c-cardiolipin complex. Biochemistry 50:8383–8391. https://doi.org/10.1021/bi2010202
Kagan VE, Bayir A, Bayir H et al (2009) Mitochondria-targeted disruptors and inhibitors of cytochrome c/cardiolipin peroxidase complexes: a new strategy in anti-apoptotic drug discovery. Mol Nutr Food Res 53:104–114
Kagan VE, Tyurin VA, Jiang J et al (2005) Cytochrome c acts as a cardiolipin oxygenase required for release of proapoptotic factors. Nat Chem Biol 1:223–232. https://doi.org/10.1038/nchembio727
Planas-Iglesias J, Dwarakanath H, Mohammadyani D et al (2015) Cardiolipin interactions with proteins. Biophys J 109:1282–1294. https://doi.org/10.1016/j.bpj.2015.07.034
Zeng L, Wu L, Liu L, Jiang X (2017) The role of water distribution controlled by transmembrane potentials in the cytochrome c–cardiolipin interaction: revealing from surface-enhanced infrared absorption spectroscopy. Chem A Eur J 23:15491–15497. https://doi.org/10.1002/chem.201703400
Thong A, Tsoukanova V (2018) Cytochrome-c-assisted escape of cardiolipin from a model mitochondrial membrane. Biochim Biophys Acta Biomembr 1860:475–480. https://doi.org/10.1016/j.bbamem.2017.10.032
Spooner PJR, Watts A (1992) Cytochrome c interactions with cardiolipin in bilayers: a multinuclear magic-angle spinning NMR study. Biochemistry 31:10129–10138. https://doi.org/10.1021/bi00156a037
Basova LV, Kurnikov IV, Wang L et al (2007) Cardiolipin switch in mitochondria: shutting off the reduction of cytochrome c and turning on the peroxidase activity. Biochemistry 46:3423–3434. https://doi.org/10.1021/bi061854k
Kapralov AA, Kurnikov IV, Vlasova II et al (2007) The hierarchy of structural transitions induced in cytochrome c by anionic phospholipids determines its peroxidase activation and selective peroxidation during apoptosis in cells. Biochemistry 46:14232–14244. https://doi.org/10.1021/bi701237b
Kawai C, Prado FM, Nunes GLC et al (2005) pH-dependent interaction of cytochrome c with mitochondrial mimetic membranes: the role of an array of positively charged amino acids. J Biol Chem 280:34709–34717. https://doi.org/10.1074/jbc.M412532200
Rytomaa M, Mustonen P, Kinnunen PKJ (1992) Reversible, nonionic, and pH-dependent association of cytochrome c with cardiolipin-phosphatidylcholine liposomes. J Biol Chem 267:22243–22248
Rytömaa M, Kinnunen PKJ (1994) Evidence for two distinct acidic phospholipid-binding sites in cytochrome c. J Biol Chem 269:1770–1774
Rytomaa M, Kinnunen PKJ (1995) Reversibility of the binding of cytochrome c to liposomes. Implications for lipid–protein interactions. J Biol Chem 270:3197–3202. https://doi.org/10.1074/jbc.270.7.3197
O’Brien ES, Nucci NV, Fuglestad B et al (2015) Defining the apoptotic trigger: the interaction of cytochrome c and cardiolipin. J Biol Chem 290:30879–30887. https://doi.org/10.1074/jbc.M115.689406
Kobayashi H, Nagao S, Hirota S (2016) Characterization of the cytochrome c membrane-binding site using cardiolipin-containing bicelles with NMR. Angew Chemie Int Ed 55:14019–14022. https://doi.org/10.1002/anie.201607419
Balakrishnan G, Hu Y, Spiro TG et al (2012) His26 protonation in cytochrome c triggers microsecond β-sheet formation and heme exposure: implications for apoptosis. J Am Chem Soc 134:19061–19069. https://doi.org/10.1021/ja307100a
Snider EJ, Muenzner J, Toffey JR et al (2013) Multifaceted effects of ATP on cardiolipin-bound cytochrome c. Biochemistry 52:993–995. https://doi.org/10.1021/bi301682c
Pandiscia LA, Schweitzer-Stenner R (2015) Coexistence of native-like and non-native partially unfolded ferricytochrome c on the surface of cardiolipin-containing liposomes. J Phys Chem B 119:1334–1349. https://doi.org/10.1021/jp5104752
Patriarca A, Eliseo T, Sinibaldi F et al (2009) ATP acts as a regulatory effector in modulating structural transitions of cytochrome c : implications for apoptotic activity. Biochemistry 9:3279–3287. https://doi.org/10.1021/bi801837e
Battistuzzi G, Borsari M, Bortolotti CA et al (2007) Effects of mutational (Lys to Ala) surface charge changes on the redox properties of electrode-immobilized cytochrome c. J Phys Chem B 111:10281–10287. https://doi.org/10.1021/jp0730343
Casalini S, Battistuzzi G, Borsari M et al (2010) Electron transfer properties and hydrogen peroxide electrocatalysis of cytochrome c variants at positions 67 and 80. J Phys Chem B 114:1698–1706. https://doi.org/10.1021/jp9090365
Battistuzzi G, Bortolotti CA, Bellei M et al (2012) Role of Met80 and Tyr67 in the low-pH conformational equilibria of cytochrome c. Biochemistry 51:5967–5978. https://doi.org/10.1021/bi3007302
Di Rocco G, Battistuzzi G, Bortolotti CA et al (2011) Cloning, expression, and physicochemical characterization of a new diheme cytochrome c from Shewanella baltica OS155. J Biol Inorg Chem 16:461–471. https://doi.org/10.1007/s00775-010-0742-y
Bellei M, Bortolotti CA, Di Rocco G et al (2018) The influence of the Cys46/Cys55 disulfide bond on the redox and spectroscopic properties of human neuroglobin. J Inorg Biochem 178:70–86. https://doi.org/10.1016/j.jinorgbio.2017.10.005
Berendsen HJC, van der Spoel D, van Drunen R (1995) GROMACS: a message-passing parallel molecular dynamics implementation. Comput Phys Commun 91:43–56. https://doi.org/10.1016/0010-4655(95)00042-E
Berendsen HJC, Grigera JR, Straatsma TP (1987) The missing term in effective pair potentials. J Phys Chem 91:6269–6271. https://doi.org/10.1021/j100308a038
Paltrinieri L, Borsari M, Ranieri A et al (2013) The active site loop modulates the reorganization energy of blue copper proteins by controlling the dynamic interplay with solvent. J Phys Chem Lett 4:710–715. https://doi.org/10.1021/jz302125k
Zanetti-Polzi L, Daidone I, Bortolotti CA, Corni S (2014) Surface packing determines the redox potential shift of cytochrome c adsorbed on gold. J Am Chem Soc 136:12929–12937. https://doi.org/10.1021/ja505251a
Bortolotti CA, Amadei A, Aschi M et al (2012) The reversible opening of water channels in cytochrome c modulates the heme iron reduction potential. J Am Chem Soc 134:13670–13678. https://doi.org/10.1021/ja3030356
Brown D, Clarke JHR (1984) Comparison of constant energy, constant temperature, and constant pressure ensembles in molecular dynamics simulations of atomic liquids. Mol Phys 51:1243–1252
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472. https://doi.org/10.1002/(SICI)1096-987X(199709)18:12%3c1463:AID-JCC4%3e3.0.CO;2-H
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N·log(N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092
Petřek M, Otyepka M, Banáš P et al (2006) CAVER: a new tool to explore routes from protein clefts, pockets and cavities. BMC Bioinform 7:1–9. https://doi.org/10.1186/1471-2105-7-316
Walker FA (2002) Magnetic spectroscopic (EPR, ESEEM, Mössbauer, MCD and NMR) studies of low-spin ferriheme centers and their corresponding heme proteins. Coord Chem Rev 185–186:471–534. https://doi.org/10.1016/s0010-8545(99)00029-6
Kirk ML, Peariso K (2003) Recent applications of MCD spectroscopy to metalloenzymes. Curr Opin Chem Biol 7:220–227
McMaster J, Oganesyan VS (2010) Magnetic circular dichroism spectroscopy as a probe of the structures of the metal sites in metalloproteins. Curr Opin Struct Biol 20:615–622
Fedurco M, Augustynski J, Indiani C et al (2004) The heme iron coordination of unfolded ferric and ferrous cytochrome c in neutral and acidic urea solutions. Spectroscopic and electrochemical studies. Biochim Biophys Acta Proteins Proteom 1703:31–41. https://doi.org/10.1016/j.bbapap.2004.09.013
Cheek J, Dawson JH (2000) Magnetic circular dichroism spectroscopy of heme proteins and model systems. In: Kadish KM, Smith KM, Guilard R (eds) Porphyrin handbook, vol 3. Academic Press, San Diego, pp 339–369
Vickery L, Nozawa T, Sauer K (1976) Magnetic circular dichroism studies of low-spin cytochromes. Temperature dependence and effects of axial coordination on the spectra of cytochrome c and cytochrome b5. J Am Chem Soc 98:351–357. https://doi.org/10.1021/ja00418a006
Vickery L, Sauer K, Nozawa T (1976) Magnetic circular dichroism studies of myoglobin complexes. Correlations with heme spin state and axial ligation. J Am Chem Soc 98:343–350. https://doi.org/10.1021/ja00418a005
Du J, Sono M, Dawson JH (2011) The H93G myoglobin cavity mutant as a versatile scaffold for modeling heme iron coordination structures in protein active sites and their characterization with magnetic circular dichroism spectroscopy. Coord Chem Rev 255:700–716
Pond AE, Roach MP, Thomas MR et al (2000) The H93G myoglobin cavity mutant as a versatile template for modeling heme proteins: ferrous, ferric, and ferryl mixed-ligand complexes with imidazole in the cavity. Inorg Chem 39:6061–6066. https://doi.org/10.1021/ic0007198
Du J, Perera R, Dawson JH (2011) Alkylamine-ligated H93G myoglobin cavity mutant: a model system for endogenous lysine and terminal amine ligation in heme proteins such as nitrite reductase and cytochrome f. Inorg Chem 50:1242–1249. https://doi.org/10.1021/ic101644u
Que LJ (2000) Physical methods in bioinorganic chemistry—spectroscopy and magnetism. University Science Books, Sausalito
McClelland LJ, Steele HBB, Whitby FG et al (2016) Cytochrome c can form a well-defined binding pocket for hydrocarbons. J Am Chem Soc 138:16770–16778. https://doi.org/10.1021/jacs.6b10745
Krishna MMG, Lin Y, Rumbley JN, Englander SW (2003) Cooperative omega loops in cytochrome c: role in folding and function. J Mol Biol 2836:29–36. https://doi.org/10.1016/S0022-2836(03)00697-1
Hoang L, Maity H, Krishna MMG et al (2003) Folding units govern the cytochrome c alkaline transition. J Mol Biol 2836:37–43. https://doi.org/10.1016/S0022-2836(03)00698-3
Milne JS, Xu Y, Mayne LC, Englander SW (1999) Experimental study of the protein folding landscape : unfolding reactions in cytochrome c. J Mol Biol 290:811–822. https://doi.org/10.1006/jmbi.1999.2924
Englander SW, Sosnick TR, Mayne LC et al (2002) Fast and slow folding in cytochrome c. Acc Chem Res 31:737–744. https://doi.org/10.1021/ar970085h
Sinibaldi F, Piro MC, Howes BD et al (2003) Rupture of the hydrogen bond linking two ω-loops induces the molten globule state at neutral pH in cytochrome c. Biochemistry 42:7604–7610. https://doi.org/10.1021/bi034132r
Baddam S, Bowler BE (2006) Mutation of asparagine 52 to glycine promotes the alkaline form of iso-1-cytochrome c and causes loss of cooperativity in acid unfolding. Biochemistry 45:4611–4619. https://doi.org/10.1021/bi0524971
Baddam S, Bowler BE (2005) Conformationally gated electron transfer in iso-1-cytochrome c: engineering the rate of a conformational switch. J Am Chem Soc 127:9702–9703. https://doi.org/10.1021/ja0527368
Bandi S, Baddam S, Bowler BE (2007) Alkaline conformational transition and gated electron transfer with a Lys 79 → His variant of iso-1-cytochrome c. Biochemistry 46:10643–10654. https://doi.org/10.1021/bi700992y
Funding
This work was supported by the University of Modena and Reggio Emilia FAR 2016 and FAR 2019 funding programs.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Paradisi, A., Bellei, M., Paltrinieri, L. et al. Binding of S. cerevisiae iso-1 cytochrome c and its surface lysine-to-alanine variants to cardiolipin: charge effects and the role of the lipid to protein ratio. J Biol Inorg Chem 25, 467–487 (2020). https://doi.org/10.1007/s00775-020-01776-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-020-01776-1