Abstract
T-cell-secreted interferon (IFN)-γ can exert pleiotropic effects on tumor cells that include induction of immune checkpoints and antigen presentation machinery components, and inhibition of cell growth. Despite its role as a key effector molecule, little is known about the spatiotemporal spreading of IFN-γ secreted by activated CD8+ T cells within the tumor environment. Using multiday intravital imaging, we demonstrate that T cell recognition of a minor fraction of tumor cells leads to sensing of IFN-γ by a large part of the tumor mass. Furthermore, imaging of tumors in which antigen-positive and antigen-negative tumor cells are separated in space reveals spreading of the IFN-γ response, reaching distances of >800 µm. Notably, long-range sensing of IFN-γ can modify tumor behavior, as shown by both induction of PD-L1 expression and inhibition of tumor growth. Collectively, these data reveal how, through IFN-γ, CD8+ T cells modulate the behavior of remote tumor cells, including antigen-loss variants.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Statistical source data for all figures and extended data figures including all independent repeats are provided online.
All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
Change history
17 June 2020
A Correction to this paper has been published: https://doi.org/10.1038/s43018-020-0092-9
References
Durgeau, A., Virk, Y., Corgnac, S. & Mami-Chouaib, F. Recent advances in targeting CD8 T-cell immunity for more effective cancer immunotherapy. Front. Immunol. 9, 14 (2018).
Hadrup, S., Donia, M. & Thor Straten, P. Effector CD4 and CD8 T cells and their role in the tumor microenvironment. Cancer Microenviron. 6, 123–133 (2013).
Galon, J. et al. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313, 1960–1964 (2006).
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Tumeh, P. C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014).
Dudley, M. E. et al. Randomized selection design trial evaluating CD8+-enriched versus unselected tumor-infiltrating lymphocytes for adoptive cell therapy for patients with melanoma. J. Clin. Oncol. 31, 2152–2159 (2013).
Rosenberg, S. A. et al. Durable complete responses in heavily pretreated patients with metastatic melanoma using T-cell transfer immunotherapy. Clin. Cancer Res. 17, 4550–4557 (2011).
Yang, J. C. & Rosenberg, S. A. Adoptive T-cell therapy for cancer. Adv. Immunol. 130, 279–294 (2016).
Rohaan, M. W., van den Berg, J. H., Kvistborg, P. & Haanen, J. Adoptive transfer of tumor-infiltrating lymphocytes in melanoma: a viable treatment option. J. Immunother. Cancer 6, 102 (2018).
Beck, R. J., Slagter, M. & Beltman, J. B. Contact-dependent killing by cytotoxic T lymphocytes is insufficient for EL4 tumor regression in vivo. Cancer Res. 79, 3406–3416 (2019).
Nakajima, C. et al. A role of interferon-gamma (IFN-gamma) in tumor immunity: T cells with the capacity to reject tumor cells are generated but fail to migrate to tumor sites in IFN-gamma-deficient mice. Cancer Res. 61, 3399–3405 (2001).
Mojic, M., Takeda, K. & Hayakawa, Y. The dark side of IFN-gamma: its role in promoting cancer immunoevasion. Int. J. Mol. Sci. 19, E89 (2017).
Castro, F., Cardoso, A. P., Goncalves, R. M., Serre, K. & Oliveira, M. J. Interferon-gamma at the crossroads of tumor immune surveillance or evasion. Front. Immunol. 9, 847 (2018).
Schroder, K., Hertzog, P. J., Ravasi, T. & Hume, D. A. Interferon-gamma: an overview of signals, mechanisms and functions. J. Leukoc. Biol. 75, 163–189 (2004).
Braumuller, H. et al. T-helper-1-cell cytokines drive cancer into senescence. Nature 494, 361–365 (2013).
Kammertoens, T. et al. Tumour ischaemia by interferon-gamma resembles physiological blood vessel regression. Nature 545, 98–102 (2017).
Shankaran, V. et al. IFNgamma and lymphocytes prevent primary tumour development and shape tumour immunogenicity. Nature 410, 1107–1111 (2001).
Manguso, R. T. et al. In vivo CRISPR screening identifies Ptpn2 as a cancer immunotherapy target. Nature 547, 413–418 (2017).
Patel, S. J. et al. Identification of essential genes for cancer immunotherapy. Nature 548, 537–542 (2017).
Kearney, C. J. et al. Tumor immune evasion arises through loss of TNF sensitivity. Sci. Immunol. 3, eaar3451 (2018).
Pan, D. et al. A major chromatin regulator determines resistance of tumor cells to T cell-mediated killing. Science 359, 770–775 (2018).
Zaretsky, J. M. et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. NEJM 375, 819–829 (2016).
Sucker, A. et al. Acquired IFNgamma resistance impairs anti-tumor immunity and gives rise to T-cell-resistant melanoma lesions. Nat. Commun. 8, 15440 (2017).
Gao, J. et al. Loss of IFN-gamma pathway genes in tumor cells as a mechanism of resistance to anti-CTLA-4 therapy. Cell 167, 397–404 e399 (2016).
Shin, D. S. et al. Primary resistance to PD-1 blockade mediated by JAK1/2 mutations. Cancer Discov. 7, 188–201 (2017).
Briesemeister, D. et al. Tumor rejection by local interferon gamma induction in established tumors is associated with blood vessel destruction and necrosis. Int. J. Cancer 128, 371–378 (2011).
Spiotto, M. T., Rowley, D. A. & Schreiber, H. Bystander elimination of antigen loss variants in established tumors. Nat. Med. 10, 294–298 (2004).
Spiotto, M. T. & Schreiber, H. Rapid destruction of the tumor microenvironment by CTLs recognizing cancer-specific antigens cross-presented by stromal cells. Cancer Immun. 5, 8 (2005).
Zhang, B. et al. Induced sensitization of tumor stroma leads to eradication of established cancer by T cells. J. Exp. Med. 204, 49–55 (2007).
Bhatia, A. & Kumar, Y. Cellular and molecular mechanisms in cancer immune escape: a comprehensive review. Expert Rev. Clin. Immunol. 10, 41–62 (2014).
Muller, A. J. et al. CD4+ T cells rely on a cytokine gradient to control intracellular pathogens beyond sites of antigen presentation. Immunity 37, 147–157 (2012).
Perona-Wright, G., Mohrs, K. & Mohrs, M. Sustained signaling by canonical helper T cell cytokines throughout the reactive lymph node. Nat. Immunol. 11, 520–526 (2010).
Barcia, C. et al. In vivo polarization of IFN-gamma at Kupfer and non-Kupfer immunological synapses during the clearance of virally infected brain cells. J. Immunol. 180, 1344–1352 (2008).
Sanderson, N. S. et al. Cytotoxic immunological synapses do not restrict the action of interferon-gamma to antigenic target cells. Proc. Natl Acad. Sci. USA 109, 7835–7840 (2012).
Stronen, E. et al. Targeting of cancer neoantigens with donor-derived T cell receptor repertoires. Science 352, 1337–1341 (2016).
Ivashkiv, L. B. & Donlin, L. T. Regulation of type I interferon responses. Nat. Rev. Immunol. 14, 36–49 (2014).
Kotredes, K. P. & Gamero, A. M. Interferons as inducers of apoptosis in malignant cells. J. Interferon Cytokine Res. 33, 162–170 (2013).
Wang, W. et al. CD8+ T cells regulate tumour ferroptosis during cancer immunotherapy. Nature 569, 270–274 (2019).
Rouwkema, J., Rivron, N. C. & van Blitterswijk, C. A. Vascularization in tissue engineering. Trends Biotechnol. 26, 434–441 (2008).
Benci, J. L. et al. Tumor interferon signaling regulates a multigenic resistance program to immune checkpoint blockade. Cell 167, 1540–1554.e1512 (2016).
Pai, C. S. et al. Clonal deletion of tumor-specific T cells by interferon-gamma confers therapeutic resistance to combination immune checkpoint blockade. Immunity 50, 477–492 e478 (2019).
Benci, J. L. et al. Opposing functions of interferon coordinate adaptive and innate immune responses to cancer immune checkpoint blockade. Cell 178, 933–948.e914 (2019).
Workel, H. H. et al. A transcriptionally distinct CXCL13+CD103+CD8+ T-cell population is associated with B-cell recruitment and neoantigen load in human cancer. Cancer Immunol. Res. 7, 784–796 (2019).
Thommen, D. S. et al. A transcriptionally and functionally distinct PD-1+ CD8+ T cell pool with predictive potential in non-small-cell lung cancer treated with PD-1 blockade. Nat. Med. 24, 994–1004 (2018).
Farhood, B., Najafi, M. & Mortezaee, K. CD8+ cytotoxic T lymphocytes in cancer immunotherapy: a review. J. Cell. Physiol. 234, 8509–8521 (2019).
Nagarsheth, N., Wicha, M. S. & Zou, W. Chemokines in the cancer microenvironment and their relevance in cancer immunotherapy. Nat. Rev. Immunol. 17, 559–572 (2017).
Platanias, L. C. Mechanisms of type-I- and type-II-interferon-mediated signalling. Nat. Rev. Immunol. 5, 375–386 (2005).
van Rooij, N. et al. Tumor exome analysis reveals neoantigen-specific T-cell reactivity in an ipilimumab-responsive melanoma. J. Clin. Oncol. 31, e439–e442 (2013).
Jorritsma, A. et al. Selecting highly affine and well-expressed TCRs for gene therapy of melanoma. Blood 110, 3564–3572 (2007).
Linnemann, C. et al. High-throughput identification of antigen-specific TCRs by TCR gene capture. Nat. Med. 19, 1534–1541 (2013).
Acknowledgements
We thank E. Beerling, M. Mertz, B. van den Broek, L. Fanchi, R. Mezzadra, D. Peters, A. Broeks and the staff at the NKI Animal facility, Flow Cytometry facility, Bioimaging facility and Molecular Pathology & Biobanking facility for technical support and input, and members of the Schumacher and van Rheenen laboratories for discussions. This work was supported by a Boehringer Ingelheim Fonds PhD Fellowship (to M.E.H.), a Swiss National Science Foundation APM fellowship (no. P300PB_177881; to D.S.T.), ERC CoG Cancer-recurrence (grant no. 648804), Cancer Genomics Netherlands and the Doctor Josef Steiner Foundation (to J.v.R.) and ERC AdG SENSIT (to T.N.M.S.).
Author information
Authors and Affiliations
Contributions
M.E.H., L.B., D.P. and I.N.P. performed experiments and analyzed data. L.B. performed multiphoton imaging, F.E.D. and M.E.H. designed and tested the IGS reporter. L.B. and M.E.H. designed imaging analyses. M.E.H. and M.T. performed LCK inhibitor experiments. D.S.T. provided and analyzed human tumor samples. M.E.H., L.B., F.E.D., J.v.R. and T.N.M.S. contributed to experimental design. M.E.H., L.B., J.v.R. and T.N.M.S. prepared the manuscript with input from all the coauthors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 In vitro IGS reporter cell characteristics.
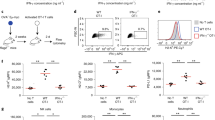
a, Median Katushka fluorescence intensity of IGS reporter-modified CFP+ OVCAR5 cells upon incubation with recombinant IFNγ under the indicated conditions. Bar graph shows mean of n=3 technical replicates, representative data of two independent experiments are depicted. b, CFP+ IGS reporter modified OVCAR5 cells proficient or deficient for the IFNγR were incubated for 48h with the indicated concentrations of recombinant IFNγ or IFNα and Katushka expression was analyzed by flow cytometry. Bar graphs show mean of technical duplicates data obtained from one experiment. c, Percentage IR-Dye positive IGS reporter-modified CFP+ OVCAR5 cells upon incubation with recombinant IFNγ under the indicated conditions. Bar graph shows mean of n=3 technical replicates. Representative data of two independent experiments are depicted. d, Percentage of IR-Dye positive Ag−CFP+ or Ag−IFNγR−/− OVCAR5 cells after 72h incubation with 100 ng/mL IFNγ. Bar graph shows mean of n=3 technical replicates. Representative data of three independent experiments are depicted.
Extended Data Fig. 2 IFNγ-induced IGS reporter and PD-L1 expression in MDA-MB-231 cells.
a, Median Katushka fluorescence intensity of IGS reporter-modified CFP+ MDA-MB-231 cells upon incubation with recombinant IFNγ under the indicated conditions. Bar graph shows mean of n=3 technical replicates. Representative data from two independent experiments are depicted. b, Median fluorescence intensity of PD-L1 staining as a function of median Katushka fluorescence intensity of CFP+Ag− IGS MDA-MB-231 reporter cells incubated for 24h with recombinant IFNγ under the indicated conditions. Plot depicts representative data three technical replicates of two independent experiments. c, 20% GFP+ Ag+ cells and 80% CFP+ IGS reporter bystander MDA-MB-231 tumor cells (5 x 105 total) were subcutaneously injected in NSG-β2m-/- mice. Mice were treated with HBSS (control), 5 x 106 control CD8+ T cells or with 5 x 106 CDK4R>L-specific CD8+ T cells, and tumors were harvested at day 3 after treatment. Bar graphs depicting mean percentage plus SD of Katushka+ reporter cells in control and tumor-specific T cell treated mice, n=5 mice per group. Representative data of two independent experiments are depicted. Two tailed Mann-Whitney U test was performed, with: p= 0.3095 (ns); p= 0.0317 (*); p= 0.0079 (**). d, Percentage of PD-L1-expressing cells of IGS reporter-modified CFP+ MDA-MB-231 cells from tumors as described in c. Bar graphs depict mean percentage of PD-L1 positive Ag- IGS cells plus SD, n=5 mice per group. Representative data of n=2 independent experiments are depicted. Two tailed Mann-Whitney U test was performed, with: p> 0.9999 (ns); p= 0.0317 (*); p= 0.0079 (**).
Extended Data Fig. 3 Distance from Ag- IGS cells to the nearest Ag+ cell upon tumor cell co-injection.
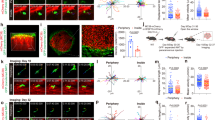
Analysis of the distance between Ag- IGS tumor cells and the nearest Ag+ tumor cell for the imaging experiments depicted in Fig. 3. a, Representative image of a tumor with intermingled Ag- and Ag+IGS cells. Scale bar is 100 μm b, Plots show the min., max., and mean of 25th and 75th percentile plus the median for n=4 mice. c, Percentage of Ag-IGS reporter cells in the indicated distance bins to the nearest Ag+ cell, depicted mean plus SD for n=4 mice. Data obtained from three independent experiments.
Extended Data Fig. 4 CD8+ T cell dependent Katushka signaling in IGS reporter cells in vivo.
a, Flow cytometric analysis of Katushka expression in CFP+ IGS reporter (left panel) and GFP+ Ag+ (right panel) cells derived from mixed tumors described in Fig. 2b. Data from mice treated with HBSS are depicted in blue, data from mice treated with CDK4R>L-specific CD8+ T cells are depicted in red, n=4 mice per condition, data obtained from one experiment. b, Representative images of tumors before and 120h after injection of CDK4R>L-specific CD8+ T cells (left panel, three independent experiments with n=1 mouse each) or HBSS (right panel, two independent experiments with n=1 mouse each), for the imaging experiments described in Fig. 3. SHG: Second-harmonic generation. Scale bar is 200 μm.
Extended Data Fig. 5 Analysis of T cell mediated loss of Ag-presenting tumor cells over time.
a, Relative GFP+ volume in tumors from imaging experiments described in Fig. 2 quantified over time. Mean and SEM are depicted for n=5 mice (n=5 mice for time 0, 16, 24, and 32 h; n=4 for 40, 48, and 72 h; n=3 for 120 h, from data obtained in all independent experiments. b, The distance between CFP+ bystander tumor cells and the closest GFP+ Ag+ tumor cell was determined at indicated time points from tumors described in Fig. 2 for n=2 mice. Data are obtained from two independent experiments, boxplot presenting the minimum, 25th percentile, median, 75th percentile and the maximum For total sample size per timepoint see Source Data ED_Fig5_source table.
Extended Data Fig. 6 CD8+ T cell quantification in Ag+ and Ag- IGS reporter tumor areas.
a, Quantification of mOrange2+ CD8+ T cells in tumors with spatially separated GFP+Ag+ (green) and CFP+Ag- IGS reporter cell (cyan) islands obtained by sequential injection, as described in Fig. 4d, e. Number of mOrange2+ T cells was determined in multiple three-dimensional stacks of 2.5*107 μm3 in either Ag+ or Ag- areas. Symbols represent individual mice, and mean and SD for n=4 mice are depicted, obtained from two independent experiments. Normal distribution was confirmed by D’Agostino and Pearson omnibus normality test. Two tailed unpaired t-tests were performed, p=0.0003 (***). b, Estimate of the ratio of tumor cells to T cells in Ag+ and Ag- areas under the assumption that the diameter of an average tumor cell is 24 μm. c, Purified CD8+ T cells were activated with plate-bound anti-CD3/anti-CD28 antibodies for 2h. Subsequently, cells were either left untreated or were treated with 5nM LCKi inhibitor for the indicated times. Cells were washed to remove previously secreted IFNγ, and fresh control medium or medium containing 5nM LCK inhibitor was added to the cells. After 3h incubation, supernatants were collected and IFNγ concentrations were analysed. Bar graph shows mean IFNγ concentrations of n=3 technical replicates. Representative data of four independent experiments are depicted. d, As in c, depicting the IFNγ concentration in supernatants obtained from 2h LCK inhibitor treated cell cultures as a percentage of IFNγ concentration in control, non treated cell cultures. Dots represent four independent experiments, using different T cell donors in each experiment.
Extended Data Fig. 7 Distinct β2m positive and negative areas in human cancers.
a, Immunohistochemical staining of β2m and b, β2m and CD8 proteins on FFPE tissue of indicated human tumors. Heterogeneous β2m signal was observed in 16/51 tumors analyzed, one representative slide per tumor (obtained from resection material) was assessed and representative images are depicted in a. a. Scale bars are 100 μm. Note that CD8+ T cells in tumors predominantly localize to β2m high regions, representative images are depicted in b. Scale bars are 250 μm.
Extended Data Fig. 8 CD8+ T cell mediated killing of bystander OVCAR5 tumor cells.
A mixture of GFP+ Ag+, CFP+ Ag- IFNγR proficient and CFP+ Ag- IFNγR deficient OVCAR5 cells (2:1:1 ratio) was treated with CDK4R>L-specific CD8+ T cells at a 2:1 T cell: tumor cell ratio, or left untreated, and cell survival was analyzed by staining with IR-Dye and subsequent flow cytometry. a, Representative plots depicting the percentage of IR-Dye+ cells for the indicated groups. b, Quantification of a, bar graph shows mean of n=3 technical replicates. Representative data of two independent experiments are depicted.
Supplementary information
Source data
Source data Fig. 1
Statistical source data of all independent experiments.
Source data Fig. 2
Statistical source data of all independent experiments.
Source data Fig. 3
Statistical source data of all independent experiments.
Source data Fig. 4
Statistical source data of all independent experiments.
Source data Fig. 5
Statistical source data of all independent experiments.
Source data Extended data Fig. 1
Statistical source data of all independent experiments.
Source data Extended data Fig. 2
Statistical source data of all independent experiments.
Source data Extended data Fig. 3
Statistical source data of all independent experiments.
Source data Extended data Fig. 5
Statistical source data of all independent experiments.
Source data Extended data Fig. 6
Statistical source data of all independent experiments.
Source data Extended data Fig. 8
Statistical source data of all independent experiments.
Rights and permissions
About this article
Cite this article
Hoekstra, M.E., Bornes, L., Dijkgraaf, F.E. et al. Long-distance modulation of bystander tumor cells by CD8+ T-cell-secreted IFN-γ. Nat Cancer 1, 291–301 (2020). https://doi.org/10.1038/s43018-020-0036-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-020-0036-4
This article is cited by
-
In vivo imaging of inflammatory response in cancer research
Inflammation and Regeneration (2023)
-
ERK mediates interferon gamma-induced melanoma cell death
Molecular Cancer (2023)
-
Tumor-intrinsic sensitivity to the pro-apoptotic effects of IFN-γ is a major determinant of CD4+ CAR T-cell antitumor activity
Nature Cancer (2023)
-
Disrupting the phase separation of KAT8–IRF1 diminishes PD-L1 expression and promotes antitumor immunity
Nature Cancer (2023)
-
Rational combinations of targeted cancer therapies: background, advances and challenges
Nature Reviews Drug Discovery (2023)