Abstract
Pyricularia oryzae is the causal agent of blast disease on staple gramineous crops. Sulphur is an essential element for the biosynthesis of cysteine and methionine in fungi. Here, we targeted the P. oryzae PoMET3 encoding the enzyme ATP sulfurylase, and PoMET14 encoding the APS (adenosine-5′-phosphosulphate) kinase that are involved in sulfate assimilation and sulphur-containing amino acids biosynthesis. In P. oryzae, deletion of PoMET3 or PoMET14 separately results in defects of conidiophore formation, significant impairments in conidiation, methionine and cysteine auxotrophy, limited invasive hypha extension, and remarkably reduced virulence on rice and barley. Furthermore, the defects of the null mutants could be restored by supplementing with exogenous cysteine or methionine. Our study explored the biological functions of sulfur assimilation and sulphur-containing amino acids biosynthesis in P. oryzae.






Similar content being viewed by others
References
Amich J, Schafferer L, Haas H, Krappmann S (2013) Regulation of sulphur assimilation is essential for virulence and affects iron homeostasis of the human-pathogenic mould Aspergillus fumigatus. PLoS Pathog 9:e1003573. https://doi.org/10.1371/journal.ppat.1003573
Ball RO, Courtney-Martin G, Pencharz PB (1693S) The in vivo sparing of methionine by cysteine in sulfur amino acid requirements in animal models and adult humans. J Nutr 136:1682S–1693S. https://doi.org/10.1093/jn/136.6.1682S
Fernandez J, Orth K (2018) Rise of a cereal killer: the biology of Magnaporthe oryzae biotrophic growth. Trends Microbiol. https://doi.org/10.1016/j.tim.2017.12.007
Fernandez J, Marroquin-Guzman M, Wilson RA (2014) Mechanisms of nutrient acquisition and utilization during fungal infections of leaves. Annu Rev Phytopathol 52:155–174. https://doi.org/10.1146/annurev-phyto-102313-050135
Gai Y, Liu B, Ma H, Li L, Chen X, Moenga S, Riely B, Fayyaz A, Wang M, Li H (2019) The methionine biosynthesis regulator AaMetR contributes to oxidative stress tolerance and virulence in Alternaria alternata. Microbiol Res 219:94–109. https://doi.org/10.1016/j.micres.2018.11.007
Howard RJ, Valent B (1996) Breaking and entering: host penetration by the fungal rice blast pathogen Magnaporthe grisea. Annu Rev Microbiol 50:491–512. https://doi.org/10.1146/annurev.micro.50.1.491
Howard RJ, Ferrari MA, Roach DH, Money NP (1991) Penetration of hard substrates by a fungus employing enormous turgor pressures. Proc Natl Acad Sci USA 88:11281–11284
Jain S, Sekonyela R, Knox BP, Palmer JM, Huttenlocher A, Kabbage M, Keller NP (2018) Selenate sensitivity of a laeA mutant is restored by overexpression of the bZIP protein MetR in Aspergillus fumigatus. Fungal Genet Biol 117:1–10. https://doi.org/10.1016/j.fgb.2018.05.001
Linder T (2017) ATP sulfurylase is essential for the utilization of sulfamate as a sulfur source in the yeast Komagataella pastoris (syn. Pichia pastoris). Curr Microbiol 74:1021–1025. https://doi.org/10.1007/s00284-017-1276-0
Liu XH, Chen SM, Gao HM, Ning GA, Shi HB, Wang Y, Dong B, Qi YY, Zhang DM, Lu GD, Wang ZH, Zhou J, Lin FC (2015) The small GTPase MoYpt7 is required for membrane fusion in autophagy and pathogenicity of Magnaporthe oryzae. Environ Microbiol 17:4495–4510. https://doi.org/10.1111/1462-2920.12903
Liu XH, Ning GA, Huang LY, Zhao YH, Dong B, Lu JP, Lin FC (2016) Calpains are involved in asexual and sexual development, cell wall integrity and pathogenicity of the rice blast fungus. Sci Rep 6:31204. https://doi.org/10.1038/srep31204
Lo SC, Hamer L, Hamer JE (2002) Molecular characterization of a cystathionine beta-synthase gene, CBS1, in Magnaporthe grisea. Eukaryot Cell 1:311–314
Lu JP, Liu XH, Feng XX, Min H, Lin FC (2009) An autophagy gene, MgATG5, is required for cell differentiation and pathogenesis in Magnaporthe oryzae. Curr Genet 55:461–473. https://doi.org/10.1007/s00294-009-0259-5
Lu J, Cao H, Zhang L, Huang P, Lin F (2014) Systematic analysis of Zn2Cys6 transcription factors required for development and pathogenicity by high-throughput gene knockout in the rice blast fungus. PLoS Pathog 10:e1004432. https://doi.org/10.1371/journal.ppat.1004432
Marzluf GA (1997) Molecular genetics of sulfur assimilation in filamentous fungi and yeast. Annu Rev Microbiol 51:73–96. https://doi.org/10.1146/annurev.micro.51.1.73
Pan Y, Pan R, Tan L, Zhang Z, Guo M (2019) Pleiotropic roles of O-mannosyltransferase MoPmt4 in development and pathogenicity of Magnaporthe oryzae. Curr Genet 65:223–239. https://doi.org/10.1007/s00294-018-0864-2
Que Y, Yue X, Yang N, Xu Z, Tang S, Wang C, Lv W, Xu L, Talbot NJ, Wang Z (2019) Leucine biosynthesis is required for infection-related morphogenesis and pathogenicity in the rice blast fungus Magnaporthe oryzae. Curr Genet. https://doi.org/10.1007/s00294-019-01009-2
Saint-Macary ME, Barbisan C, Gagey MJ, Frelin O, Beffa R, Lebrun MH, Droux M (2015) Methionine biosynthesis is essential for infection in the rice blast fungus Magnaporthe oryzae. PLoS ONE 10:e0111108. https://doi.org/10.1371/journal.pone.0111108
Schaible UE, Kaufmann SH (2005) A nutritive view on the host-pathogen interplay. Trends Microbiol 13:373–380. https://doi.org/10.1016/j.tim.2005.06.009
Shipman EN, Jones K, Jenkinson CB, Kim DW, Zhu J, Khang CH (2017) Nuclear and structural dynamics during the establishment of a specialized effector-secreting cell by Magnaporthe oryzae in living rice cells. BMC Cell Biol 18:11. https://doi.org/10.1186/s12860-017-0126-z
Sweigard JA, Carroll AM, Farrall L, Valent B (1997) A series of vectors for fungal transformation. Fungal Genet Rep. https://doi.org/10.4148/1941-4765.1287
Talbot NJ (2003) On the trail of a cereal killer: exploring the biology of Magnaporthe grisea. Annu Rev Microbiol 57:177–202. https://doi.org/10.1146/annurev.micro.57.030502.090957
Tang W, Ru Y, Hong L, Zhu Q, Zuo R, Guo X, Wang J, Zhang H, Zheng X, Wang P, Zhang Z (2014) System-wide characterization of bZIP transcription factor proteins involved in infection-related morphogenesis of Magnaporthe oryzae. Environ Microbiol. https://doi.org/10.1111/1462-2920.12618
Tang W, Jiang H, Zheng Q, Chen X, Wang R, Yang S, Zhao G, Liu J, Norvienyeku J, Wang Z (2019) Isopropylmalate isomerase MoLeu1 orchestrates leucine biosynthesis, fungal development, and pathogenicity in Magnaporthe oryzae. Appl Microbiol Biotechnol 103:327–337. https://doi.org/10.1007/s00253-018-9456-9
Toh EA, Ohkusu M, Shimizu K, Ishiwada N, Watanabe A, Kamei K (2018) Novel biosynthetic pathway for sulfur amino acids in Cryptococcus neoformans. Curr Genet 64:681–696. https://doi.org/10.1007/s00294-017-0783-7
Wilson RA, Fernandez J, Quispe CF, Gradnigo J, Seng A, Moriyama E, Wright JD (2012) Towards defining nutrient conditions encountered by the rice blast fungus during host infection. PLoS ONE 7:e47392. https://doi.org/10.1371/journal.pone.0047392
Yan X, Que Y, Wang H, Wang C, Li Y, Yue X, Ma Z, Talbot NJ, Wang Z (2013) The MET13 methylenetetrahydrofolate reductase gene is essential for infection-related morphogenesis in the rice blast fungus Magnaporthe oryzae. PLoS ONE 8:e76914. https://doi.org/10.1371/journal.pone.0076914
Zhang Y, Shi H, Liang S, Ning G, Xu N, Lu J, Liu X, Lin F (2015) MoARG1, MoARG5,6 and MoARG7 involved in arginine biosynthesis are essential for growth, conidiogenesis, sexual reproduction, and pathogenicity in Magnaporthe oryzae. Microbiol Res 180:11–22. https://doi.org/10.1016/j.micres.2015.07.002
Acknowledgements
This study was supported by grants from the Ministry of Agriculture of China (2016ZX08009003-001-005), National Science and Technology Major Project (2018ZX08001-03B), and the National Natural Science Foundation of China (No. 31770154).
Author information
Authors and Affiliations
Contributions
X-HL, BD, J-PL, and F-CL contributed to the study conception and design. Material preparation, data collection and analysis were performed by YL, MW, QY, Z-ZS, and Q-SL. The manuscript was written by YL, MW, BD, and X-HL. All authors read and approved the final manuscript.
Corresponding author
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
294_2020_1055_MOESM1_ESM.tif
Supplementary file1 Figure S1. Sequence alignment of PoMet3 and PoMet14. The amino acid alignment of the PoMet3 of Saccharomyces cerevisiae (NP_012543.3), Fusarium graminearum (XP_011319833.1), Eremothecium gossypii (NP_986988.1), Kluyveromyces lactis (XP_454393.1), Schizosaccharomyces pombe (NP_595662.2), and Neurospora crassa (XP_964349.1) was performed using ClustalW. The amino acid alignments of the PoMet14 are S. cerevisiae (NP_012925.3), F. graminearum (XP_011317116.1), K. lactis (XP_452124.1), S. pombe (NP_594718.1), N. crassa (XP_958621.1) (TIF 6530 kb)
294_2020_1055_MOESM3_ESM.tif
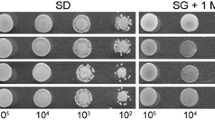
Supplementary file3 Figure S3. Growth of M. oryzae mutants on selected sulfur sources. Growth of the strains on MM supplemented with 1 mM selected sulfur sources for 8 days (TIF 8208 kb)
Rights and permissions
About this article
Cite this article
Li, Y., Wu, M., Yu, Q. et al. PoMet3 and PoMet14 associated with sulfate assimilation are essential for conidiogenesis and pathogenicity in Pyricularia oryzae. Curr Genet 66, 765–774 (2020). https://doi.org/10.1007/s00294-020-01055-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-020-01055-1