Abstract
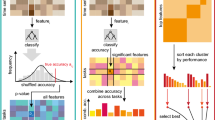
Experimental time series provide an informative window into the underlying dynamical system, and the timing of the extrema of a time series (or its derivative) contains information about its structure. However, the time series often contain significant measurement errors. We describe a method for characterizing a time series for any assumed level of measurement error \(\varepsilon \) by a sequence of intervals, each of which is guaranteed to contain an extremum for any function that \(\varepsilon \)-approximates the time series. Based on the merge tree of a continuous function, we define a new object called the normalized branch decomposition, which allows us to compute intervals for any level \(\varepsilon \). We show that there is a well-defined total order on these intervals for a single time series, and that it is naturally extended to a partial order across a collection of time series comprising a dataset. We use the order of the extracted intervals in two applications. First, the partial order describing a single dataset can be used to pattern match against switching model output (Cummins et al. in SIAM J Appl Dyn Syst 17(2):1589–1616, 2018), which allows the rejection of a network model. Second, the comparison between graph distances of the partial orders of different datasets can be used to quantify similarity between biological replicates.















Similar content being viewed by others
References
Akutsu T, Miyano S, Kuhara S (2000) Inferring qualitative relations in genetic networks and metabolic pathways. Bioinformatics 16(8):727–734. https://doi.org/10.1093/bioinformatics/16.8.727
Albert R (2007) Network inference, analysis, and modeling in systems biology. Plant Cell 19(11):3327–3338
Albert R, Collins JJ, Glass L (2013) Introduction to focus issue: quantitative approaches to genetic networks. Chaos 23(2):025001
Barker NA, Myers CJ, Kuwahara H (2011) Learning genetic regulatory network connectivity from time series data. IEEE/ACM Trans Comput Biol Bioinform 8(1):152–165. https://doi.org/10.1109/TCBB.2009.48
Bristow SL, Leman AR, Simmons Kovacs LA, Deckard A, Harer J, Haase SB (2014) Checkpoints couple transcription network oscillator dynamics to cell-cycle progression. Genome Biol 15(9):446. https://doi.org/10.1186/s13059-014-0446-7
Brunton SL, Proctor JL, Kutz JN (2016) Discovering governing equations from data by sparse identification of nonlinear dynamical systems. Proc Natl Acad Sci 113(15):3932–3937. https://doi.org/10.1073/pnas.1517384113
Bunke H, Riesen K (2011) Recent advances in graph-based pattern recognition with applications in document analysis. Pattern Recognit 44(5):1057–1067. https://doi.org/10.1016/j.patcog.2010.11.015
Carlsson G (2009) Topology and data. Bull Am Math Soc (NS) 46(2):255–308
Carré C, Mas A, Krouk G (2017) Reverse engineering highlights potential principles of large gene regulatory network design and learning. NPJ Syst Biol Appl. https://doi.org/10.1038/s41540-017-0019-y
Cho CY, Motta FC, Kelliher CM, Deckard A, Haase SB (2017) Reconciling conflicting models for global control of cell-cycle transcription. Cell Cycle. https://doi.org/10.1080/15384101.2017.1367073
Cho CY, Kelliher CM, Haase SB (2019) The cell-cycle transcriptional network generates and transmits a pulse of transcription once each cell cycle. Cell Cycle. https://doi.org/10.1080/15384101.2019.1570655
Conte D, Foggia P, Sansone C, Vento M (2004) Thirty years of graph matching in pattern recognition. Int J Pattern Recognit Artif Intell 18(03):265–298. https://doi.org/10.1142/S0218001404003228
Cummins B, Nerem R (2019) \(\epsilon \)-minimal interval software v0.4. https://doi.org/10.5281/zenodo.3405579; https://github.com/breecummins/min_interval_posets. Accessed Sept 2019
Cummins B, Gedeon T, Spendlove K (2015) On the efficacy of state space reconstruction methods in determining causality. SIAM J Appl Dyn Syst 14(1):335–381
Cummins B, Gedeon T, Harker S, Mischaikow K, Mok K (2016) Combinatorial representation of parameter space for switching systems. SIAM J Appl Dyn Syst 15(4):2176–2212
Cummins B, Gedeon T, Harker S, Mischaikow K (2018) Model rejection and parameter reduction via time series. SIAM J Appl Dyn Syst 17(2):1589–1616
Davey BA, Priestley HA (2002) Introduction to lattices and order. Cambridge University Press, Cambridge
Edelsbrunner H, Harer JL (2010) Computational topology. American Mathematical Society, Providence
Edwards R (2001) Chaos in neural and gene networks with hard switching. Differ Equ Dyn Syst 9:187–220
Fu JJ (1996) Approximate pattern matching in directed graphs. In: Hirschberg D, Myers G (eds) Combinatorial pattern matching, no. 1075 in Lecture Notes in Computer Science. Springer, Berlin, pp 373–383. https://doi.org/10.1007/3-540-61258-0_27
Glass L, Kauffman SA (1973) The logical analysis of continuous, non-linear biochemical control networks. J Theor Biol 39(1):103–29
Günther D, Salmon J, Tierny J (2014) Mandatory critical points of 2d uncertain scalar fields. In: Computer graphics forum, vol 33. Wiley Online Library, pp 31–40
Haase SB, Wittenberg C (2014) Topology and control of the cell-cycle-regulated transcriptional circuitry. Genetics 196(1):65–90. https://doi.org/10.1534/genetics.113.152595
Harker S (2018) DSGRN software. https://doi.org/10.5281/zenodo.1210003; https://github.com/shaunharker/DSGRN. Accessed June 2019
Kelliher CM, Leman AR, Sierra CS, Haase SB (2016) Investigating conservation of the cell-cycle-regulated transcriptional program in the fungal pathogen, cryptococcus neoformans. PLoS Genet 12(12):e1006453
Lähdesmäki H, Shmulevich I, Yli-Harja O (2003) On learning gene regulatory networks under the boolean network model. Mach Learn 52(1):147–167. https://doi.org/10.1023/A:1023905711304
Livi L, Rizzi A (2012) Parallel algorithms for tensor product-based inexact graph matching. In: The 2012 international joint conference on neural networks (IJCNN), pp 1–8. https://doi.org/10.1109/IJCNN.2012.6252681
Maucher M, Kracher B, Khl M, Kestler HA (2011) Inferring Boolean network structure via correlation. Bioinformatics 27(11):1529–1536. https://doi.org/10.1093/bioinformatics/btr166
McGoff K, Guo X, Deckard A, Kelliher C, Leman A, Francey L, Hogenesch J, Haase S, Harer J (2016) The Local Edge Machine: inference of dynamic models of gene regulation. Genome Biol 17(1):214
Morozov D, Weber G (2013) Distributed merge trees. In: Proceedings of the annual symposium on principles and practice of parallel programming, pp 93–102
Morozov D, Beketayev K, Weber G (2013) Interleaving distance between merge trees. Discrete Comput Geom 49:22–45
Mure LS, Le HD, Benegiamo G, Chang MW, Rios L, Jillani N, Ngotho M, Kariuki T, Dkhissi-Benyahya O, Cooper HM, Panda S (2018) Diurnal transcriptome atlas of a primate across major neural and peripheral tissues. Science. https://doi.org/10.1126/science.aao0318
Nerem R, Crawford-Kahrl P, Cummins B, Gedeon T (2019) A poset metric from the directed maximum common edge subgraph. arXiv:1910.14638
Orlando DA, Lin CY, Bernard A, Iversen ES, Hartemink AJ, Haase SB (2007) A probabilistic model for cell cycle distributions in synchrony experiments. Cell Cycle 6:478–488
Orlando DA, Lin CY, Bernard A, Wang JY, Socolar JE, Iversen ES, Hartemink AJ, Haase SB (2008) Global control of cell-cycle transcription by coupled cdk and network oscillators. Nature 453(7197):944–7
Pascucci V, Cole-Mclaughlin K, Scorzelli G (2004) Multi-resolution computation and presentation of contour trees. In: IASTED conference on visualization, imaging, and image processing
Pramila T, Wu W, Miles S, Noble WS, Breeden LL (2006) The forkhead transcription factor HCM1 regulates chromosome segregation genes and fills the S-phase gap in the transcriptional circuitry of the cell cycle. Genes Dev 20(16):2266–78
Rahi SJ, Pecani K, Ondracka A, Oikonomou C, Cross FR (2016) The CDK-APC/C oscillator predominantly entrains periodic cell-cycle transcription. Cell 165(2):475–87. https://doi.org/10.1016/j.cell.2016.02.060
Shedden K, Cooper S (2002) Analysis of cell-cycle gene expression in saccharomyces cerevisiae using microarrays and multiple synchronization methods. Nucleic Acids Res 30(13):2920–9
Simmons Kovacs LA, Orlando DA, Haase SB (2008) Transcription networks and cyclin/cdks: the yin and yang of cell cycle oscillators. Cell Cycle 7(17):2626–9
Simmons Kovacs LA, Mayhew MB, Orlando DA, Jin Y, Li Q, Huang C, Reed SI, Mukherjee S, Haase SB (2012) Cyclin-dependent kinases are regulators and effectors of oscillations driven by a transcription factor network. Mol Cell 45(5):669–79. https://doi.org/10.1016/j.molcel.2011.12.033
Simon I, Barnett J, Hannett N, Harbison CT, Rinaldi NJ, Volkert TL, Wyrick JJ, Zeitlinger J, Gifford DK, Jaakkola TS, Young RA (2001) Serial regulation of transcriptional regulators in the yeast cell cycle. Cell 106(6):697–708
Smirnov D, Morozov D (2017) Triplet merge trees. In: Topological methods in data analysis and visualization V (proceedings of TopoInVis 2017) (to appear)
Sugihara G, May R, Ye H, Hsieh C, Deyle E, Fogarty M, Munch S (2016) Detecting causality in complex ecosystems. Science 338:496
Thomas R (1991) Regulatory networks seen as asynchronous automata: a logical description. J Theor Biol 153:1–23. https://doi.org/10.1016/S0022-5193(05)80350-9
Zhang R, Lahens NF, Ballance HI, Hughes ME, Hogenesch JB (2014) A circadian gene expression atlas in mammals: implications for biology and medicine. Proc Natl Acad Sci U S A 111(45):16219–24. https://doi.org/10.1073/pnas.1408886111
Zomorodian A, Carlsson G (2005) Computing persistent homology. Discrete Comput Geom 33(2):249–274
Acknowledgements
T. G. was partially supported by NSF Grant DMS-1361240, USDA 2015-51106-23970, DARPA Grant FA8750-17-C-0054, NIH Grant 1R01GM126555-01, and NSF TRIPODS+X Grant 1839299. B.C. was partially supported by Grants USDA 2015-51106-23970, DARPA Grant FA8750-17-C-0054, NIH 1R01GM126555-01, and NSF TRIPODS+X Grant 1839299. E. B. was partially supported by NSF Grants DMS-1508040, DMS-1664858, DMS-1557716, DMS-1812055 and DMS-1945639. R. N. was supported by Montana State University’s Undergraduate Scholars Program (USP) during the Fall 2018 funding cycle. S.H. was partially supported by NIH 5 R01 GM126555-03 and DARPA FA8750-17-C-0054. L.S. was supported by NIH 5 R01 GM126555-03. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the National Science Foundation. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Yeast data analysis
Yeast data analysis
Time-series transcriptomic data for one replicate of wild-type yeast Saccharomyes cerevisiae were previously published in Orlando et al. (2008). Microarray (Affymetrix Yeast Genome 2.0) expression data were normalized as previously described (Orlando et al. 2008), although for this study Affymetrix probe IDs were re-annotated using Affymetrix Yeast Genome 2.0 microarray annotation 35. Expression data were aligned to a common cell-cycle time line using the CLOCCS (Characterizing Loss of Cell Cycle Synchrony) (Orlando et al. 2007) population synchrony model, as previously described (Orlando et al. 2008). Briefly, the CLOCCS model allows multiple time-series experiments to be aligned to a common cell-cycle timeline, using experimentally-derived yeast budding data. The CLOCCS model converts time points in the series to life points, which indicate the progression through the cell cycle. Expression data for both replicates were interpolated to integer life points with an interval of one using a Piecewise Cubic Hermite Interpolating Polynomial (PCHIP) spline. Life points were then trimmed so both replicate time series were of identical location and duration in the cell cycle.
Rights and permissions
About this article
Cite this article
Berry, E., Cummins, B., Nerem, R.R. et al. Using extremal events to characterize noisy time series. J. Math. Biol. 80, 1523–1557 (2020). https://doi.org/10.1007/s00285-020-01471-4
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00285-020-01471-4