Abstract
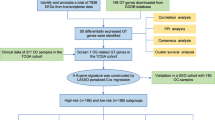
The complex enzyme network responsible for glycan synthesis suffers significant changes during the first steps of tumor development, leading to the early formation of tumor-associated glycan signatures. Among the glycosylation pathways, changes in fucosylation emerged as one of most important features in cancer. Αlpha-1,3/4-fucosyltransferase (FUT3) has been linked to pro-tumor and anti-tumor pathways depending on the cancer type. The present study aimed to understand the gene and protein expression profiles of FUT3 in three different and independent cohorts composed by invasive breast cancer patients: Local Brazilian population, METABRIC and TCGA. FUT3 transcripts and protein were measured in the Brazilian population by real-time PCR and Western blotting, respectively. Clinical records and FUT3 levels from public METABRIC and TCGA cohorts were accessed through CBioPortal database. FUT3 expression was analyzed in each cohort using the appropriated statistic tools. Survival meta-analysis in triple negative patients was performed using five independent cohorts including GSE41119, GSE47994 and GSE86945, data obtained from GEO repository available at NCBI database, and METABRIC and TCGA. Our analysis showed that high FUT3 levels were consistently associated to reduced invasive breast cancer patients overall survival. This finding is particularly significant in triple negative patients. These results together with the previously knowledge regarding the involvement of FUT3 in pro-tumor and anti-tumor mechanisms led us to purpose a model for FUT3 expression regulation throughout breast cancer establishment and progression.



Similar content being viewed by others
References
Bray, F., Ferlay, J., Soerjomataram, I., Siegel, R.L., Torre, L.A., Jemal, A.: Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68(6), 394–424 (2018)
Ferlay, J., Soerjomataram, I., Dikshit, R., Eser, S., Mathers, C., Rebelo, M., Parkin, D.M., Forman, D., Bray, F.: Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer. 136(5), E359–E386 (2015)
Colomer, R., Aranda-López, I., Albanell, J., García-Caballero, T., Ciruelos, E., López-García, M.Á., Cortés, J., Rojo, F., Martín, M., Palacios-Calvo, J.: Biomarkers in breast cancer: a consensus statement by the Spanish Society of Medical Oncology and the Spanish Society of Pathology. Clin. Transl. Oncol. 20(7), 815–826 (2018)
Yadav, B.S., Chanana, P., Jhamb, S.: Biomarkers in triple negative breast cancer: a review. World J. Clin. Oncol. 6(6), 252–263 (2015)
Dube, D.H., Bertozzi, C.R.: Glycans in cancer and inflammation-potential for therapeutics and diagnostics. Nat. Rev. Drug Discov. 4(6), 477–488 (2005)
Pinho, S.S., Reis, A.R.: Glycosylation in cancer: mechanisms and clinical implications. Nat. Rev. Cancer. 15(9), 540–555 (2015)
Colomb, F., Robbe-Masselot, C., Groux-Degroote, S., Bouckaert, J., Delannoy, P., Michalski, J.: Epithelial mucins and bacterial adhesion. J. Carbohyd. Chem. 40, 596–623 (2014)
Jeschke, U., Mylonas, I., Shabani, N., Kunert-Keil, C., Schindlbeck, C., Gerber, B., Friese, K.: Expression of Sialyl Lewis X. Sialyl Lewis A, E-Cadherin and Cathepsin-D in Human Breast Cancer: Immunohistochemical Analysis in Mammary Carcinoma In Situ. Anticancer Res. 25(3A), 1615–1622 (2005)
Czerwinski, M.J., Desiderio, V., Shkeir, O., Papagerakis, P., Lapadatescu, M.C., Owen, J.H., Athanassiou-Papaefthymiou, M., Zheng, L., Papaccio, G., Prince, M.E., Papagerakis, S.: In vitro evaluation of sialyl Lewis X relationship with head and neck cancer stem cells. Otolaryngol. Head Neck Surg. 149(1), 97–104 (2013)
Borzym-Kluczyk, M., Radziejewska, I., Cechowska-Pasko, M.: Increased expression of MUC1 and sialyl Lewis antigensin different areas of clear renal cell carcinoma. Clin. Exp. Nephrol. 19(4), 732–737 (2015)
Burdick, M.M., Henson, K.A., Delgadillo, L.F., Choi, Y.E., Goetz, D.J., Tees, D.F., Benencia, F.: Expression of E-selectin ligands on circulating tumor cells: cross-regulation with cancer stem cell regulatory pathways? Front. Oncol. 2, 103 (2012)
Hirakawa, M., Takimoto, R., Tamura, F., Yoshida, M., Ono, M., Murase, K., Sato, Y., Osuga, T., Sato, T., Iyama, S., Miyanishi, K., Takada, K., Hayashi, T., Kobune, M., Kato, J.: Fucosylated TGF-β receptors transduces a signal for epithelial-mesenchymal transition in colorectal cancer cells. Br. J. Cancer. 110(1), 156–163 (2014)
Higai, K., Ichikawa, A., Matsumoto, K.: Binding of sialyl Lewis X antigen to lectin-like receptors on NK cells induces cytotoxicity and tyrosine phosphorylation of a 17-kDa protein. Biochim. Biophys. Acta. 1760(9), 1355–1363 (2006)
Zhang, B., Van Roosmalen, I.A.M., Reis, C.R., Setroikromo, R., Quax, W.J.: Death receptor 5 is activated by fucosylation in colon cancer cells. FEBS J. 286(3), 555–571 (2019)
Vasconcelos, J.L.A., Ferreira, S.A., Lima, A.L., Rêgo, M.J.M., Bandeira, A.R., Cavalcanti, C.L.B., Lira, M.M.M., Beltrão, E.I.: Comparing the Immunoexpression of FUT3 and FUT6 between prostatic adenocarcinoma and benign prostatic hyperplasia. Acta Histochem. Cytochem. 46(3), 105–109 (2013)
Vajaria, B.N., Patel, K.R., Begum, R., Shah, F.D., Patel, J.B., Joshi, G.M., Patel, P.S.: Expression of Glycosyltransferases; ST3GAL1, FUT3, FUT5, and FUT6b transcripts in Oral Cancer. Glycobiol. Insights. 4, 7–14 (2014)
Nascimento, J.C.F., Ferreira, S.A., Vasconcelos, J.L.A., Silva-Filho, J.L., Barbosa, B.T., Bezerra, M.F., Rocha, C.R., Beltrão, E.I.: Fut3 role in breast invasive ductal carcinoma: investigating its gene promoter and protein expression. Exp. Mol. Pathol. 99(3), 409–415 (2015)
Meng, L., Xu, L., Yang, Y., Zhou, L., Chang, Y., Shi, T., Tan, C., An, H., Zhu, Y., Xu, J.: High expression of FUT3 is linked to poor prognosis in clear cell renal cell carcinoma. Oncotarget. 8(37), 61036–61047 (2017)
Aubert, M., Panicot-Dubois, L., Crotte, C., Sbarra, V., Lombardo, D., Sadoulet, M.O., Mas, E.: Peritoneal colonization by human pancreatic cancer cells is inhibited by antisense FUT3 sequence. Int. J. Cancer. 88(4), 558–565 (2000)
Barthel, S.R., Wiese, G.K., Cho, J., Opperman, M.J., Hays, D.L., Siddiqui, J., Pienta, K.J., Furie, B., Dimitroff, C.J.: Alpha 1,3 fucosyltransferases are master regulators of prostate cancer cell trafficking. Proc. Natl. Acad. Sci. U. S. A. 106(46), 19491–19496 (2009)
Carrascal, M.A., Silva, M., Ramalho, J.S., Pen, C., Martins, M., Pascoal, C., Amaral, C., Serrano, I., Oliveira, M.J., Sackstein, R., Videira, P.A.: Inhibition of fucosylation in human invasive ductal carcinoma reduces E-selectin ligand expression, cell proliferation and ERK1/2 and p38 MAPK activation. Mol. Oncol. 12(5), 579–593 (2018)
Ma, H., Miao, X., Ma, Q., Zheng, W., Zhou, H., Jia, L.: Functional roles of glycogene and N-glycan in multidrug resistance of human breast cancer cells. IUBMB Life. 65(5), 409–422 (2013)
Breiman, A., Robles, M.D.L., Trécesson, S.C., Echasserieau, K., Bernardeau, K., Drickamer, K., Imberty, A., Barillé-Nion, S., Altare, F., Le Pendu, J.: Carcinoma-associated fucosylated antigens are markers of the epithelial state and can contribute to cell adhesion through CLEC17A (Prolectin). Oncotarget. 7(12), 14064–14082 (2016)
Julien, S., Ivetic, A., Grigoriadis, A., QiZe, D., Burford, B., Sproviero, D., Picco, G., Gillett, C., Papp, S.L., Schaffer, L., Tutt, A., Taylor-Papadimitriou, J., Pinder, S.E., Burchell, J.M.: Selectin ligand sialyl-Lewis x antigen drives metastasis of hormone-dependent breast cancers. Cancer Res. 71(24), 7683–7693 (2011)
Liu, L.L., Zhao, H., Ma, T.F., Ge, F., Chen, C.S., Zhang, Y.P.: Identification of valid reference genes for the normalization of RT-qPCR expression studies in human breast Cancer cell lines treated with and without transient transfection. PLoS One. 10(1), e0117058 (2015)
Nascimento, J.C.F., Vasconcelos, A.O., Seabra, M.A.B.L., Beltrão, E.I.C., Rocha, C.R.C.: The challenge of determining the impact of FUT3 tumor-associated polymorphism rs2306969 (−6951 C> T) in invasive breast cancer cells. Mol. Biol. Rep. 46(3), 3531–3536 (2019)
Pereira, B., Chin, S.F., Rueda, O.M., Vollan, H.K., Provenzano, E., Bardwell, H.A., Pugh, M., Jones, L., Russell, R., Sammut, S.J., Tsui, D.W., Liu, B., Dawson, S.J., Abraham, J., Northen, H., Peden, J.F., Mukherjee, A., Turashvili, G., Green, A.R., McKinney, S., Oloumi, A., Shah, S., Rosenfeld, N., Murphy, L., Bentley, D.R., Ellis, I.O., Purushotham, A., Pinder, S.E., Børresen-Dale, A.L., Earl, H.M., Pharoah, P.D., Ross, M.T., Aparicio, S., Caldas, C.: The somatic mutation profiles of 2,433 breast cancers refine their genomic and transcriptomic landscapes. Nat. Commun. 7, 11479 (2016)
Ciriello, G., Gatza, M.L., Beck, A.H., Wilkerson, M.D., Rhie, S.K., Pastore, A., Zhang, H., McLellan, M., Yau, C., Kandoth, C., Bowlby, R., Shen, H., Hayat, S., Fieldhouse, R., Lester, S.C., Tse, G.M., Factor, R.E., Collins, L.C., Allison, K.H., Chen, Y.Y., Jensen, K., Johnson, N.B., Oesterreich, S., Mills, G.B., Cherniack, A.D., Robertson, G., Benz, C., Sander, C., Laird, P.W., Hoadley, K.A., King, T.A.: Comprehensive molecular portraits of invasive lobular breast cancer. Cell. 163(2), 506–519 (2015)
Prat, A., Adamo, B., Cheang, M.C.U., Anders, C.K., Carey, L.A., Perou, C.M.: Molecular characterization of basal-like and non-basal-like triple-negative breast Cancer. Oncologist. 18, 123–133 (2013)
Romero-Cordoba, S.L., Rodriguez-Cuevas, S., Bautista-Pina, V., Maffuz-Aziz, A., D'Ippolito, E., Cosentino, G., Baroni, S., Iorio, M.V., Hidalgo-Miranda, A.: Loss of function of miR-342-3p results in MCT1 over-expression and contributes to oncogenic metabolic reprogramming in triple negative breast cancer. Sci. Rep. 8, 12252 (2018)
Camp, R.L., Dolled-Filhart, M., Rimm, D.L.: X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin. Can. Res. 10, 7252–7259 (2004)
Guo, Y., Sheng, Q., Li, J., Ye, F., Samuels, D.C., Shyr, Y.: Large scale comparison of gene expression levels by microarrays and RNAseq using TCGA data. PLoS One. 8(8), e71462 (2013)
Duffy, M.J., Sturgeon, C.M., Sölétormos, G., Barak, V., Molina, R., Hayes, D.F., Diamandis, E.P., Bossuyt, P.M.: Validation of new cancer biomarkers: a position statement from the European group on tumor markers. Clin. Chem. 61(6), 809–820 (2015)
Xu, Z., Marko, N.F., Angelov, L., Barnett, G.H., Chao, S.T., Vogelbaum, M.A., Suh, J.H., Weil, R.J.: Impact of preexisting tumor necrosis on the efficacy of stereotactic radiosurgery in the treatment of brain metastases in women with breast cancer. Cancer. 118(5), 1323–1333 (2012)
Fujii, T., Yajima, R., Hirakata, T., Miyamoto, T., Fujisawa, T., Tsutsumi, S., Ynagita, Y., Iijima, M., Kuwano, H.: Impact of the prognostic value of vascular invasion, but not lymphatic invasion, of the primary tumor in patients with breast cancer. Anticancer Res. 34(3), 1255–1259 (2014)
Tang, C., Wang, P., Li, X., Zhao, B., Yang, H., Yu, H., Li, C.: Lymph node status have a prognostic impact in breast cancer patients with distant metastasis. PLoS One. 12(8), e0182953 (2017)
Li, Y., Steppi, A., Zhou, Y., Mao, F., Miller, P.C., He, M.M., Zhao, T., Sun, Q., Zhang, J.: Tumoral expression of drug and xenobiotic metabolizing enzymes in breast cancer patients of different ethnicities with implications to personalized medicine. Sci. Rep. 7(1), 4747 (2017)
Salzano, F.M., Sans, M.: Interethnic admixture and the evolution of Latin American populations. Genet. Mol. Biol. 37(1), 151–170 (2014)
Favé, M., Lamaze, F.C., Soave, D., Hodgkinson, A., Gauvin, H., Bruat, V., Grenier, J., Gbeha, E., Skead, K., Smargiassi, A., Johnson, M., Idaghdour, Y., Awadalla, P.: Gene-by-environment interactions in urban populations modulate risk phenotypes. Nat. Commun. 9(1), 827 (2018)
Duarte, H.O., Balmaña, M., Mereiter, S., Osório, H., Gomes, J., Reis, C.A.: Gastric Cancer cell glycosylation as a modulator of the ErbB2 oncogenic receptor. Int. J. Mol. Sci. 18(11), E2262 (2017)
Castellano, L., Giamas, G., Jacob, J., Coombes, R.C., Lucchesi, W., Thiruchelvam, P., Barton, G., Jiao, L.R., Wait, R., Waxman, J., Hannon, G.J., Stebbing, J.: The estrogen receptor-alpha-induced microRNA signature regulates itself and its transcriptional response. Proc. Natl. Acad. Sci. U. S. A. 106(37), 15732–15737 (2009)
Zheng, Y., Shao, X., Huang, Y., Shi, L., Chen, B., Wang, X., Yang, H., Chen, Z., Zhang, X.: Role of estrogen receptor in breast cancer cell gene expression. Mol. Med. Rep. 13(5), 4046–4050 (2016)
Rahmatpanah, F.B., Jia, Z., Chen, X., Char, J.E., Men, B., Franke, A.C., Jones, F.E., McClelland, M., Mercola, D.: A class of genes in the HER2 regulon that is poised for transcription in breast cancer cell lines and expressed in human breast tumors. Oncotarget. 6(2), 1286–1301 (2015)
Gnant, M., Harbeck, N., Thomssen, C.: St. Gallen 2011: Summary of the Consensus Discussion. Breast Care. 6, 136–141 (2011)
Balic, M., Thomssen, C., Würstlein, R., Gnant, M., Harbeck, N.: St. Gallen/Vienna 2019: A Brief Summary of the Consensus Discussion on the Optimal Primary Breast Cancer Treatment. Breast Care. 14, 103–110 (2019)
Park, Y.H., Lee, S.J., Cho, E.Y., Choi, Y.L., Lee, J.E., Nam, S.J., Yang, J.H., Shin, J.H., Ko, E.Y., Han, B.K., Ahn, S.J., Im, Y.H.: Clinical relevance of TNM staging system according tobreast cancer subtypes. Ann. Oncol. 22, 1554–1560 (2011)
Gonçalves Junior, H., Guerra, M.R., Cintra, J.R.D., Fayer, V.A., Brum, I.V., Teixeira, A.T.B.: Survival study of triple-negative and non–triple-negative breast Cancer in a Brazilian cohort. Clin. Med. Insights Oncol. 12, 1179554918790563 (2018)
Milde-Langosch, K., Karn, T., Schmidt, M., Zu Eulenburg, C., Oliveira-Ferrer, L., Wirtz, R.M., Schumacher, U., Witzel, I., Schütze, D., Müller, V.: Prognostic relevance of glycosylation-associated genes in breast cancer. Breast Cancer Res. Treat. 145(2), 295–305 (2014)
Cheng, T., Tu, S., Chen, L., Chen, M.Y., Chen, W.Y., Lin, Y.K., Ho, C.T., Lin, S.Y., Wu, C.H., Ho, Y.S.: Down-regulation of α-L-fucosidase 1 expression confers inferior survival for triple-negative breast cancer patients by modulating the glycosylation status of the tumor cell surface. Oncotarget. 6(25), 21283–21300 (2015)
Bonin, S., Parascandolo, A., Aversa, C., Barbazza, R., Tsuchida, N., Castellone, M.D., Stanta, G., Vecchio, G.: Reduced expression of α-L-Fucosidase-1 (FUCA-1) predicts recurrence and shorter cancer specific survival in luminal B LN+ breast cancer patients. Oncotarget. 9(20), 15228–15238 (2018)
Esposito, M., Mondal, N., Greco, T.M., Wei, Y., Spadazzi, C., Lin, S.C., Zheng, H., Cheung, C., Magnani, J.L., Lin, S.H., Cristea, I.M., Sackstein, R., Kang, Y.: Bone vascular niche E-selectin induces mesenchymal–epithelial transition and Wnt activation in cancer cells to promote bone metastasis. Nat. Cell Biol. 21, 627–639 (2019)
Scully, O.J., Bay, B.H., Yip, G., Yu, Y.: Breast Cancer Metastasis. Cancer Genomics Proteomics. 9(5), 311–320 (2012)
Acknowledgments
We gratefully acknowledge Dr. George Siqueira and all the clinical and technical team from Hospital das Clínicas da Universidade Federal de Pernambuco (HC-UFPE, Brazil) for the support during the volunteers’ screening and sample collection. We also thank Professor Dayane Gomes from Universidade Federal de Pernambuco (Brazil) for the support during the execution of the Western blotting platform assays and Dr. Sandra L. Romero Cordoba from Instituto Nacional de Ciencias Médicas y Nutrición Salvador Zubiran (Mexico) to provide us the clinical data from GSE86945 cohort.
Funding
This study was funded by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) [grant number 480291/2011–0]; and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Jessica Catarine Frutuoso do Nascimento. The first draft of the manuscript was written by Jessica Catarine Frutuoso do Nascimento and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in this study involving human participants were in accordance with the ethical standards of the institutional research committee (Comité de Ética em Pesquisa Envolvendo Seres Humanos do Centro de Ciências da Saúde da Universidade Federal de Pernambuco, CAAE 06586612.9.0000.5208 — Num. 140.876) and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
do Nascimento, J.C.F., Beltrão, E.I.C. & Rocha, C.R.C. High FUT3 expression is a marker of lower overall survival of breast cancer patients. Glycoconj J 37, 263–275 (2020). https://doi.org/10.1007/s10719-020-09914-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10719-020-09914-2