Abstract
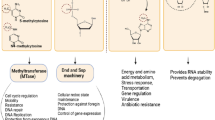
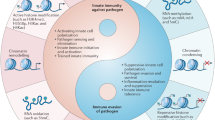
In the long co-evolution of host-pathogen interaction, bacteria have developed sophisticated strategies to manipulate host cell mechanisms and reprogram host transcription. Targeting chromatin, mainly through post-translational modification (PTM) of histone proteins, is one strategy that has been revealed over the last decade. Indeed, histone modifications play a crucial role in regulating transcription during cell type and stimulus specific responses, making them good targets during infection. Therefore, the study of host-pathogen interactions provides breakthroughs in understanding virulence mechanisms, but also in host cell mechanisms. Although chromatin is regulated by DNA methylation, noncoding RNAs, and post-translational modifications of histones, most studies have concentrated on bacteria-induced histone modifications, which will be the focus of this review. We will discuss the different mechanisms used by bacteria to induce histone PTMs, whether it is through direct targeting of pathogen effector enzymes, or indirectly through modulation of cellular signaling cascade. We will summarize the concepts we learned in cell biology from exploring bacteria-triggered histone modifications, by focusing on the signaling cascades modified by bacteria, bacterial mimics of eukaryotic enzymes, and the novel histone marks imposed upon infection.




Similar content being viewed by others
References
Zhou K, Gaullier G, Luger K (2019) Nucleosome structure and dynamics are coming of age. Nat Struct Mol Biol 26(1):3–13
Bednar J, Horowitz RA, Grigoryev SA, Carruthers LM, Hansen JC, Koster AJ, Woodcock CL (1998) Nucleosomes, linker DNA, and linker histone form a unique structural motif that directs the higher-order folding and compaction of chromatin. Proc Natl Acad Sci U S A 95(24):14173–14178
Narlikar GJ, Sundaramoorthy R, Owen-Hughes T (2013) Mechanisms and functions of ATP-dependent chromatin-remodeling enzymes. Cell 154(3):490–503
Li E (2002) Chromatin modification and epigenetic reprogramming in mammalian development. Nat Rev Genet 3(9):662–673
Tan M, Luo H, Lee S, Jin F, Yang JS, Montellier E, Buchou T, Cheng Z, Rousseaux S, Rajagopal N, Lu Z, Ye Z, Zhu Q, Wysocka J, Ye Y, Khochbin S, Ren B, Zhao Y (2011) Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell 146(6):1016–1028
Strahl BD, Allis CD (2000) The language of covalent histone modifications. Nature 403(6765):41–45
Kouzarides T (2007) Chromatin modifications and their function. Cell 128(4):693–705
Musselman CA, Lalonde ME, Cote J, Kutateladze TG (2012) Perceiving the epigenetic landscape through histone readers. Nat Struct Mol Biol 19(12):1218–1227
Agalioti T, Chen G, Thanos D (2002) Deciphering the transcriptional histone acetylation code for a human gene. Cell 111(3):381–392
Lauberth SM, Nakayama T, Wu X, Ferris AL, Tang Z, Hughes SH, Roeder RG (2013) H3K4me3 interactions with TAF3 regulate preinitiation complex assembly and selective gene activation. Cell 152(5):1021–1036
Lindroth AM, Shultis D, Jasencakova Z, Fuchs J, Johnson L, Schubert D, Patnaik D, Pradhan S, Goodrich J, Schubert I, Jenuwein T, Khorasanizadeh S, Jacobsen SE (2004) Dual histone H3 methylation marks at lysines 9 and 27 required for interaction with CHROMOMETHYLASE3. EMBO J 23(21):4146–4155
Schubert D, Primavesi L, Bishopp A, Roberts G, Doonan J, Jenuwein T, Goodrich J (2006) Silencing by plant Polycomb-group genes requires dispersed trimethylation of histone H3 at lysine 27. EMBO J 25(19):4638–4649
Creyghton MP, Cheng AW, Welstead GG, Kooistra T, Carey BW, Steine EJ, Hanna J, Lodato MA, Frampton GM, Sharp PA, Boyer LA, Young RA, Jaenisch R (2010) Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc Natl Acad Sci U S A 107(50):21931–21936
Henikoff S, Shilatifard A (2011) Histone modification: cause or cog? Trends in genetics : TIG 27(10):389–396
Gates LA, Foulds CE, O’Malley BW (2017) Histone marks in the ‘driver’s seat’: functional roles in steering the transcription cycle. Trends Biochem Sci 42(12):977–989
Koul A, Herget T, Klebl B, Ullrich A (2004) Interplay between mycobacteria and host signalling pathways. Nat Rev Microbiol 2(3):189–202
Haraga A, Ohlson MB, Miller SI (2008) Salmonellae interplay with host cells. Nat Rev Microbiol 6(1):53–66
Lebeer S, Vanderleyden J, De Keersmaecker SC (2010) Host interactions of probiotic bacterial surface molecules: comparison with commensals and pathogens. Nat Rev Microbiol 8(3):171–184
Bierne H, Hamon M, Cossart P (2012) Epigenetics and bacterial infections. Cold Spring Harbor perspectives in medicine 2(12):a010272
Eberharter A, Becker PB (2002) Histone acetylation: a switch between repressive and permissive chromatin-second in review series on chromatin dynamics. EMBO Rep 3(3):224–229
Filippakopoulos P, Picaud S, Mangos M, Keates T, Lambert J-P, Barsyte-Lovejoy D, Felletar I, Volkmer R, Müller S, Pawson T (2012) Histone recognition and large-scale structural analysis of the human bromodomain family. Cell 149(1):214–231
Cheung P, Tanner KG, Cheung WL, Sassone-Corsi P, Denu JM, Allis CD (2000) Synergistic coupling of histone H3 phosphorylation and acetylation in response to epidermal growth factor stimulation. Mol Cell 5(6):905–915
Lo WS, Trievel RC, Rojas JR, Duggan L, Hsu JY, Allis CD, Marmorstein R, Berger SL (2000) Phosphorylation of serine 10 in histone H3 is functionally linked in vitro and in vivo to Gcn5-mediated acetylation at lysine 14. Mol Cell 5(6):917–926
Josefowicz SZ, Shimada M, Armache A, Li CH, Miller RM, Lin S, Yang A, Dill BD, Molina H, Park HS, Garcia BA, Taunton J, Roeder RG, Allis CD (2016) Chromatin kinases act on transcription factors and histone tails in regulation of inducible transcription. Mol Cell 64(2):347–361
Yamamoto Y, Verma UN, Prajapati S, Kwak YT, Gaynor RB (2003) Histone H3 phosphorylation by IKK-alpha is critical for cytokine-induced gene expression. Nature 423(6940):655–659
Dyson MH, Thomson S, Inagaki M, Goto H, Arthur SJ, Nightingale K, Iborra FJ, Mahadevan LC (2005) MAP kinase-mediated phosphorylation of distinct pools of histone H3 at S10 or S28 via mitogen- and stress-activated kinase 1/2. J Cell Sci 118(Pt 10):2247–2259
Tiwari VK, Stadler MB, Wirbelauer C, Paro R, Schubeler D, Beisel C (2011) A chromatin-modifying function of JNK during stem cell differentiation. Nat Genet 44(1):94–100
Sassone-Corsi P, Mizzen CA, Cheung P, Crosio C, Monaco L, Jacquot S, Hanauer A, Allis CD (1999) Requirement of Rsk-2 for epidermal growth factor-activated phosphorylation of histone H3. Science 285(5429):886–891
Akira S, Uematsu S, Takeuchi O (2006) Pathogen recognition and innate immunity. Cell 124(4):783–801
Arthur JS, Ley SC (2013) Mitogen-activated protein kinases in innate immunity. Nat Rev Immunol 13(9):679–692
Anest V, Hanson JL, Cogswell PC, Steinbrecher KA, Strahl BD, Baldwin AS (2003) A nucleosomal function for IkappaB kinase-alpha in NF-kappaB-dependent gene expression. Nature 423(6940):659–663
L.C. Poulsen, R.J. Edelmann, S. Kruger, R. Dieguez-Hurtado, A. Shah, T.E. Stav-Noraas, A. Renzi, M. Szymanska, J. Wang, M. Ehling, R. Benedito, M. Kasprzycka, E. Baekkevold, O. Sundnes, K.S. Midwood, H. Scott, P. Collas, C.W. Siebel, R.H. Adams, G. Haraldsen, E. Sundlisaeter, J. Hol. Inhibition of endothelial NOTCH1 signaling attenuates inflammation by reducing cytokine-mediated histone acetylation at inflammatory enhancers, Arteriosclerosis, thrombosis, and vascular biology 38(4) (2018) 854–869
Weinmann AS, Mitchell DM, Sanjabi S, Bradley MN, Hoffmann A, Liou H-C, Smale ST (2001) Nucleosome remodeling at the IL-12 p40 promoter is a TLR-dependent, Rel-independent event. Nat Immunol 2(1):51–57
Rigillo G, Vilella A, Benatti C, Schaeffer L, Brunello N, Blom JMC, Zoli M, Tascedda F (2018) LPS-induced histone H3 phospho (Ser10)-acetylation (Lys14) regulates neuronal and microglial neuroinflammatory response. Brain Behav Immun 74:277–290
Saccani S, Pantano S, Natoli G (2002) p38-Dependent marking of inflammatory genes for increased NF-kappa B recruitment. Nat Immunol 3(1):69–75
Iglesias MJ, Reilly SJ, Emanuelsson O, Sennblad B, Pirmoradian Najafabadi M, Folkersen L, Malarstig A, Lagergren J, Eriksson P, Hamsten A, Odeberg J (2012) Combined chromatin and expression analysis reveals specific regulatory mechanisms within cytokine genes in the macrophage early immune response. PLoS One 7(2):e32306
Zippo A, Serafini R, Rocchigiani M, Pennacchini S, Krepelova A, Oliviero S (2009) Histone crosstalk between H3S10ph and H4K16ac generates a histone code that mediates transcription elongation. Cell 138(6):1122–1136
Schmeck B, Beermann W, van Laak V, Zahlten J, Opitz B, Witzenrath M, Hocke AC, Chakraborty T, Kracht M, Rosseau S, Suttorp N, Hippenstiel S (2005) Intracellular bacteria differentially regulated endothelial cytokine release by MAPK-dependent histone modification. J Immunol 175(5):2843–2850
B. Schmeck, J. Lorenz, D. N’Guessan P, B. Opitz, V. van Laak, J. Zahlten, H. Slevogt, M. Witzenrath, A. Flieger, N. Suttorp, S. Hippenstiel. Histone acetylation and flagellin are essential for Legionella pneumophila-induced cytokine expression, Journal of immunology 181(2) (2008) 940–7
Raymond B, Batsche E, Boutillon F, Wu YZ, Leduc D, Balloy V, Raoust E, Muchardt C, Goossens PL, Touqui L (2009) Anthrax lethal toxin impairs IL-8 expression in epithelial cells through inhibition of histone H3 modification. PLoS Pathog 5(4):e1000359
Bardwell AJ, Abdollahi M, Bardwell L (2004) Anthrax lethal factor-cleavage products of MAPK (mitogen-activated protein kinase) kinases exhibit reduced binding to their cognate MAPKs. The Biochemical journal 378(Pt 2):569–577
Gouin E, Adib-Conquy M, Balestrino D, Nahori MA, Villiers V, Colland F, Dramsi S, Dussurget O, Cossart P (2010) The Listeria monocytogenes InlC protein interferes with innate immune responses by targeting the I{kappa} B kinase subunit IKK{alpha}. Proc Natl Acad Sci U S A 107(40):17333–17338
Orth K, Palmer LE, Bao ZQ, Stewart S, Rudolph AE, Bliska JB, Dixon JE (1999) Inhibition of the mitogen-activated protein kinase kinase superfamily by a Yersinia effector. Science 285(5435):1920–1923
Mukherjee S, Keitany G, Li Y, Wang Y, Ball HL, Goldsmith EJ, Orth K (2006) Yersinia YopJ acetylates and inhibits kinase activation by blocking phosphorylation. Science 312(5777):1211–1214
Paquette N, Conlon J, Sweet C, Rus F, Wilson L, Pereira A, Rosadini CV, Goutagny N, Weber AN, Lane WS, Shaffer SA, Maniatis S, Fitzgerald KA, Stuart L, Silverman N (2012) Serine/threonine acetylation of TGFbeta-activated kinase (TAK1) by Yersinia pestis YopJ inhibits innate immune signaling. Proc Natl Acad Sci U S A 109(31):12710–12715
Jones RM, Wu H, Wentworth C, Luo L, Collier-Hyams L, Neish AS (2008) Salmonella AvrA coordinates suppression of host immune and apoptotic defenses via JNK pathway blockade. Cell Host Microbe 3(4):233–244
Lin SL, Le TX, Cowen DS (2003) SptP, a Salmonella typhimurium type III-secreted protein, inhibits the mitogen-activated protein kinase pathway by inhibiting Raf activation. Cell Microbiol 5(4):267–275
Mazurkiewicz P, Thomas J, Thompson JA, Liu M, Arbibe L, Sansonetti P, Holden DW (2008) SpvC is a Salmonella effector with phosphothreonine lyase activity on host mitogen-activated protein kinases. Mol Microbiol 67(6):1371–1383
Rolhion N, Furniss RC, Grabe G, Ryan A, Liu M, Matthews SA, Holden DW (2016) Inhibition of nuclear transport of NF-kB p65 by the Salmonella type III secretion system effector SpvD. PLoS Pathog 12(5):e1005653
Sun H, Kamanova J, Lara-Tejero M, Galan JE (2016) A family of Salmonella type III secretion effector proteins selectively targets the NF-kappaB signaling pathway to preserve host homeostasis. PLoS Pathog 12(3):e1005484
Kramer RW, Slagowski NL, Eze NA, Giddings KS, Morrison MF, Siggers KA, Starnbach MN, Lesser CF (2007) Yeast functional genomic screens lead to identification of a role for a bacterial effector in innate immunity regulation. PLoS Pathog 3(2):e21
de Jong MF, Liu Z, Chen D, Alto NM (2016) Shigella flexneri suppresses NF-kappaB activation by inhibiting linear ubiquitin chain ligation. Nat Microbiol 1(7):16084
H. Ashida, M. Kim, M. Schmidt-Supprian, A. Ma, M. Ogawa, C. Sasakawa, A bacterial E3 ubiquitin ligase IpaH9.8 targets NEMO/IKKgamma to dampen the host NF-kappaB-mediated inflammatory response. Nature cell biology 12(1) (2010) 66–73; sup pp 1–9
Kim DW, Lenzen G, Page AL, Legrain P, Sansonetti PJ, Parsot C (2005) The Shigella flexneri effector OspG interferes with innate immune responses by targeting ubiquitin-conjugating enzymes. Proc Natl Acad Sci U S A 102(39):14046–14051
Sanada T, Kim M, Mimuro H, Suzuki M, Ogawa M, Oyama A, Ashida H, Kobayashi T, Koyama T, Nagai S, Shibata Y, Gohda J, Inoue J, Mizushima T, Sasakawa C (2012) The Shigella flexneri effector OspI deamidates UBC13 to dampen the inflammatory response. Nature 483(7391):623–626
Zhang Y, Muhlen S, Oates CV, Pearson JS, Hartland EL (2016) Identification of a distinct substrate-binding domain in the bacterial cysteine methyltransferase effectors NleE and OspZ. J Biol Chem 291(38):20149–20162
Lad SP, Yang G, Scott DA, Wang G, Nair P, Mathison J, Reddy VS, Li E (2007) Chlamydial CT441 is a PDZ domain-containing tail-specific protease that interferes with the NF-kappa B pathway of immune response. J Bacteriol 189(18):6619–6625
Trosky JE, Li Y, Mukherjee S, Keitany G, Ball H, Orth K (2007) VopA inhibits ATP binding by acetylating the catalytic loop of MAPK kinases. J Biol Chem 282(47):34299–34305
Nadler C, Baruch K, Kobi S, Mills E, Haviv G, Farago M, Alkalay I, Bartfeld S, Meyer TF, Ben-Neriah Y, Rosenshine I (2010) The type III secretion effector NleE inhibits NF-kappaB activation. PLoS Pathog 6(1):e1000743
Yen H, Ooka T, Iguchi A, Hayashi T, Sugimoto N, Tobe T (2010) NleC, a type III secretion protease, compromises NF-kappaB activation by targeting p65/RelA. PLoS Pathog 6(12):e1001231
Tweten RK (2005) Cholesterol-dependent cytolysins, a family of versatile pore-forming toxins. Infect Immun 73(10):6199–6209
Hamon MA, Batsche E, Regnault B, Tham TN, Seveau S, Muchardt C, Cossart P (2007) Histone modifications induced by a family of bacterial toxins. Proc Natl Acad Sci U S A 104(33):13467–13472
Hamon MA, Cossart P (2011) K+ efflux is required for histone H3 dephosphorylation by listeria monocytogenes listeriolysin O and other pore-forming toxins. Infect Immun 79(7):2839–2846
Dortet L, Lombardi C, Cretin F, Dessen A, Filloux A (2018) Pore-forming activity of the Pseudomonas aeruginosa type III secretion system translocon alters the host epigenome. Nat Microbiol 3(3):378–386
Brennan DF, Barford D (2009) Eliminylation: a post-translational modification catalyzed by phosphothreonine lyases. Trends Biochem Sci 34(3):108–114
Li H, Xu H, Zhou Y, Zhang J, Long C, Li S, Chen S, Zhou JM, Shao F (2007) The phosphothreonine lyase activity of a bacterial type III effector family. Science 315(5814):1000–1003
Arbibe L, Kim DW, Batsche E, Pedron T, Mateescu B, Muchardt C, Parsot C, Sansonetti PJ (2007) An injected bacterial effector targets chromatin access for transcription factor NF-kappaB to alter transcription of host genes involved in immune responses. Nat Immunol 8(1):47–56
Harouz H, Rachez C, Meijer BM, Marteyn B, Donnadieu F, Cammas F, Muchardt C, Sansonetti P, Arbibe L (2014) Shigella flexneri targets the HP1 gamma subcode through the phosphothreonine lyase OspF. EMBO J 33(22):2606–2622
Bierne H, Tham TN, Batsche E, Dumay A, Leguillou M, Kerneis-Golsteyn S, Regnault B, Seeler JS, Muchardt C, Feunteun J, Cossart P (2009) Human BAHD1 promotes heterochromatic gene silencing. Proc Natl Acad Sci U S A 106(33):13826–13831
Lebreton A, Cossart P, Bierne H (2012) Bacteria tune interferon responses by playing with chromatin. Virulence 3(1):87–91
Lakisic G, Lebreton A, Pourpre R, Wendling O, Libertini E, Radford EJ, Le Guillou M, Champy MF, Wattenhofer-Donze M, Soubigou G, Ait-Si-Ali S, Feunteun J, Sorg T, Coppee JY, Ferguson-Smith AC, Cossart P, Bierne H (2016) Role of the BAHD1 chromatin-repressive complex in placental development and regulation of steroid metabolism. PLoS Genet 12(3):e1005898
Eskandarian HA, Impens F, Nahori MA, Soubigou G, Coppee JY, Cossart P, Hamon MA (2013) A role for SIRT2-dependent histone H3K18 deacetylation in bacterial infection. Science 341(6145):1238858
Pereira JM, Chevalier C, Chaze T, Gianetto Q, Impens F, Matondo M, Cossart P, Hamon MA (2018) Infection reveals a modification of SIRT2 critical for chromatin association. Cell Rep 23(4):1124–1137
Pai RK, Pennini ME, Tobian AA, Canaday DH, Boom WH, Harding CV (2004) Prolonged toll-like receptor signaling by Mycobacterium tuberculosis and its 19-kilodalton lipoprotein inhibits gamma interferon-induced regulation of selected genes in macrophages. Infect Immun 72(11):6603–6614
Gehring AJ, Dobos KM, Belisle JT, Harding CV, Boom WH (2004) Mycobacterium tuberculosis LprG (Rv1411c): a novel TLR-2 ligand that inhibits human macrophage class II MHC antigen processing. J Immunol 173(4):2660–2668
Noss EH, Pai RK, Sellati TJ, Radolf JD, Belisle J, Golenbock DT, Boom WH, Harding CV (2001) Toll-like receptor 2-dependent inhibition of macrophage class II MHC expression and antigen processing by 19-kDa lipoprotein of Mycobacterium tuberculosis. J Immunol 167(2):910–918
Pennini ME, Pai RK, Schultz DC, Boom WH, Harding CV (2006) Mycobacterium tuberculosis 19-kDa lipoprotein inhibits IFN-gamma-induced chromatin remodeling of MHC2TA by TLR2 and MAPK signaling. J Immunol 176(7):4323–4330
Wang Y, Curry HM, Zwilling BS, Lafuse WP (2005) Mycobacteria inhibition of IFN-gamma induced HLA-DR gene expression by up-regulating histone deacetylation at the promoter region in human THP-1 monocytic cells. J Immunol 174(9):5687–5694
F. Lan, Y. Shi, Epigenetic regulation: methylation of histone and non-histone proteins. Science in China. Series C, Life sciences 52(4) (2009) 311–322
Lehnertz B, Ueda Y, Derijck AA, Braunschweig U, Perez-Burgos L, Kubicek S, Chen T, Li E, Jenuwein T, Peters AH (2003) Suv39h-mediated histone H3 lysine 9 methylation directs DNA methylation to major satellite repeats at pericentric heterochromatin. Current biology : CB 13(14):1192–1200
Sanz LA, Chamberlain S, Sabourin JC, Henckel A, Magnuson T, Hugnot JP, Feil R, Arnaud P (2008) A mono-allelic bivalent chromatin domain controls tissue-specific imprinting at Grb10. EMBO J 27(19):2523–2532
Cheng X, Zhang X (2007) Structural dynamics of protein lysine methylation and demethylation. Mutat Res 618(1–2):102–115
Li T, Lu Q, Wang G, Xu H, Huang H, Cai T, Kan B, Ge J, Shao F (2013) SET-domain bacterial effectors target heterochromatin protein 1 to activate host rDNA transcription. EMBO Rep 14(8):733–740
Pennini ME, Perrinet S, Dautry-Varsat A, Subtil A (2010) Histone methylation by NUE, a novel nuclear effector of the intracellular pathogen Chlamydia trachomatis. PLoS Pathog 6(7):e1000995
Mujtaba S, Winer BY, Jaganathan A, Patel J, Sgobba M, Schuch R, Gupta YK, Haider S, Wang R, Fischetti VA (2013) Anthrax SET protein: a potential virulence determinant that epigenetically represses NF-κB activation in infected macrophages. J Biol Chem 288(32):23458–23472
Yaseen I, Kaur P, Nandicoori VK, Khosla S (2015) Mycobacteria modulate host epigenetic machinery by Rv1988 methylation of a non-tail arginine of histone H3. Nat Commun 6:8922
Casadio F, Lu XD, Pollock SB, LeRoy G, Garcia BA, Muir TW, Roeder RG, Allis CD (2013) H3R42me2a is a histone modification with positive transcriptional effects. Proc Natl Acad Sci U S A 110(37):14894–14899
Hyland EM, Molina H, Poorey K, Jie C, Xie Z, Dai J, Qian J, Bekiranov S, Auble DT, Pandey A, Boeke JD (2011) An evolutionarily ‘young’ lysine residue in histone H3 attenuates transcriptional output in Saccharomyces cerevisiae. Genes Dev 25(12):1306–1319
L. Jose, R. Ramachandran, R. Bhagavat, R.L. Gomez, A. Chandran, S. Raghunandanan, R.V. Omkumar, N. Chandra, S. Mundayoor, R.A. Kumar, Hypothetical protein Rv3423.1 of Mycobacterium tuberculosis is a histone acetyltransferase. The FEBS journal 283(2) (2016) 265–81
Chopra P, Singh B, Singh R, Vohra R, Koul A, Meena LS, Koduri H, Ghildiyal M, Deol P, Das TK, Tyagi AK, Singh Y (2003) Phosphoprotein phosphatase of Mycobacterium tuberculosis dephosphorylates serine-threonine kinases PknA and PknB. Biochem Biophys Res Commun 311(1):112–120
Agarwal S, Agarwal S, Jin H, Pancholi P, Pancholi V (2012) Serine/threonine phosphatase (SP-STP), secreted from Streptococcus pyogenes, is a pro-apoptotic protein. J Biol Chem 287(12):9147–9167
Rolando M, Sanulli S, Rusniok C, Gomez-Valero L, Bertholet C, Sahr T, Margueron R, Buchrieser C (2013) Legionella pneumophila effector RomA uniquely modifies host chromatin to repress gene expression and promote intracellular bacterial replication. Cell Host Microbe 13(4):395–405
Schuhmacher MK, Rolando M, Brohm A, Weirich S, Kudithipudi S, Buchrieser C, Jeltsch A (2018) The Legionella pneumophila methyltransferase RomA methylates also non-histone proteins during infection. J Mol Biol 430(13):1912–1925
Lin H, Su X, He B (2012) Protein lysine acylation and cysteine succination by intermediates of energy metabolism. ACS Chem Biol 7(6):947–960
Sabari BR, Zhang D, Allis CD, Zhao YM (2017) Metabolic regulation of gene expression through histone acylations. Nat Rev Mol Cell Bio 18(2):90–101
Wei W, Liu X, Chen J, Gao S, Lu L, Zhang H, Ding G, Wang Z, Chen Z, Shi T, Li J, Yu J, Wong J (2017) Class I histone deacetylases are major histone decrotonylases: evidence for critical and broad function of histone crotonylation in transcription. Cell Res 27(7):898–915
B.R. Sabari, Z.Y. Tang, H. Huang, V. Yong-Gonzalez, H. Molina, H.E. Kong, L.Z. Dai, M. Shimada, J.R. Cross, Y.M. Zhao, R.G. Roeder, C.D. Allis, Intracellular crotonyl-CoA stimulates transcription through p300-catalyzed histone crotonylation. (vol 58, pg 203, 2015), Mol Cell 69(3) (2018) 533–533
Koh A, De Vadder F, Kovatcheva-Datchary P, Backhed F (2016) From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell 165(6):1332–1345
Fellows R, Denizot J, Stellato C, Cuomo A, Jain P, Stoyanova E, Balazsi S, Hajnady Z, Liebert A, Kazakevych J, Blackburn H, Correa RO, Fachi JL, Sato FT, Ribeiro WR, Ferreira CM, Peree H, Spagnuolo M, Mattiuz R, Matolcsi C, Guedes J, Clark J, Veldhoen M, Bonaldi T, Vinolo MAR, Varga-Weisz P (2018) Microbiota derived short chain fatty acids promote histone crotonylation in the colon through histone deacetylases. Nat Commun 9(1):105
Simon JA, Kingston RE (2013) Occupying chromatin: Polycomb mechanisms for getting to genomic targets, stopping transcriptional traffic, and staying put. Mol Cell 49(5):808–824
Cavaillon JM, Adib-Conquy M (2006) Bench-to-bedside review: endotoxin tolerance as a model of leukocyte reprogramming in sepsis. Crit Care 10(5)
Chan C, Li L, McCall CE, Yoza BK (2005) Endotoxin tolerance disrupts chromatin remodeling and NF-kappaB transactivation at the IL-1beta promoter. J Immunol 175(1):461–468
El Gazzar M, Yoza BK, Hu JY, Cousart SL, McCall CE (2007) Epigenetic silencing of tumor necrosis factor alpha during endotoxin tolerance. J Biol Chem 282(37):26857–26864
Chen X, El Gazzar M, Yoza BK, McCall CE (2009) The NF-kappaB factor RelB and histone H3 lysine methyltransferase G9a directly interact to generate epigenetic silencing in endotoxin tolerance. J Biol Chem 284(41):27857–27865
B. Novakovic, E. Habibi, S.Y. Wang, R.J.W. Arts, R. Davar, W. Megchelenbrink, B. Kim, T. Kuznetsova, M. Kox, J. Zwaag, F. Matarese, S.J. van Heeringen, E.M. Janssen-Megens, N. Sharifi, C. Wang, F. Keramati, V. Schoonenberg, P. Flicek, L. Clarke, P. Pickkers, S. Heath, I. Gut, M.G. Netea, J.H.A. Martens, C. Logie, H.G. Stunnenberg, Beta-Glucan reverses the epigenetic state of LPS-induced immunological tolerance. Cell 167(5) (2016) 1354-+
Netea MG, Quintin J, van der Meer JW (2011) Trained immunity: a memory for innate host defense. Cell Host Microbe 9(5):355–361
Yoshida K, Maekawa T, Zhu Y, Renard-Guillet C, Chatton B, Inoue K, Uchiyama T, Ishibashi K, Yamada T, Ohno N, Shirahige K, Okada-Hatakeyama M, Ishii S (2015) The transcription factor ATF7 mediates lipopolysaccharide-induced epigenetic changes in macrophages involved in innate immunological memory. Nat Immunol 16(10):1034–1043
Quintin J, Saeed S, Martens JHA, Giamarellos-Bourboulis EJ, Ifrim DC, Logie C, Jacobs L, Jansen T, Kullberg BJ, Wijmenga C, Joosten LAB, Xavier RJ, van der Meer JWM, Stunnenberg HG, Netea MG (2012) Candida albicans infection affords protection against reinfection via functional reprogramming of monocytes. Cell Host Microbe 12(2):223–232
Ostuni R, Piccolo V, Barozzi I, Polletti S, Termanini A, Bonifacio S, Curina A, Prosperini E, Ghisletti S, Natoli G (2013) Latent enhancers activated by stimulation in differentiated cells. Cell 152(1–2):157–171
Rasid O, Chevalier C, Camarasa T, Fitting C, Cavaillon J-M, Hamon MA (2019) H3K4me1 supports memory-like NK cells induced by systemic inflammation. Cell Rep 29(12):3933–3945
Acknowledgments
We apologize to any colleagues whose work was not included in this review due to space limitations. We thank Michael Connor for critical reading of the manuscript.
Funding
Work in the Chromatin and Infection Group is supported by the Pasteur Institute and the Agence National de la Recherche (ANR-EpiBActIn). Wenyang Dong is part of the Pasteur - Paris University (PPU) International PhD Program, a project which has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Sklodowska-Curie grant agreement no. 665807. Wenyang Dong is supported by the EUR G.E.N.E. (reference #ANR-17-EURE-0013) and is part of the Université de Paris IdEx #ANR-18-IDEX-0001 funded by the French Government through its “Investments for the Future” program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
This article is a contribution to the special issue on Infection-induced epigenetic changes and the pathogenesis of diseases - Guest Editor: Nicole Fischer
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dong, W., Hamon, M.A. Revealing eukaryotic histone-modifying mechanisms through bacterial infection. Semin Immunopathol 42, 201–213 (2020). https://doi.org/10.1007/s00281-019-00778-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00281-019-00778-9