Abstract
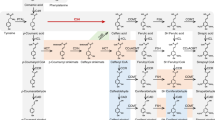
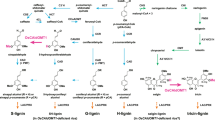
Plant laccases catalyse the oxidation of monolignols in lignification, a process reinforcing the cell wall of many different cell types that provide mechanical support, nutrient transportation and defence against pathogens in plants1. The isozymes display a broad range of substrate preferences. Here, the substrate preference of a laccase (ZmLac3) from Zea mays (maize) was characterized. The crystal structure of ZmLac3 revealed a compact and deep substrate-binding pocket, and the binding modes of sinapyl alcohol (SinA) and coniferyl alcohol (ConA) were solved. On the basis of structural data and kinetics analysis, we propose that the regionalization of polar and hydrophobic surfaces in the binding pocket of ZmLac3 is vital for defining the orientation of SinA/ConA binding. The extra methoxyl group in SinA makes substantial contributions to interactions between SinA and ZmLac3, which are absent in the ZmLac3–ConA complex. In summary, the polar and hydrophobic interactions between SinA/ConA and ZmLac3 determine the binding positions of the monolignols in ZmLac3. These results provide valuable insight about ZmLac3 catalysis and should aid industrial processes that use plant laccases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Barros, J., Serk, H., Granlund, I. & Pesquet, E. The cell biology of lignification in higher plants. Ann. Bot. 115, 1053–1074 (2015).
Jones, S. M. & Solomon, E. I. Electron transfer and reaction mechanism of laccases. Cell. Mol. Life Sci. 72, 869–883 (2015).
Caparros-Ruiz, D., Fornale, S., Civardi, L., Puigdomenech, P. & Rigau, J. Isolation and characterisation of a family of laccases in maize. Plant Sci. 171, 217–225 (2006).
McCaig, B. C., Meagher, R. B. & Dean, J. F. D. Gene structure and molecular analysis of the laccase-like multicopper oxidase (LMCO) gene family in Arabidopsis thaliana. Planta 221, 619–636 (2005).
Berthet, S. et al. Disruption of LACCASE4 and 17 results in tissue-specific alterations to lignification of Arabidopsis thaliana stems. Plant Cell 23, 1124–1137 (2011).
Zhao, Q. et al. Laccase is necessary and nonredundant with peroxidase for lignin polymerization during vascular development in Arabidopsis. Plant Cell 25, 3976–3987 (2013).
Zhang, Y. et al. The cotton laccase gene GhLAC15 enhances Verticillium wilt resistance via an increase in defence-induced lignification and lignin components in the cell walls of plants. Mol. Plant Pathol. 20, 309–322 (2019).
Wildhagen, H. et al. Genes and gene clusters related to genotype and drought-induced variation in saccharification potential, lignin content and wood anatomical traits in Populus nigra. Tree Physiol. 38, 320–339 (2018).
Sun, Q. et al. MicroRNA528 affects lodging resistance of maize by regulating lignin biosynthesis under nitrogen-luxury conditions. Mol. Plant 11, 806–814 (2018).
Bao, W., O’Malley, D. M., Whetten, R. & Sederoff, R. R. A laccase associated with lignification in loblolly pine xylem. Science 260, 672–674 (1993).
Mcdougall, G. J. A comparison of proteins from the developing xylem of compression and non-compression wood of branches of Sitka spruce (Picea sitchensis) reveals a differentially expressed laccase. J. Exp. Bot. 51, 1395–1401 (2000).
Fang, F. et al. An intracellular laccase is responsible for the epicatechin mediated anthocyanin degradation in litchi fruit pericarp. Plant Physiol. 169, 2391–2408 (2015).
Kallio, J. P. et al. Structure–function studies of a Melanocarpus albomyces laccase suggest a pathway for oxidation of phenolic compounds. J. Mol. Biol. 392, 895–909 (2009).
Enguita, F. J., Martins, L. O., Henriques, A. O. & Carrondo, M. A. Crystal structure of a bacterial endospore coat component a laccase with enhanced thermostability properties. J. Biol. Chem. 278, 19416–19425 (2003).
Adzhubei, A. A., Sternberg, M. J. & Makarov, A. A. Polyproline-II helix in proteins: structure and function. J. Mol. Biol. 425, 2100–2132 (2013).
Chou, E. Y. et al. Distribution, mobility, and anchoring of lignin-related oxidative enzymes in Arabidopsis secondary cell walls. J. Exp. Bot. 69, 1849–1859 (2018).
Liu, Z., Xie, T., Zhong, Q. & Wang, G. Crystal structure of CotA laccase complexed with 2,2-azinobis-(3-ethylbenzothiazoline-6-sulfonate) at a novel binding site. Acta Crystallogr. F 72, 328–335 (2016).
Ducros, V. et al. Crystal structure of the type-2 Cu depleted laccase from Coprinus cinereus at 2.2 Å resolution. Nat. Struct. Mol. Biol. 5, 310–316 (1998).
Maestre-Reyna, M. et al. Structural and functional roles of glycosylation in fungal laccase from Lentinus sp. PLoS ONE 10, e0120601 (2015).
Jubb, H. C. et al. Arpeggio: a web server for calculating and visualising interatomic interactions in protein structures. J. Mol. Biol. 429, 365–371 (2017).
Biot, C., Buisine, E., Kwasigroch, J.-M., Wintjens, R. & Rooman, M. Probing the energetic and structural role of amino acid/nucleobase cation–π interactions in protein–ligand complexes. J. Biol. Chem. 277, 40816–40822 (2002).
Gupta, N. & Farinas, E. T. Directed evolution of CotA laccase for increased substrate specificity using Bacillus subtilis spores. Protein Eng. Des. Sel. 23, 679–682 (2010).
Andberg, M. et al. Essential role of the C terminus in Melanocarpus albomyces laccase for enzyme production, catalytic properties and structure. FEBS J. 276, 6285–6300 (2009).
Jiao, Y. et al. Improved maize reference genome with single-molecule technologies. Nature 546, 524–527 (2017).
Bosch, M., Mayer, C. D., Cookson, A. & Donnison, I. S. Identification of genes involved in cell wall biogenesis in grasses by differential gene expression profiling of elongating and non-elongating maize internodes. J. Exp. Bot. 62, 3545–3561 (2011).
Barriere, Y., Courtial, A., Chateigner-Boutin, A. L., Denoue, D. & Grima-Pettenati, J. Breeding maize for silage and biofuel production, an illustration of a step forward with the genome sequence. Plant Sci. 242, 310–329 (2016).
Hoopes, J. T. & Dean, J. F. Ferroxidase activity in a laccase-like multicopper oxidase from Liriodendron tulipifera. Plant Physiol. Biochem. 42, 27–33 (2004).
Shen, Y. et al. Genome expression profile analysis reveals important transcripts in maize roots responding to the stress of heavy metal Pb. Physiol. Plant. 147, 270–282 (2013).
Pourcel, L. et al. TRANSPARENT TESTA10 encodes a laccase-like enzyme involved in oxidative polymerization of flavonoids in Arabidopsis seed coat. Plant Cell 17, 2966–2980 (2005).
Sterjiades, R., Dean, J. F. D., Gamble, G., Himmelsbach, D. S. & Eriksson, K. E. L. Extracellular laccases and peroxidases from sycamore maple (Acer pseudoplatanus) cell-suspension cultures: reactions with monolignols and lignin model compounds. Planta 190, 75–87 (1993).
Lee, Y., Rubio, M. C., Alassimone, J. & Geldner, N. A mechanism for localized lignin deposition in the endodermis. Cell 153, 402–412 (2013).
Munk, L., Sitarz, A. K., Kalyani, D. C., Mikkelsen, J. D. & Meyer, A. S. Can laccases catalyze bond cleavage in lignin? Biotechnol. Adv. 33, 13–24 (2015).
pPIC9K a Pichia vector for multicopy integration and secreted expression (Thermo Fisher Scientific, 2010); http://tools.thermofisher.com/content/sfs/manuals/ppic9k_man.pdf
Munteanu, B., Braun, M. & Boonrod, K. Improvement of PCR reaction conditions for site-directed mutagenesis of big plasmids. J. Zhejiang Univ. Sci. B 13, 244–247 (2012).
Pichia fermentation process guidelines (Invitrogen, 2002); http://tools.thermofisher.com/content/sfs/manuals/pichiaferm_prot.pdf
O’Callaghan, J., O’Brien, M. M., McClean, K. & Dobson, A. D. W. Optimisation of the expression of a Trametes versicolor laccase gene in Pichia pastoris. J. Ind. Microbiol. Biotechnol. 29, 55–59 (2002).
Wang, Q. S. et al. The macromolecular crystallography beamline of SSRF. Nucl. Sci. Tech. 26, 12–17 (2015).
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL-3000: the integration of data reduction and structure solution—from diffraction images to an initial model in minutes. Acta Crystallogr. D 62, 859–866 (2010).
Mccoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
Adams, P. D. et al. The Phenix software for automated determination of macromolecular structures. Methods 55, 94–106 (2011).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Dundas, J. et al. CASTp: computed atlas of surface topography of proteins with structural and topographical mapping of functionally annotated residues. Nucleic Acids Res. 34, W116–W118 (2006).
Tan, K. P., Varadarajan, R. & Madhusudhan, M. S. DEPTH: a web server to compute depth and predict small-molecule binding cavities in proteins. Nucleic Acids Res. 39, W242–W248 (2011).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein using the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976).
Bond, C. S. TopDraw: a sketchpad for protein structure topology cartoons. Bioinformatics 19, 311–312 (2003).
Lafayette, P. R., Eriksson, K. E. L. & Dean, J. F. D. Nucleotide-sequence of a cDNA clone encoding an acidic laccase from sycamore maple (Acer pseudoplatanus L.). Plant Physiol. 107, 667–668 (1995).
Wang, G. D., Li, Q. J., Luo, B. & Chen, X. Y. Ex planta phytoremediation of trichlorophenol and phenolic allelochemicals via an engineered secretory laccase. Nat. Biotechnol. 22, 893–897 (2004).
Cesarino, I. et al. Expression of SofLAC, a new laccase in sugarcane, restores lignin content but not S:G ratio of Arabidopsis lac17 mutant. J. Exp. Bot. 64, 1769–1781 (2013).
Sato, Y. & Whetten, R. W. Characterization of two laccases of loblolly pine (Pinus taeda) expressed in tobacco BY-2 cells. J. Plant Res. 119, 581–588 (2006).
Ranocha, P. et al. Biochemical characterization, molecular cloning and expression of laccases—a divergent gene family—in poplar. Eur. J. Biochem. 259, 485–495 (1999).
Koutaniemi, S., Malmberg, H. A., Simola, L. K., Teeri, T. H. & Karkonen, A. Norway spruce (Picea abies) laccases: characterization of a laccase in a lignin-forming tissue culture. J. Integr. Plant Biol. 57, 341–348 (2015).
Bond, C. S. & Schuettelkopf, A. W. ALINE: a WYSIWYG protein-sequence alignment editor for publication-quality alignments. Acta Crystallogr. D 65, 510–512 (2009).
Bertrand, T. et al. Crystal structure of a four-copper laccase complexed with an arylamine: insights into substrate recognition and correlation with kinetics. Biochemistry 41, 7325–7333 (2002).
Kallio, J. P. et al. Crystal structure of an ascomycete fungal laccase from Thielavia arenaria—common structural features of asco-laccases. FEBS J. 278, 2283–2295 (2011).
Holm, L. & Rosenstrom, P. DAIL server: conservation mapping in 3D. Nucleic Acids Res. 38, 545–549 (2010).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Chengzhi, L. et al. Gramene: a growing plant comparative genomics resource. Nucleic Acids Res. 36, 947–953 (2008).
Hoegger, P. J., Kilaru, S., James, T. Y., Thacker, J. R. & Kues, U. Phylogenetic comparison and classification of laccase and related multicopper oxidase protein sequences. FEBS J. 273, 2308–2326 (2006).
Lawrence, C. J. et al. MaizeGDB: the maize model organism database for basic, translational, and applied research. Int. J. Plant Genomics 2008, 496957 (2008).
Acknowledgements
This work was supported by grants from the Innovative Academy of Seed Design (INASEED) at the Chinese Academy of Science, initial grants from the 100 Talents Program of the Chinese Academy of Sciences and the open fund from the Key Laboratory of Environmental and Applied Microbiology at the Chinese Academy of Sciences (grant nos. KLCAS-2016-09 and KLCAS-2017-05). We thank the Shanghai Synchrotron Radiation Facilities (SSRF) and the National Center for Protein Science Shanghai (NCPSS) for the provision of synchrotron radiation facilities and efficient support.
Author information
Authors and Affiliations
Contributions
T.X., Z.L. and G.W. designed the research. T.X. expressed, purified and crystallized ZmLac3 and performed the biochemical assays. T.X. and Z.L. collected and processed the diffraction data. Z.L. determined the structures. T.X., Z.L. and G.W. wrote the manuscript. G.W. directed and supervised all of the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Jane Agger, Kurt Fagerstedt and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
Data collection and refinement statistics.
Extended Data Fig. 2 Topology structure of Zmlac3.
Sheets in domains I, II and III are coloured red, blue and green, respectively. Alpha-helices and 310 helices are shown as cyan and magenta cylinders, respectively. The figure was prepared by TopDraw software47.
Extended Data Fig. 3 Sequence alignment between zmlac3 and other plant laccases.
Similarities are depicted in grey-scale shading and residues that are identical indicated by a black background. Four substrate-binding loops are marked by blue boxes. Orange stars placed above the sequences indicate amino acids that may interact with substrates. 17 A. thaliana laccases (AtLacs)4, 5 Z. mays W64A laccases (ZmLacs)3, A.pseudoplatanus laccase (ApLac)48, Litchi chinensis laccase (LcLac)12, Gossypium arboretum laccase (GaLac)49, Gossypium hirsutum-15 (GhLac15)6, Saccharum officinarum laccase (SoLac)50, two P. taeda laccases (PtLacs)51, Populus euramericana laccase-90 (PeLac90)52 and Picea abies laccase-4 (PaLac4)53 were included in the alignment. The alignment was generated by ALINE54.
Extended Data Fig. 5
SAA, SAV and depth of substrate binding pockets in laccases.
Extended Data Fig. 6 Electron density map around SinA/ConA located in the binding pocket.
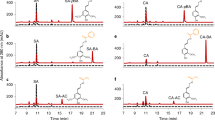
The 2Fo-Fc electron density map (1σ level, blue) and the omit Fo-Fc map (2σ level, red) around SinA (a) and ConA (b) in the binding pocket are represented in mesh.
Extended Data Fig. 7
Laccase candidates identified in maize B73. To explore the laccase candidates in Z. mays B73 genome (Refgen_V4)24, protein sequences of 17 AtLacs from A. thaliana and 5 ZmLacs from Z. mays WB64A were used as queries for BLASTP58 on Gramene59 online. The hits without three cupredoxin domains or copper binding sites were excluded. Then, the phylogenetic analysis of the candidate proteins with AtLacs, ZmLacs and plant ascorbate oxidases60 was performed in MEGA software (http://www.megasoftware.net). The hits clustered with plant ascorbate oxidases were discarded. Finally, a total of 19 laccase candidates (Gene IDs in Gramene59 and MaizeGDB61 are listed) were found in Z. mays B73 genome, which showed 36–99% sequence identity to the counterparts in Z. mays WB64A (Extended Data Fig. 9).
Extended Data Fig. 8 Neighbor-joining phylogenetic analysis of Zmlac3 with other laccases in Maize W64A and B73.
The phylogenetic tree was constructed by MEGA software (http://www.megasoftware.net/) with bootstrap tests for 1,000 replicates. 10 AtLacs4, 5 ZmLacs3 and 19 predicted ZmLacs_B73 (Extended Data Fig. 7) were clustered into five groups.
Extended Data Fig. 9
Sequence identity of laccases in maize W64A and B73.
Supplementary information
Rights and permissions
About this article
Cite this article
Xie, T., Liu, Z. & Wang, G. Structural basis for monolignol oxidation by a maize laccase. Nat. Plants 6, 231–237 (2020). https://doi.org/10.1038/s41477-020-0595-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-0595-5