Abstract
Angiosarcoma (AS) is the most frequent primary sarcoma of the breast but nevertheless remains uncommon, accounting for <0.05% of breast malignancies. Secondary mammary AS arise following radiation therapy for breast cancer, in contrast to primary AS which occur sporadically. Essentially all show aggressive clinical behavior independent of histologic grade and most are treated by mastectomy. MYC amplification is frequently identified in radiation-induced AS but only rarely in primary mammary AS (PMAS). As a heterogeneous group, AS from various anatomic sites have been shown to harbor recurrent alterations in TP53, MAP kinase pathway genes, and genes involved in angiogenic signaling including KDR (VEGFR2) and PTPRB. In part due to its rarity, the pathogenesis of PMAS has not been fully characterized. In this study, we examined the clinical, pathologic, and genomic features of ten cases of PMAS, including one patient with bilateral disease. Recurrent genomic alterations were identified in KDR (70%), PIK3CA/PIK3R1 (70%), and PTPRB (30%), each at higher frequencies than reported in AS across all sites. Six tumors harbored a KDR p.T771R hotspot mutation, and all seven KDR-mutant cases showed evidence suggestive of biallelism (four with loss of heterozygosity and three with two aberrations). Of the seven tumors with PI3K alterations, six harbored pathogenic mutations other than in the canonical PIK3CA residues which are most frequent in breast cancer. Three AS were hypermutated (≥10 mutations/megabase (Mb)); hypermutation was seen concurrent with KDR or PIK3CA mutations. The patient with bilateral disease demonstrated shared alterations, indicative of contralateral metastasis. No MYC or TP53 aberrations were detected in this series. Immunohistochemistry for VEGFR2 was unable to discriminate between KDR-mutant tumors and benign vascular lesions of the breast. These findings highlight the underrecognized frequency of KDR and PIK3CA mutation in PMAS, and a significant subset with hypermutation, suggesting a pathogenesis distinct from other AS.
Similar content being viewed by others
Introduction
Angiosarcomas (AS) represent a heterogeneous group of malignant vascular tumors expressing the morphologic and phenotypic properties of endothelial cells. These tumors represent 1–2% of soft tissue sarcomas and arise within different anatomic sites, among a wide age range, and in a variety of clinical settings, such as prior radiation therapy or chronic lymphedema [1,2,3]. Surgery and possible radiotherapy are the mainstay of treatment for patients with localized disease [2]. Metastasis is reported in ~50% of cases, often treated with adjuvant chemotherapy [4]. Median overall survival time is ~50 months for local disease and ~10 months for metastatic cases [5]. Treatment with targeted agents, such as tyrosine kinase inhibitors, has shown partial responses in selected patients, but overall prognosis remains poor [6,7,8,9].
AS of the breast is rare, accounting for 0.04% of all breast malignancies, yet remains the most common breast sarcoma [10]. Primary mammary AS (PMAS) arise spontaneously in women with no relevant breast history, typically in their third and fourth decades. Secondary mammary AS (SMAS) occur either in the setting of radiation therapy following breast conservation surgery or lymphedema following mastectomy (Stewart–Treves syndrome); postradiation AS are by far the most frequent contemporaneously [11, 12]. On average, patients with SMAS present at an older age, usually within the sixth or seventh decade and a median postradiation latency of 6–7 years [13, 14]. Primary AS typically involve the mammary parenchyma and may secondarily involve the skin, whereas postradiation AS most often primarily arise in the skin and may invade deeper into the breast [15,16,17,18]. AS may be classified into low, intermediate, and high grades based on a combination of histologic features including degree of vasoformative growth, cytologic atypia, mitotic activity, and presence of endothelial multilayering/tufting, solid growth, necrosis, and hemorrhage (blood lakes) [19]. Historically thought to be prognostic, more recent studies have failed to show an association between tumor grade and clinical outcome [20, 21]. Most cases of mammary AS are treated by mastectomy, although the use of breast conserving surgery may be increasing [16, 17, 22, 23]. Frequent sites of metastasis include the lung, liver, contralateral breast, bone, and other skin and soft tissue sites [16, 17, 23].
Distinct differences have emerged among AS subgroups, and prognosis has been shown to vary depending on primary site [2]. Among breast tumors, some reports suggest that SMAS has a worse prognosis than PMAS, while others find no difference [16, 22, 24]. Genomic studies of AS across anatomic sites have shown a heterogeneous mutational spectrum. Activating alterations in mitogen-activated protein kinase (MAPK) pathway genes, such as KRAS, HRAS, NRAS, BRAF, and MAPK1, were detected in over half of AS cases in one series [25]. Reports of TP53 mutation vary from 4 to 35% [25,26,27]. In the breast and other sites, amplification of MYC is the hallmark of most postradiation secondary AS (54–100% of cases), while only sparingly reported in primary AS [28,29,30,31,32,33,34]. Amplification of FLT4 (VEGFR3, vascular endothelial growth factor receptor 3) has also been identified in SMAS (18–25% of cases), typically coamplified with MYC [28, 31]. Additional recurrently mutated genes in SMAS include PTPRB (45%), a tyrosine phosphatase specific to endothelial cells that inhibits angiogenesis, and PLCG1 (9%), which encodes phospholipase C gamma 1 [27, 35]. In one series of AS from various sites, Antonescu et al. identified mutations in KDR, the gene that encodes VEGFR2, in 10% of cases (4/44); all KDR-mutant tumors originated in the breast/chest wall (4/17) and included both PMAS and SMAS (two cases each) [36].
As a specific subgroup, the molecular landscape of PMAS has not been fully characterized. Rare cases of PMAS have been analyzed in the aforementioned case series among larger numbers of SMAS or nonmammary AS. Italiano et al. included nine PMAS in their study reporting the absence of both deleterious TP53 mutations and activating PIK3CA mutations (using hotspot analysis on a subset of cases) [26]. To our knowledge, the largest series to date with genomic analysis included 22 PMAS, of which five (23%) harbored KDR mutations and four (18%) demonstrated PLCG1 mutations [35].
In this study, we comprehensively characterize a cohort of PMAS by capture-based next-generation sequencing of 479 cancer-related genes. We sought to define their molecular drivers and determine whether these rare tumors have a pathogenesis distinct from other AS. Although limited in number of cases, our work highlights a high frequency of KDR, PIK3CA, and PTPRB mutations in PMAS, as well as a subset demonstrating hypermutation. Such novel genomic findings may have much-needed therapeutic implications for these aggressive tumors.
Materials and methods
Study population
With institutional review board approval, the pathology archives of Stanford University and the University of California San Francisco were searched for cases of PMAS. Tumors were selected from patients with no known prior breast cancer or radiation treatment. Cases were reviewed to confirm that the tumors were centered in the mammary parenchyma; AS appearing to originate in the dermis were excluded. The series is comprised of ten cases total, including one with bilateral tumors. Clinical information was obtained from online electronic medical records when available.
Capture-based next-generation DNA sequencing
Matched normal and tumor tissue was selected from nine cases for capture-based next-generation DNA sequencing. The bilateral case consisted of separate tumor-only specimens. Sequencing libraries were prepared from genomic DNA extracted from punch biopsies or macrodissected unstained sections from formalin fixed paraffin embedded tissue. Target enrichment was performed by hybrid capture using a custom oligonucleotide library. Capture-based next-generation sequencing was performed at the UCSF Clinical Cancer Genomics Laboratory, using an assay (UCSF500 panel) that targets the coding regions of 479 cancer-related genes, select introns from ~40 genes, and the TERT promoter with a total sequencing footprint of 2.8 Mb (Supplementary Table S1). Sequencing was performed on a HiSeq 2500 (Illumina, San Diego, CA). Duplicate sequencing reads were removed computationally to allow for accurate allele frequency determination and copy number calling. The analysis was based on the human reference sequence UCSC build hg19 (NCBI build 37), using the following software packages: BWA: 0.7.10-r789, Samtools: 1.1 (using htslib 1.1), Picard tools: 1.97 (1504), GATK: 2014.4–3.3.0–0-ga3711, CNVkit: 0.3.3, Pindel: 0.2.5a7, SATK: 2013.1–10- gd6fa6c3, Annovar: v2015Mar22, Freebayes: 0.9.20, and Delly: 0.5.9 [37,38,39,40,41,42,43,44,45,46,47]. Only insertions/deletions (indels) up to 100 bp in length were included in the mutational analysis. Somatic single nucleotide variants and indels were visualized and verified using Integrated Genome Viewer. Genome-wide copy number analysis based on on-target and off-target reads was performed by CNVkit and Nexus Copy Number (Biodiscovery, Hawthorne, CA, USA). Lollipop plots were modified from MutationMapper [48, 49]. Tumor mutational burden was quantified for matched tumor-normal cases. For hypermutated tumors, signatures were delineated by deconstructSigs using nonsynonymous mutations; only alterations with ≥5 mutant reads were reported [50]. Microsatellite instability analysis was performed with MSIsensor and interpreted using revised Bethesda guidelines [51, 52].
Immunohistochemistry
Immunohistochemistry for VEGFR2 was performed on whole slide sections of PMAS and benign vascular lesions using the rabbit monoclonal 55B11 antibody (Cell Signaling Technology, Beverly, MA) at 1:200 with Tris–EDTA antigen retrieval (pH 9).
Results
Clinicopathologic features of primary mammary angiosarcomas
Clinicopathologic features of PMAS included in this study are shown in Table 1 and Fig. 1. All patients were female, with ages ranging from 31 to 64 years (mean 45 years). Nine of 11 tumors were right sided. The majority presented as palpable masses (8/11, 73%). Tumor size at initial excision ranged from 2 to 13.6 cm (mean 5.2 cm). Histologic grades ranged from low to high. All patients ultimately underwent mastectomy; 6 of 11 (55%) mastectomies followed an earlier excision or lumpectomy. Six of 10 (60%) patients were treated with adjuvant chemotherapy and radiation. One woman had germline testing which showed a pathogenic FANCA c.987_990delTCAC (p.T329fs) mutation. This patient initially presented with AS of the right breast (Case 9R), and 2 years later subsequently presented with AS of the left breast (Case 9L). She underwent mastectomy for each diagnosis with no adjuvant chemotherapy; the left breast tumor was clinically considered a separate primary AS. Three women demonstrated definite metastasis (Cases 2, 6, and 7); metastatic sites included liver and lung. In patients with available follow-up clinical data (9/10 women, average 22 months), two women died of disease (at 11 and 27 months following excision), one woman is alive with disease, and the remainder have no evidence of residual disease. Although limited in number, no associations were evident between patient age, tumor size, histologic grade, initial surgery, adjuvant therapy, metastasis, and outcome.
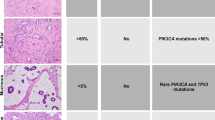
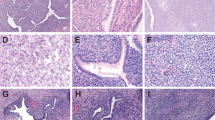
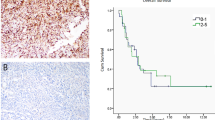
Representative photomicrographs of hematoxylin and eosin (H&E)-stained PMAS: (a) complex anastomosing channels and “blood lakes” of high grade tumor, (b) inset at higher power showing cellular pleomorphism, (c) papillary tufts and invasion into adjuvant adipose tissue, (d) angiosarcoma involving fibroadenoma (*). e Immunohistochemistry for CD34 showing strong and diffuse expression in AS (blue arrowhead) with negative staining in adjacent breast epithelium (red arrowhead). f Immunohistochemistry for VEFGR2 was diffusely positive in all PMAS (n = 8) and benign vascular lesions (n = 4, inset with angiolipoma) tested. Magnification, ×100 (a, d–f), ×200 (c), ×400 (b).
Genomic features and immunohistochemistry of primary mammary angiosarcomas
Genomic data are depicted in Fig. 2 and Supplementary Table S2. The mean target sequencing coverage was 548 (± 279) unique reads per target interval (Supplementary Table S3). The number of identified nonsynonymous coding mutations across the 2.8 Mb footprint of the panel ranged from 1 to 74 per Mb (median 3, mean 16 ± 24).
KDR was the most frequently altered gene in the series, detected in seven PMAS (of ten, 70%). Six of these tumors harbored a KDR p.T771R hotspot mutation. All seven KDR-mutant AS showed evidence suggestive of biallelism: four with loss of heterozygosity (LOH) and three with two separate aberrations (cis vs. trans orientation could not be definitively determined). Pathogenic alterations in PI3K were also common (7/10, 70%), including six cases with PIK3CA mutations (specifically p.P104L, p.P539R, p.E545A, p.M1004V in two cases, and p.M1043I) and one case with two PIK3R1 mutations. Among the PIK3CA alterations, only one mutation (p.E545A of Case 2) was in the most common canonical residue in exon 9 (E542 and E545) and none were in the common canonical residue in exon 20 (H1047) [53]. In addition, inactivating mutations in PTPRB were identified (3/10, 30%), with no association with KDR or PIK3CA. No alterations in MYC, FLT4, or TP53 were identified. PLCG1 is not included on the UCSF500 panel and could not be evaluated.
For the patient with bilateral but asynchronous disease, differential tumor sequencing indicated shared mutations in PTPRB (two alterations) and KMT2D, at mutant allele frequencies not compatible with germline findings (Supplementary Table S4). These results are virtually confirmative of metastasis to the contralateral breast. Interestingly, a PIK3CA p.M1004V mutation was identified in the primary AS of the right breast, but this was not detected in the subsequent left breast tumor. As expected, this patient’s previously identified germline FANCA inactivating mutation was also identified in both specimens (at appropriately ~50% mutant allele frequency). Treated with consecutive mastectomies and no adjuvant therapy, she remains with no evidence of disease nearly 4 years after initial diagnosis. No pathogenic germline alterations were identified in the remaining patients.
Copy number analysis revealed two cases with concomitant amplification of the KDR locus, both in mutant-positive tumors. Case 5 demonstrated mutant allele-specific amplification (10×) of 4q11-q13.1 including KDR, KIT, and PDGFRA genes, and Case 7 harbored 4q12 amplification with KDR (5×) and KIT (2×). Recurrent copy number gains included chromosome 7 (7/10, 70%), distal 17q (6/10, 60%), and the majority of chromosome 8 (5/10, 50%). The bilateral case showed identical copy number changes, with gains in distal 15q and distal 17q in both specimens. Overall, no associations between chromosomal gain and other genomic aberrations were seen.
Hypermutation was detected in three tumors (Cases 6–8), defined as ≥10 mutations/Mb, each with a prevalence of C > T transitions (Supplementary Fig. 1) [54]. Signature analysis of hypermutated cases was inconclusive, with dominant COSMIC Signatures 1 and 6 identified; these are typically ascribed to age and DNA mismatch repair deficiency, respectively. However, there was no association between patient age and hypermutation status, and all tumors tested were microsatellite stable by MSIsensor (cutoff = 10). In addition, hypermutation was seen concurrent with KDR or PIK3CA alterations.
Based on the high frequency of KDR mutations in PMAS, immunohistochemistry for VEGFR2 was performed on the majority of cases (n = 9), supplemented with a limited number of benign vascular lesions of the breast including hemangiomas and angiolipomas (n = 4). Prior works have published conflicting data, with both VEGFR2 protein overexpression by immunohistochemistry reported in KDR-mutant AS, and paradoxically others reporting AS were negative for VEGFR2 expression [36, 55, 56]. Here, we found that VEGFR2 was positive in all vascular lesions tested, with no discrimination between KDR-positive and -negative AS, as well as benign entities (Fig. 1f).
Discussion
This work revealed an unexpectedly high frequency of KDR and PI3K activating mutations in PMAS, each occurring at ~70%. In fact, every tumor was found to have an alteration in KDR and/or PIK3CA; the mutations were not mutually exclusive. Here, we confirm the previously reported association between KDR mutation and AS of the breast and chest wall, but determination of this high occurrence is made possible by exclusively focusing on PMAS in our series [35, 36]. The hotspot mutation KDR p.T771R in exon 16 is particularly prevalent, occurring in the transmembrane domain of VEGFR2. To our knowledge, this alteration has only been reported in vascular lesions: overwhelmingly cases of AS, with a lone exception of a sporadic angioma arising in a patient on anti-VEGFR2 therapy (ramucirumab) for metastatic rectal cancer [48, 49, 57, 58]. Of note, the KDR mutation in this angioma case report was a single copy with no other alteration or evidence of LOH [58]. In contrast, biallelism was present in each of the KDR-mutant tumors in our series, a novel finding which suggests it may be necessary for PMAS pathogenesis.
Activating PI3K mutations are common in breast carcinomas but relatively rare in sarcomas. Specific to the breast, PIK3CA alterations have been reported in higher grade phyllodes tumors and in one case each of undifferentiated pleomorphic sarcoma and osteosarcoma [59,60,61,62]. Yet PI3KCA mutations were not identified in AS of the breast or other sites in a prior study despite evidence of PI3K/AKT/mTOR pathway activation [26]. In contrast, PIK3CA (and one case of PIK3R1) alterations were frequent in our series. Prior work focused on selected exons with canonical PIK3CA hotspot mutations, whereas our study more comprehensively assessed the entire coding sequence [26]. Specifically, alterations were detected at P104, P539, M1004 (two cases), and M1043, occurring in exons 2 (adaptor binding domain), 10 (helical domain), and 21 (kinase domain), respectively. Each of these mutations have reported in vitro oncogenic properties and recurrent alteration in breast and other cancers [48, 49, 63,64,65]. The identification of PIK3CA mutations in PMAS may have significant clinical importance, especially with the recent approval of PI3K inhibition for the treatment of PIK3CA-mutant advanced breast carcinoma [66].
Other recurrently altered genes in this study include PTPRB (n = 3) and KMT2D (n = 2). Mutations in KMT2D, a histone methyltransferase, have been previously identified in breast AS among various cancers [59]. Inactivating mutations in PTPRB are rare in other tumor types yet frequent in AS, although heretofore had not been reported in PMAS [27]. Each PTPRB-mutant PMAS demonstrated at least one truncating alteration, predicted to disrupt the coding sequence before the tyrosine phosphatase domain. Two of three tumors showed a second nontruncating alteration, suggestive of possible biallelism; no features of LOH were observed in the third case with a single frameshift mutation. This pattern of mutation, suggestive but not definitive for a recessive driver mechanism of pathogenesis, is similar to a prior report [27]. Moreover, alterations in other protein phosphatases, such as PTPRT and PPP2R1A, have been reported in human and canine AS [67]. Although no KRAS, HRAS, or NRAS aberrations were detected in this series, one PMAS showed inactivating somatic alterations in RASA1, a negative regulator of the RAS and MAPK pathways, and similarly reported in human and canine AS [67]. Notably, no TP53, MYC, or FLT4 aberrations were identified.
Regarding Case 9, the genomic data of bilateral disease is essentially confirmatory of metastasis, despite the 2-year time interval between tumors, absence of adjuvant therapy, and no evidence of additional disease following surgery alone for the past 1.5 years. Clinical germline testing of this patient had shown a pathogenic FANCA mutation, and she was managed under the assumption of bilateral primary AS. However, identical somatic nonrecurrent indels in PTPRB and KMT2D were identified in each tumor, a finding incompatible with two independent malignancies. Mutations in FANCA are the most common cause of Fanconi’s anemia and have been implicated in increased risk of various cancers, but a specific association with AS is unclear and no evidence of LOH at this locus is identified in either of the patient’s tumors. Notably, a PIK3CA mutation was detected in the initial right-sided PMAS but not the left-sided tumor; discordance in PIK3CA status between primary and metastatic breast cancer is a known phenomenon and may be applicable in this context [68]. Such a case may illustrate the importance of the endogenous microenvironment of the breast that contains niche-promoting elements conducive to contralateral metastasis. The relatively indolent behavior of this case of PMAS is remarkable; in contrast sequencing studies that have positively identified metastasis among metachronous contralateral breast carcinomas have shown poor outcomes [69, 70].
Intriguingly, 3 of 10 PMAS (30%) demonstrated somatic hypermutation. Hypermutation has been recently reported in a subset of sarcomas, primarily with high levels of ultraviolet (UV)-associated mutations and categorized as Signature 7 (or cluster C6) in the hypermutant tumor classification [54, 71]. In fact, in a large series interrogating the tumor mutational burden of 100,000 diverse cancer genomes, hypermutation was detected in ~13% of soft tissue AS, although anatomic site and radiation history were not provided [72]. To our knowledge, the finding of somatic hypermutation specifically in PMAS is novel. Cases were too few to identify any associations between hypermutation and other clinicopathologic features, but it is noted that the two patients who died of disease demonstrated hypermutant tumors.
Although limited in number, our series nonetheless clearly suggests a number of therapeutic options for PMAS. The outcome data show that some tumors may require no more treatment beyond surgery, even curiously including Case 9. On the other hand, aggressive or recurrent tumors may benefit from targeted therapies implicated by the high frequency of KDR (VEGFR2) and PIK3CA mutations in PMAS, as well as a subset showing hypermutation. Our findings coincide with recently reported therapeutic efforts. For AS, data on targeting the PI3K pathway are limited to in vitro work in cell lines, but this potential treatment avenue may indirectly benefit from the rapidly expanding targeting of PIK3CA-mutant breast carcinomas [73]. A number of agents targeting the VEGFR pathway have been reported, including the anti-VEGF monoclonal antibody bevacizumab and tyrosine kinase inhibitors pazopanib, sorafenib, sunitinib, and axitinib [74]. Results in AS patients have been underwhelming, but such trials have been primarily comprised of tumors of heterogeneous origin and unknown KDR status. Limited individual case reports of therapeutic responses to VEGFR inhibition in which the tumor had been first shown to harbor KDR amplification/overexpression have been described [75, 76]. How PMAS with pathogenic KDR mutations specifically respond to such inhibitors has not been fully characterized, and is an avenue for further clinical investigation. For the subset of hypermutated tumors, immune checkpoint inhibitors may show promise. A small series of heterogeneous AS without mutational burden testing demonstrated mixed responses to immune checkpoint blockade [77]. Yet a recent case report of a hypermutated AS with a UV-induced signature treated with pembrolizumab showed a dramatic clinical response [78].
In conclusion, by exclusively focusing on a series of PMASs, our work expands on the earlier work of Antonescu et al. by identifying a high frequency of recurrent KDR and PIK3CA mutations in these rare tumors [26, 35, 36]. Comprehensive genomic sequencing of PMAS also revealed alterations in PTPRB and a subset of tumors with somatic hypermutation. Importantly, we note an ongoing patient-partnered research platform termed the Angiosarcoma Project as part of the Count Me In initiative, which is accumulating genetic data on a series of AS including PMAS. Ad hoc review of this publicly available data on cBioPortal reveals a similar prevalence of KDR alterations (9/14 cases, 64%) in PMAS, as well as frequent PIK3CA mutations (5/14 cases, 36%), providing additional support of our findings [48, 49]. Large-scale endeavors such as this have an incredible potential for unraveling the genomic underpinnings of especially rare tumors. Together, our findings provide evidence that PMAS demonstrate a pathogenesis distinct from AS of other sites, with the hope of guiding future therapies.
References
Young RJ, Brown NJ, Reed MW, Hughes D, Woll PJ. Angiosarcoma. Lancet Oncol. 2010;11:983–91.
Fayette J, Martin E, Piperno-Neumann S, Le Cesne A, Robert C, Bonvalot S, et al. Angiosarcomas, a heterogeneous group of sarcomas with specific behavior depending on primary site: a retrospective study of 161 cases. Ann Oncol. 2007;18:2030–6.
Naka N, Ohsawa M, Tomita Y, Kanno H, Uchida A, Myoui A, et al. Prognostic factors in angiosarcoma: a multivariate analysis of 55 cases. J Surg Oncol. 1996;61:170–6.
Mark RJ, Poen JC, Tran LM, Fu YS, Juillard GF. Angiosarcoma. A report of 67 patients and a review of the literature. Cancer. 1996;77:2400–6.
Lahat G, Dhuka AR, Hallevi H, Xiao L, Zou C, Smith KD, et al. Angiosarcoma: clinical and molecular insights. Ann Surg. 2010;251:1098–106.
Yoo KH, Kim HS, Lee SJ, Park SH, Kim SJ, Kim SH, et al. Efficacy of pazopanib monotherapy in patients who had been heavily pretreated for metastatic soft tissue sarcoma: a retrospective case series. BMC Cancer. 2015;15:154.
Azzariti A, Porcelli L, Mangia A, Saponaro C, Quatrale AE, Popescu OS, et al. Irradiation-induced angiosarcoma and anti-angiogenic therapy: a therapeutic hope? Exp Cell Res. 2014;321:240–7.
Ray-Coquard I, Italiano A, Bompas E, Le Cesne A, Robin YM, Chevreau C, et al. Sorafenib for patients with advanced angiosarcoma: a phase II trial from the French Sarcoma Group (GSF/GETO). Oncologist. 2012;17:260–6.
Taylor BS, Barretina J, Maki RG, Antonescu CR, Singer S, Ladanyi M. Advances in sarcoma genomics and new therapeutic targets. Nat Rev Cancer. 2011;11:541–57.
Brodie C, Provenzano E. Vascular proliferations of the breast. Histopathology. 2008;52:30–44.
Body G, Sauvanet E, Calais G, Fignon A, Fetissof F, Lansac J. [Cutaneous angiosarcoma of the breast following surgery and irradiation of breast adenocarcinoma]. J Gynecol Obstet Biol Reprod. 1987;16:479–83.
Stewart FW, Treves N. Lymphangiosarcoma in postmastectomy lymphedema; a report of six cases in elephantiasis chirurgica. Cancer. 1948;1:64–81.
Depla AL, Scharloo-Karels CH, de Jong MAA, Oldenborg S, Kolff MW, Oei SB, et al. Treatment and prognostic factors of radiation-associated angiosarcoma (RAAS) after primary breast cancer: a systematic review. Eur J Cancer. 2014;50:1779–88.
Brenn T, Fletcher CD. Radiation-associated cutaneous atypical vascular lesions and angiosarcoma: clinicopathologic analysis of 42 cases. Am J Surg Pathol. 2005;29:983–96.
Baker GM, Schnitt SJ. Vascular lesions of the breast. Semin Diagn Pathol. 2017;34:410–9.
Fraga-Guedes C, Gobbi H, Mastropasqua MG, Botteri E, Luini A, Viale G. Primary and secondary angiosarcomas of the breast: a single institution experience. Breast Cancer Res Treat. 2012;132:1081–8.
Scow JS, Reynolds CA, Degnim AC, Petersen IA, Jakub JW, Boughey JC. Primary and secondary angiosarcoma of the breast: the Mayo Clinic experience. J Surg Oncol. 2010;101:401–7.
Lucas DR. Angiosarcoma, radiation-associated angiosarcoma, and atypical vascular lesion. Arch Pathol Lab Med. 2009;133:1804–9.
Donnell RM, Rosen PP, Lieberman PH, Kaufman RJ, Kay S, Braun DW Jr, et al. Angiosarcoma and other vascular tumors of the breast. Am J Surg Pathol. 1981;5:629–42.
Nascimento AF, Raut CP, Fletcher CD. Primary angiosarcoma of the breast: clinicopathologic analysis of 49 cases, suggesting that grade is not prognostic. Am J Surg Pathol. 2008;32:1896–904.
Rosen PP, Kimmel M, Ernsberger D. Mammary angiosarcoma. The prognostic significance of tumor differentiation. Cancer. 1988;62:2145–51.
Abdou Y, Elkhanany A, Attwood K, Ji W, Takabe K, Opyrchal M. Primary and secondary breast angiosarcoma: single center report and a meta-analysis. Breast Cancer Res Treat. 2019;178:523–33.
Kunkiel M, Maczkiewicz M, Jagiello-Gruszfeld A, Nowecki Z. Primary angiosarcoma of the breast-series of 11 consecutive cases-a single-centre experience. Curr Oncol. 2018;25:e50–3.
Luini A, Gatti G, Diaz J, Botteri E, Oliveira E, Cecilio Sahium de Almeida R, et al. Angiosarcoma of the breast: the experience of the European Institute of Oncology and a review of the literature. Breast Cancer Res Treat. 2007;105:81–5.
Murali R, Chandramohan R, Möller I, Scholz SL, Berger M, Huberman K, et al. Targeted massively parallel sequencing of angiosarcomas reveals frequent activation of the mitogen activated protein kinase pathway. Oncotarget. 2015;6:36041–52.
Italiano A, Chen CL, Thomas R, Breen M, Bonnet F, Sevenet N, et al. Alterations of the p53 and PIK3CA/AKT/mTOR pathways in angiosarcomas: a pattern distinct from other sarcomas with complex genomics. Cancer. 2012;118:5878–87.
Behjati S, Tarpey PS, Sheldon H, Martincorena I, Van Loo P, Gundem G, et al. Recurrent PTPRB and PLCG1 mutations in angiosarcoma. Nat Genet. 2014;46:376–9.
Cornejo KM, Deng A, Wu H, Cosar EF, Khan A, St Cyr M, et al. The utility of MYC and FLT4 in the diagnosis and treatment of postradiation atypical vascular lesion and angiosarcoma of the breast. Hum Pathol. 2015;46:868–75.
Ginter PS, Mosquera JM, MacDonald TY, D’Alfonso TM, Rubin MA, Shin SJ. Diagnostic utility of MYC amplification and anti-MYC immunohistochemistry in atypical vascular lesions, primary or radiation-induced mammary angiosarcomas, and primary angiosarcomas of other sites. Hum Pathol. 2014;45:709–16.
Mentzel T, Schildhaus HU, Palmedo G, Buttner R, Kutzner H. Postradiation cutaneous angiosarcoma after treatment of breast carcinoma is characterized by MYC amplification in contrast to atypical vascular lesions after radiotherapy and control cases: clinicopathological, immunohistochemical and molecular analysis of 66 cases. Mod Pathol. 2012;25:75–85.
Guo T, Zhang L, Chang NE, Singer S, Maki RG, Antonescu CR. Consistent MYC and FLT4 gene amplification in radiation-induced angiosarcoma but not in other radiation-associated atypical vascular lesions. Genes Chromosomes Cancer. 2011;50:25–33.
Manner J, Radlwimmer B, Hohenberger P, Mössinger K, Küffer S, Sauer C, et al. MYC high level gene amplification is a distinctive feature of angiosarcomas after irradiation or chronic lymphedema. Am J Pathol. 2010;176:34–9.
Fernandez AP, Sun Y, Tubbs RR, Goldblum JR, Billings SD. FISH for MYC amplification and anti-MYC immunohistochemistry: useful diagnostic tools in the assessment of secondary angiosarcoma and atypical vascular proliferations. J Cutan Pathol. 2012;39:234–42.
Fraga-Guedes C, André S, Mastropasqua MG, Botteri E, Toesca A, Rocha RM, et al. Angiosarcoma and atypical vascular lesions of the breast: diagnostic and prognostic role of MYC gene amplification and protein expression. Breast Cancer Res Treat. 2015;151:131–40.
Huang SC, Zhang L, Sung YS, Chen CL, Kao YC, Agaram NP, et al. Recurrent CIC gene abnormalities in angiosarcomas: a molecular study of 120 cases with concurrent investigation of PLCG1, KDR, MYC, and FLT4 gene alterations. Am J Surg Pathol. 2016;40:645–55.
Antonescu CR, Yoshida A, Guo T, Chang NE, Zhang L, Agaram NP, et al. KDR activating mutations in human angiosarcomas are sensitive to specific kinase inhibitors. Cancer Res. 2009;69:7175–9.
Picard: a set of tools (in Java) for working with next generation sequencing data in the BAM: Broad Institute. http://broadinstitute.github.io/picard.
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43:491–8.
Garrison E, Marth G. Haplotype-based variant detection from short-read sequencing. 2012. http://arxiv.org/abs/1207.3907.
Li H, Durbin R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics. 2010;26:589–95.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics. 2009;25:2078–9.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–303.
Rausch T, Zichner T, Schlattl A, Stutz AM, Benes V, Korbel JO. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics. 2012;28:i333–9.
Talevich E, Shain AH, Botton T, Bastian BC. CNVkit: genome-wide copy number detection and visualization from targeted DNA sequencing. PLoS Comput Biol. 2016;12:e1004873.
Van der Auwera GA, Carneiro MO, Hartl C, Poplin R, Del Angel G, Levy-Moonshine A, et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr Protoc Bioinforma. 2013;43:11–33.
Yang H, Wang K. Genomic variant annotation and prioritization with ANNOVAR and wANNOVAR. Nat Protoc. 2015;10:1556–66.
Ye K, Schulz MH, Long Q, Apweiler R, Ning Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics. 2009;25:2865–71.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2:401–4.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
Rosenthal R, McGranahan N, Herrero J, Taylor BS, Swanton C. DeconstructSigs: delineating mutational processes in single tumors distinguishes DNA repair deficiencies and patterns of carcinoma evolution. Genome Biol. 2016;17:31.
Niu B, Ye K, Zhang Q, Lu C, Xie M, McLellan MD, et al. MSIsensor: microsatellite instability detection using paired tumor-normal sequence data. Bioinformatics. 2014;30:1015–6.
Umar A, Boland CR, Terdiman JP, Syngal S, de la Chapelle A, Rüschoff J, et al. Revised Bethesda guidelines for hereditary nonpolyposis colorectal cancer (Lynch syndrome) and microsatellite instability. J Natl Cancer Inst. 2004;96:261–8.
Ligresti G, Militello L, Steelman LS, Cavallaro A, Basile F, Nicoletti F, et al. PIK3CA mutations in human solid tumors: role in sensitivity to various therapeutic approaches. Cell Cycle. 2009;8:1352–8.
Campbell BB, Light N, Fabrizio D, Zatzman M, Fuligni F, de Borja R, et al. Comprehensive analysis of hypermutation in human cancer. Cell. 2017;171:1042–56.e10.
Brar R, West R, Witten D, Raman B, Jacobs C, Ganjoo K. Breast angiosarcoma: case series and expression of vascular endothelial growth factor. Case Rep Oncol. 2009;2:242–50.
Itakura E, Yamamoto H, Oda Y, Tsuneyoshi M. Detection and characterization of vascular endothelial growth factors and their receptors in a series of angiosarcomas. J Surg Oncol. 2008;97:74–81.
Jauhri M, Bhatnagar A, Gupta S, Bp M, Minhas S, Shokeen Y, et al. Prevalence and coexistence of KRAS, BRAF, PIK3CA, NRAS, TP53, and APC mutations in Indian colorectal cancer patients: next-generation sequencing-based cohort study. Tumour Biol. 2017;39:1–11.
Lim YH, Odell ID, Ko CJ, Choate KA. Somatic p.T771R KDR (VEGFR2) mutation arising in a sporadic angioma during ramucirumab therapy. JAMA Dermatol. 2015;151:1240–3.
Lim SZ, Ng CCY, Rajasegaran V, Guan P, Selvarajan S, Thike AA, et al. Genomic profile of breast sarcomas: a comparison with malignant phyllodes tumours. Breast Cancer Res Treat. 2019;174:365–73.
Tan J, Ong CK, Lim WK, Ng CC, Thike AA, Ng LM, et al. Genomic landscapes of breast fibroepithelial tumors. Nat Genet. 2015;47:1341–5.
Liu SY, Joseph NM, Ravindranathan A, Stohr BA, Greenland NY, Vohra P, et al. Genomic profiling of malignant phyllodes tumors reveals aberrations in FGFR1 and PI-3 kinase/RAS signaling pathways and provides insights into intratumoral heterogeneity. Mod Pathol. 2016;29:1012–27.
Tsang JY, Go EM, Tse GM. Identification of clinically relevant alterations in phyllodes tumor of the breast by amplicon-based next-generation sequencing. Breast Cancer Res Treat. 2015;151:717–9.
Dogruluk T, Tsang YH, Espitia M, Chen F, Chen T, Chong Z, et al. Identification of variant-specific functions of PIK3CA by rapid phenotyping of rare mutations. Cancer Res. 2015;75:5341–54.
Gymnopoulos M, Elsliger MA, Vogt PK. Rare cancer-specific mutations in PIK3CA show gain of function. Proc Natl Acad Sci USA. 2007;104:5569–74.
Ng PK, Li J, Jeong KJ, Shao S, Chen H, Tsang YH, et al. Systematic functional annotation of somatic mutations in cancer. Cancer Cell. 2018;33:450–62.e10.
André F, Ciruelos E, Rubovszky G, Campone M, Loibl S, Rugo HS, et al. Alpelisib for PIK3CA-mutated, hormone receptor-positive advanced breast cancer. N Engl J Med. 2019;380:1929–40.
Megquier K, Turner-Maier J, Swofford R, Kim JH, Sarver AL, Wang C, et al. Comparative genomics reveals shared mutational landscape in canine hemangiosarcoma and human angiosarcoma. Mol Cancer Res. 2019;17:2410–21.
Dupont Jensen J, Laenkholm AV, Knoop A, Ewertz M, Bandaru R, Liu W, et al. PIK3CA mutations may be discordant between primary and corresponding metastatic disease in breast cancer. Clin Cancer Res. 2011;17:667–77.
Begg CB, Ostrovnaya I, Geyer FC, Papanastasiou AD, Ng CKY, Sakr RA, et al. Contralateral breast cancers: independent cancers or metastases? Int J Cancer. 2018;142:347–56.
Klevebring D, Lindberg J, Rockberg J, Hilliges C, Hall P, Sandberg M, et al. Exome sequencing of contralateral breast cancer identifies metastatic disease. Breast Cancer Res Treat. 2015;151:319–24.
Alexandrov LB, Nik-Zainal S, Wedge DC, Aparicio SA, Behjati S, Biankin AV, et al. Signatures of mutational processes in human cancer. Nature. 2013;500:415–21.
Chalmers ZR, Connelly CF, Fabrizio D, Gay L, Ali SM, Ennis R, et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 2017;9:34.
Wada M, Horinaka M, Yasuda S, Masuzawa M, Sakai T, Katoh N. PDK1 is a potential therapeutic target against angiosarcoma cells. J Dermatol Sci. 2015;78:44–50.
Weidema ME, Versleijen-Jonkers YMH, Flucke UE, Desar IME, van der Graaf WTA. Targeting angiosarcomas of the soft tissues: a challenging effort in a heterogeneous and rare disease. Crit Rev Oncol Hematol. 2019;138:120–31.
Silva E, Gatalica Z, Vranic S, Basu G, Reddy SK, Voss A. Refractory angiosarcoma of the breast with VEGFR2 upregulation successfully treated with sunitinib. Breast J. 2015;21:205–7.
Yang L, Liu L, Han B, Han W, Zhao M. Apatinib treatment for KIT- and KDR-amplified angiosarcoma: a case report. BMC Cancer. 2018;18:618.
Florou V, Rosenberg AE, Wieder E, Komanduri KV, Kolonias D, Uduman M, et al. Angiosarcoma patients treated with immune checkpoint inhibitors: a case series of seven patients from a single institution. J Immunother Cancer. 2019;7:213.
Momen S, Fassihi H, Davies HR, Nikolaou C, Degasperi A, Stefanato CM, et al. Dramatic response of metastatic cutaneous angiosarcoma to an immune checkpoint inhibitor in a patient with xeroderma pigmentosum: whole-genome sequencing aids treatment decision in end-stage disease. Cold Spring Harb Mol Case Stud. 2019;5:a004408.
Acknowledgements
The authors wish to thank Shirley Kwok for technical assistance in histology and immunohistochemistry, as well as members of the UCSF Clinical Cancer Genomics Laboratory for sequencing work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Beca, F., Krings, G., Chen, YY. et al. Primary mammary angiosarcomas harbor frequent mutations in KDR and PIK3CA and show evidence of distinct pathogenesis. Mod Pathol 33, 1518–1526 (2020). https://doi.org/10.1038/s41379-020-0511-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-020-0511-6
This article is cited by
-
PIK3CA is recurrently mutated in canine mammary tumors, similarly to in human mammary neoplasia
Scientific Reports (2023)