Abstract
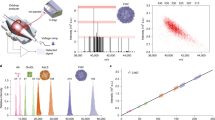
An Orbitrap-based ion analysis procedure determines the direct charge for numerous individual protein ions to generate true mass spectra. This individual ion mass spectrometry (I2MS) method for charge detection enables the characterization of highly complicated mixtures of proteoforms and their complexes in both denatured and native modes of operation, revealing information not obtainable by typical measurements of ensembles of ions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Datasets utilized for the I2MS analyses can be found on the MassiVE repository, MSV000083840. Source data are available for Figs. 2 and 3 and Supplementary Figs. 2–4 and 9. Additional desired data that support the charge-determination findings of this study are available from the corresponding authors upon request.
Code availability
Custom compiled code associated with the I2MS creation process is available via Supplementary Software or from the corresponding authors upon reasonable request.
References
Aebersold, R. & Mann, M. Mass spectrometry-based proteomics. Nature 422, 198–207 (2003).
Elliott, A. G., Harper, C. C., Lin, H.-W. & Williams, E. R. Mass, mobility and MSn measurements of single ions using charge detection mass spectrometry. Analyst 142, 2760–2769 (2017).
Keifer, D. Z., Pierson, E. E. & Jarrold, M. F. Charge detection mass spectrometry: weighing heavier things. Analyst 142, 1654–1671 (2017).
Benner, W. H. A gated electrostatic ion trap to repetitiously measure the charge and m/z of large electrospray ions. Anal. Chem. 69, 4162–4168 (1997).
Schmidt, H. T., Cederquist, H., Jensen, J. & Fardi, A. Conetrap: a compact electrostatic ion trap. Nucl. Instrum. Methods Phys. Res., Sect. B 173, 523–527 (2001).
Makarov, A. & Denisov, E. Dynamics of ions of intact proteins in the orbitrap mass analyzer. J. Am. Soc. Mass. Spectrom. 20, 1486–1495 (2009).
Rose, R. J., Damoc, E., Denisov, E., Makarov, A. & Heck, A. J. R. High-sensitivity orbitrap mass analysis of intact macromolecular assemblies. Nat. Methods 9, 1084 (2012).
Kafader, J. O. et al. Measurement of individual ions sharply increases the resolution of orbitrap mass spectra of proteins. Anal. Chem. 91, 2776–2783 (2019).
Contino, N. C. & Jarrold, M. F. Charge detection mass spectrometry for single ions with a limit of detection of 30 charges. Int. J. Mass Spectrom. 345-347, 153–159 (2013).
Kafader, J. O. et al. STORI plots enable accurate tracking of individual ion signals. J. Am. Soc. Mass. Spectrom. 30, 2200–2203 (2019).
Tran, J. C. et al. Mapping intact protein isoforms in discovery mode using top-down proteomics. Nature 480, 254 (2011).
Gómez, S. M., Nishio, J. N., Faull, K. F. & Whitelegge, J. P. The chloroplast grana proteome defined by intact mass measurements from liquid chromatography mass spectrometry. Mol. Cell Proteom. 1, 46–59 (2002).
Smith, R. D. et al. An accurate mass tag strategy for quantitative and high-throughput proteome measurements. Proteomics 2, 513–523 (2002).
The Human Protein Atlas https://www.proteinatlas.org (2019).
Lermyte, F., Tsybin, Y. O., O’Connor, P. B. & Loo, J. A. Top or Middle? Up or Down? toward a standard lexicon for protein top-down and allied mass spectrometry approaches. J. Am. Soc. Mass. Spectrom. 30, 1149–1157 (2019).
Schachner, L. F. et al. Standard proteoforms and their complexes for native mass spectrometry. J. Am. Soc. Mass. Spectrom. 30, 1190–1198 (2019).
Sobott, F. & Robinson, C. V. Characterising electrosprayed biomolecules using tandem-MS—the noncovalent GroEL chaperonin assembly. Int. J. Mass Spectrom. 236, 25–32 (2004).
Valegård, K., Liljas, L., Fridborg, K. & Unge, T. The three-dimensional structure of the bacterial virus MS2. Nature 345, 36–41 (1990).
Asensio, M. A. et al. A selection for assembly reveals that a single amino acid mutant of the bacteriophage MS2 coat protein forms a smaller virus-like particle. Nano Lett. 16, 5944–5950 (2016).
Hartman, E. C. et al. Quantitative characterization of all single amino acid variants of a viral capsid-based drug delivery vehicle. Nat. Commun. 9, 1385–1385 (2018).
Fornelli, L. et al. Advancing top-down analysis of the human proteome using a benchtop quadrupole-orbitrap mass spectrometer. J. Proteome Res. 16, 609–618 (2017).
Anderson, L. C. et al. Identification and characterization of human proteoforms by top-down LC-21 Tesla FT-ICR mass spectrometry. J. Proteome Res. 16, 1087–1096 (2017).
LeDuc, R. D. et al. Accurate estimation of context-dependent false discovery rates in top-down proteomics. Mol. Cell Proteom. 18, 796–805 (2019).
Ntai, I. et al. Applying label-free quantitation to top down proteomics. Anal. Chem. 86, 4961–4968 (2014).
Durbin, K. R., Skinner, O. S., Fellers, R. T. & Kelleher, N. L. Analyzing internal fragmentation of electrosprayed ubiquitin ions during beam-type collisional dissociation. J. Am. Soc. Mass. Spectrom. 26, 782–787 (2015).
Freeke, J., Robinson, C. V. & Ruotolo, B. T. Residual counter ions can stabilise a large protein complex in the gas phase. Int. J. Mass spectrom. 298, 91–98 (2010).
Skinner, O. S. et al. An informatic framework for decoding protein complexes by top-down mass spectrometry. Nat. Methods 13, 237 (2016).
Acknowledgements
This work was funded by the Intensifying Innovation program from Thermo Fisher Scientific and was carried out in collaboration with the National Resource for Translational and Developmental Proteomics under Grant P41 GM108569 from the National Institute of General Medical Sciences, National Institutes of Health with additional support from the Sherman Fairchild Foundation, and the instrumentation award (S10OD025194) from NIH Office of Director. In addition, we would like to thank Luca Fornelli and Timothy K. Toby for collecting and analyzing the HEK-293 LC–MS runs utilized for our intact-mass-tag I2MS analysis.
Author information
Authors and Affiliations
Contributions
Data acquisition and analysis was performed by J.O.K. with assistance from R.D.M. and K.R.D. and with additional input from all authors. R.D.M. prepared samples with help from J.O.K., and B.I. and D.T.-E. assembled and purified the MS2 virus-like particles. J.O.K., B.P.E., K.R.D., R.T.F., S.C.B., P.F.Y., M.W.S. and P.D.C. developed models and created programs to analyze the data. K.R.D., V.Z., A.A.M., J.T.M., D.L.S., M.W.S., P.D.C. and N.L.K. provided additional technical support and guidance. N.L.K., P.D.C. and M.W.S. collected funding support and oversaw the project. J.O.K. and N.L.K. wrote the manuscript. All authors critically reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
V.Z., A.A.M., J.T.M., D.L.S., P.F.Y. and M.W.S. are employees of Thermo Fisher Scientific.
Additional information
Peer review information Allison Doerr was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Ion Survival Histograms.
Histogram of individual myoglobin +20 ions collected that demonstrate the increased ion survival with decreased Orbitrap central electrode (CE) voltages. The solid blue lines indicate separation of low-intensity noise peaks, mid-level intensity real ions that have decayed during the acquisition event, and high-intensity real ions that did not decay during the detection event. The high intensity ions are used to create an I2MS spectrum. As the CE voltage was reduced from -5 kV to -1 kV, ion velocity decreased and this increased ion survival over a 4 second acquisition from just 3% (CE = -5 kV) to 71% (CE = -1 kV). This finding drastically increased the efficiency of I2MS during its development.
Supplementary Figure 2 4 Protein Mix.
Spectra of a synthetic mixture of intact proteins created by tradition ensemble ion collection plotted in m/z space (a), or via the I2MS process (b). The table in panel (c) lists the known proteins in the mixture along with their masses and corresponding figures of merit determined from the I2MS spectrum in panel (b). The number assigned to each protein in the table correspond to the labeled features in the m/z spectrum and corresponding I2MS spectrum in panels (a) and (b), respectively.
Supplementary Figure 3 Reduced Antibody Standard.
Heavy and Light chain of a monoclonal IgG measured by conventional ensemble FT-MS analysis (a) compared to I2MS (b). Various forms of the Heavy and Light chain species including guanidine adducts and glycosylations caused a rise in baseline in the ensemble spectra were easily distinguished by I2MS and caused no such rise in spectral baseline. Blue arrows are indicative of approximately a 60 Da guanidine adduct and red arrows indicate the addition of a 162 Da hexose-type glycosylation. Peaks arising from assignment of +1 or -1 charge state to individual ions are labeled in the I2MS spectrum with an asterisk*.
Supplementary Figure 4 Native Protein Complexes.
Ensemble ion spectra plotted in m/z space (a,d) and their corresponding I2MS mass spectra (b,e) for the ~232 kDa pyruvate kinase (left) and 801 kDa groEL (right) complexes with labeled charge states in panels (a) and (d). The implementation of I2MS yields accurate spectra in the mass domain with resolution (Res) comparable to the various charge states collected in m/z space. Charge state values and relative charge state intensity assignments (c,f) agreed on average within <5% with their corresponding m/z-domain spectra (a,d).
Supplementary Figure 5 VLP Subunit Analysis.
Characterization of the capsid coat proteins (CPs) used to assemble MS2 virus-like particles analyzed in Fig. 3. The WT (top) and MINI (bottom) CP masses are within 4 part-per-million of their theoretical values. Mass measurements from denaturing the VLP capsids produced intact mass values that match and confirm the sequence differences of each species. The one differing amino acid (proline vs. serine at position 37) between the two proteoforms results in the structural differences between the WT and MINI VLPs.
Supplementary Figure 6 Individual Ion STORI Plots.
Individual Ion STORI plots and their slope values for +13 (black), +25 (red), and +39 (blue) charge states of ubiquitin, myoglobin, and carbonic anhydrase, respectively. The slope of the Selective Temporal Overview of Resonant Ion (STORI) plot of an individual ion on the outer electrode of the Orbitrap is proportional to the charge of the ion signal. The STORI slope is calibrated externally to provide a single, integer result. See reference10 for a complete description of the STORI process and Fig. 1 for an overview of the entire I2MS process.
Supplementary Figure 7 Charge Calibration Function.
Linear calibration of STORI slope values as a function of charge state. Calibration is utilized to determine the charge of an unknown individual ion signal with a calculated STORI slope. The rigorously linear statistical relationship between individual ion STORI slopes and quantized charge states independent of ion m/z is the foundation I2MS is built upon. The proteins utilized to produce the various charge states are labeled by their respective points on the graph. The highlighted red points/trace correspond to the calibration function on a Q-Exactive Plus instrument while the blue points/trace correspond to the calibration function on a Q-Exactive UHMR instrument.
Supplementary Figure 8 STORI Slope Averaging/Charge Assignment.
Correct assignment of known +20 myoglobin single ions. As a larger number STORI slope values are averaged prior to the assignment of an ion’s charge (top to bottom) the percentage of ions assigned to the correct (+20) charge state increases markedly. Other examples across the range of charge states used in this study give results from 94.0% to 98.5% in the rate of correct charge states for knowns analyzed by I2MS in either denatured or native modes.
Supplementary Figure 9 Myoglobin Example.
The ensemble spectrum in the m/z domain (a), I2MS mass spectrum (b), and corresponding individual ion charge assignment histogram (c) for the myoglobin charge state envelope. The insets in the m/z spectrum and I2MS mass spectrum demonstrate that although the charges have been condensed in the I2MS spectrum, the features within each charge state remain unchanged.
Supplementary information
Supplementary Information
Supplementary Figures 1–9, Supplementary Protocol and Supplementary Table 1
Supplementary Software
Software and demo examples
Supplementary Dataset 1
HEK-293 I2 MS Proteoform Assignments; GELFrEE HEK-293 I2MS mass assignments (Fig. 2) with identified proteoform mass, TDReport ID no., PFR no., accession no., modifications and PPM error
Source data
Source Data Fig. 2
Source data for Fig. 2
Source Data Fig. 3
Source data for Fig. 3
Source Data Supplementary Fig. 2
Source data for Supplementary Fig. 2
Source Data Supplementary Fig. 3
Source data for Supplementary Fig.3
Source Data Supplementary Fig. 4
Source data for Supplementary Fig.4
Source Data Supplementary Fig. 9
Source data for Supplementary Fig.9
Rights and permissions
About this article
Cite this article
Kafader, J.O., Melani, R.D., Durbin, K.R. et al. Multiplexed mass spectrometry of individual ions improves measurement of proteoforms and their complexes. Nat Methods 17, 391–394 (2020). https://doi.org/10.1038/s41592-020-0764-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0764-5
This article is cited by
-
Nanopore DNA sequencing technologies and their applications towards single-molecule proteomics
Nature Chemistry (2024)
-
Ultralong transients enhance sensitivity and resolution in Orbitrap-based single-ion mass spectrometry
Nature Methods (2024)
-
A critical evaluation of ultrasensitive single-cell proteomics strategies
Analytical and Bioanalytical Chemistry (2024)
-
Sampling the proteome by emerging single-molecule and mass spectrometry methods
Nature Methods (2023)
-
Automated imaging and identification of proteoforms directly from ovarian cancer tissue
Nature Communications (2023)