Abstract
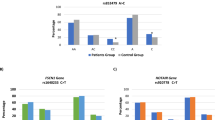
The Hippo pathway participates in development of numerous tumors through regulating tissue growth and cell fate. This study aimed to detect the association between the genetic variants in Hippo pathway genes and bladder cancer risk in a Chinese population. A case–control study of 580 cases and 1101 controls was performed to evaluate the association of single nucleotide polymorphisms (SNPs) in 39 candidate genes involved in the Hippo pathway with bladder cancer risk. A logistic regression model was used to assess the effects of SNPs on bladder cancer susceptibility. Candidate gene expression in human bladder cancer samples was detected using The Cancer Genome Atlas (TCGA) database and the Gene Expression Omnibus (GEO) datasets. We found that SNP rs755813 in WWC1 was significantly associated with a decreased risk of bladder cancer [odds ratio (OR) = 0.76, 95% confidence interval (CI) = 0.66–0.88, P = 3.63 × 10–4], which was more common in patients with low grade and non-muscle invasive tumors. Younger subjects (age ≤ 65) (OR = 0.70, 95% CI = 0.56–0.86), females (0.35, 0.23–0.52) and non-smokers (0.72, 0.58–0.88) showed a pronounced association between the rs755813 C allele and risk of bladder cancer by stratified analysis. The WWC1 was upregulated in bladder cancer tissues according to TCGA and GEO datasets. These findings indicated that genetic variant of WWC1 gene in Hippo signaling pathway contributes to the decreased risk of bladder cancer in the Chinese population and may have the protective effect against the development of bladder cancer.


Similar content being viewed by others
References
Al-Zalabani AH, Stewart KF, Wesselius A, Schols AM, Zeegers MP (2016) Modifiable risk factors for the prevention of bladder cancer: a systematic review of meta-analyses. Eur J Epidemiol 31(9):811–851. https://doi.org/10.1007/s10654-016-0138-6
An Y, Zhang Q, Li X, Wang Z, Li Y, Tang X (2018) Upregulated microRNA miR-21 promotes the progression of lung adenocarcinoma through inhibition of KIBRA and the Hippo signaling pathway. Biomed Pharmacother 108:1845–1855. https://doi.org/10.1016/j.biopha.2018.09.125
Chen W, Zheng R, Baade PD et al (2016) Cancer statistics in China, 2016. CA 66(2):115–132. https://doi.org/10.3322/caac.21338
Cucci MA, Compagnone A, Daga M et al (2019) Post-translational inhibition of YAP oncogene expression by 4-hydroxynonenal in bladder cancer cells. Free Radical Biol Med 141:205–219. https://doi.org/10.1016/j.freeradbiomed.2019.06.009
Cumberbatch MGK, Noon AP (2019) Epidemiology, aetiology and screening of bladder cancer. Trans Androl Urol 8(1):5–11. https://doi.org/10.21037/tau.2018.09.11
Cumberbatch MG, Cox A, Teare D, Catto JW (2015) Contemporary occupational carcinogen exposure and bladder cancer: a systematic review and meta-analysis. JAMA Oncol 1(9):1282–1290. https://doi.org/10.1001/jamaoncol.2015.3209
Dong L, Lin F, Wu W, Huang W, Cai Z (2016) Transcriptional cofactor Mask2 is required for YAP-induced cell growth and migration in bladder cancer cell. J Cancer 7(14):2132–2138. https://doi.org/10.7150/jca.16438
Duning K, Schurek EM, Schluter M et al (2008) KIBRA modulates directional migration of podocytes. J Am Soc Nephrol 19(10):1891–1903. https://doi.org/10.1681/ASN.2007080916
Han Y (2019) Analysis of the role of the Hippo pathway in cancer. J Trans Med 17(1):116. https://doi.org/10.1186/s12967-019-1869-4
Harvey KF, Zhang X, Thomas DM (2013) The Hippo pathway and human cancer. Nat Rev Cancer 13(4):246–257. https://doi.org/10.1038/nrc3458
Huang CY, Huang SP, Lin VC et al (2015) Genetic variants in the Hippo pathway predict biochemical recurrence after radical prostatectomy for localized prostate cancer. Sci Rep 5:8556. https://doi.org/10.1038/srep08556
Ke HL, Lin J, Ye Y et al (2015) Genetic variations in glutathione pathway genes predict cancer recurrence in patients treated with transurethral resection and Bacillus Calmette-Guerin instillation for non-muscle invasive bladder cancer. Ann Surg Oncol 22(12):4104–4110. https://doi.org/10.1245/s10434-015-4431-5
Li S, Yu Z, Chen SS et al (2015) The YAP1 oncogene contributes to bladder cancer cell proliferation and migration by regulating the H19 long noncoding RNA. Urol Oncol 33:4271
Lin Y, Ge Y, Wang Y et al (2017) The association of rs710886 in lncRNA PCAT1 with bladder cancer risk in a Chinese population. Gene 627:226–232. https://doi.org/10.1016/j.gene.2017.06.021
Liu JY, Li YH, Lin HX et al (2013) Overexpression of YAP 1 contributes to progressive features and poor prognosis of human urothelial carcinoma of the bladder. BMC Cancer 13:349. https://doi.org/10.1186/1471-2407-13-349
Meng Z, Moroishi T, Guan KL (2016) Mechanisms of Hippo pathway regulation. Genes Dev 30(1):1–17. https://doi.org/10.1101/gad.274027.115
Moleirinho S, Chang N, Sims AH et al (2013) KIBRA exhibits MST-independent functional regulation of the Hippo signaling pathway in mammals. Oncogene 32(14):1821–1830. https://doi.org/10.1038/onc.2012.196
Moroishi T, Hansen CG, Guan KL (2015) The emerging roles of YAP and TAZ in cancer. Nat Rev Cancer 15(2):73–79. https://doi.org/10.1038/nrc3876
Pan D (2010) The hippo signaling pathway in development and cancer. Dev Cell 19(4):491–505. https://doi.org/10.1016/j.devcel.2010.09.011
Schelleckes K, Schmitz B, Ciarimboli G et al (2017) Promoter methylation inhibits expression of tumor suppressor KIBRA in human clear cell renal cell carcinoma. Clin Epigenet 9:109. https://doi.org/10.1186/s13148-017-0415-6
Sebio A, Matsusaka S, Zhang W et al (2016) Germline polymorphisms in genes involved in the Hippo pathway as recurrence biomarkers in stages II/III colon cancer. Pharmacogenom J 16(4):312–319. https://doi.org/10.1038/tpj.2015.64
Siegel RL, Miller KD, Jemal A (2019) Cancer statistics, 2019. CA 69(1):7–34. https://doi.org/10.3322/caac.21551
Stauffer S, Chen X, Zhang L, Chen Y, Dong J (2016) KIBRA promotes prostate cancer cell proliferation and motility. FEBS J 283(10):1800–1811. https://doi.org/10.1111/febs.13718
Wang M, Li Z, Chu H et al (2016) Genome-wide association study of bladder cancer in a Chinese cohort reveals a new susceptibility locus at 5q123. Cancer Res 76(11):3277–3284. https://doi.org/10.1158/0008-5472.CAN-15-2564
Wang S, Huo D, Ogundiran TO et al (2018) Genetic variation in the Hippo pathway and breast cancer risk in women of African ancestry. Mol Carcinog 57(10):1311–1318. https://doi.org/10.1002/mc.22845
Wang Z, Katsaros D, Biglia N et al (2019) Low expression of WWC1, a tumor suppressor gene, is associated with aggressive breast cancer and poor survival outcome. FEBS Open Bio 9(7):1270–1280. https://doi.org/10.1002/2211-5463.12659
Xiao L, Chen Y, Ji M, Dong J (2011) KIBRA regulates Hippo signaling activity via interactions with large tumor suppressor kinases. J Biol Chem 286(10):7788–7796. https://doi.org/10.1074/jbc.M110.173468
Xin J, Du M, Gu D et al (2019) Combinations of single nucleotide polymorphisms identified in genome-wide association studies determine risk for colorectal cancer. Int J Cancer. https://doi.org/10.1002/ijc.32267
Yang S, Ji M, Zhang L et al (2014) Phosphorylation of KIBRA by the extracellular signal-regulated kinase (ERK)-ribosomal S6 kinase (RSK) cascade modulates cell proliferation and migration. Cell Signal 26(2):343–351. https://doi.org/10.1016/j.cellsig.2013.11.012
Yoshihama Y, Izumisawa Y, Akimoto K et al (2013) High expression of KIBRA in low atypical protein kinase C-expressing gastric cancer correlates with lymphatic invasion and poor prognosis. Cancer Sci 104(2):259–265. https://doi.org/10.1111/cas.12066
Yu J, Zheng Y, Dong J, Klusza S, Deng WM, Pan D (2010) Kibra functions as a tumor suppressor protein that regulates Hippo signaling in conjunction with Merlin and Expanded. Dev Cell 18(2):288–299. https://doi.org/10.1016/j.devcel.2009.12.012
Yu FX, Zhao B, Guan KL (2015) Hippo pathway in organ size control, tissue homeostasis, and cancer. Cell 163(4):811–828. https://doi.org/10.1016/j.cell.2015.10.044
Yuan H, Liu H, Liu Z et al (2015) Genetic variants in Hippo pathway genes YAP1, TEAD1 and TEAD4 are associated with melanoma-specific survival. Int J Cancer 137(3):638–645. https://doi.org/10.1002/ijc.29429
Zhang J, Yao S, Hu Q et al (2016) Genetic variations in the Hippo signaling pathway and breast cancer risk in African American women in the AMBER consortium. Carcinogenesis 37(10):951–956. https://doi.org/10.1093/carcin/bgw077
Zhao R, Liu K, Huang Z et al (2015) Genetic variants in caveolin-1 and RhoA/ROCK1 are associated with clear cell renal cell carcinoma risk in a Chinese population. PLoS ONE 10(6):e0128771. https://doi.org/10.1371/journal.pone.0128771
Acknowledgements
This study was supported in part by the National Natural Science Foundation of China (81872691), Collaborative Innovation Center for Cancer Personalized Medicine, and Priority Academic Program Development of Jiangsu Higher Education Institutions (Public Health and Preventive Medicine).
Author information
Authors and Affiliations
Contributions
ZZ and ZW conceived and designed the study. ZH and XW performed the experiments. ZH, LM, ZG, HL and MD were responsible for data acquisition and data analysis. ZH wrote the paper. MW, HC and ZZ critically revised the manuscript. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
The study was approved by the institution review board of Nanjing Medical University. The research protocol was performed in accordance with the approved guidelines and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Huang, Z., Wang, X., Ma, L. et al. Genetic variations in Hippo pathway genes influence bladder cancer risk in a Chinese population. Arch Toxicol 94, 785–794 (2020). https://doi.org/10.1007/s00204-020-02663-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-020-02663-z