Abstract
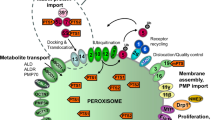
Peroxisomes are ubiquitous organelles formed by peroxisome biogenesis (PB). During PB, peroxisomal matrix proteins harboring a peroxisome targeting signal (PTS) are imported inside peroxisomes by peroxins, encoded by PEX genes. Genetic alterations in PEX genes lead to a spectrum of incurable diseases called Zellweger spectrum disorders (ZSD). In vitro drug screening is part of the quest for a cure in ZSD by restoring PB in ZSD cell models. In vitro PB evaluation is commonly achieved by immunofluorescent staining or transient peroxisome fluorescent reporter expression. Both techniques have several drawbacks (cost, time-consuming technique, etc.) which we overcame by developing a third-generation lentiviral transfer plasmid expressing an enhanced green fluorescent protein fused to PTS1 (eGFP–PTS1). By eGFP–PTS1 lentiviral transduction, we quantified PB and peroxisome motility in ZSD and control mouse and human fibroblasts. We confirmed the stable eGFP–PTS1 expression along cell passages. eGFP signal analysis distinguished ZSD from control eGFP–PTS1-transduced cells. Live eGFP–PTS1 transduced cells imaging quantified peroxisomes motility. In conclusion, we developed a lentiviral transfer plasmid allowing stable eGFP–PTS1 expression to study PB (deposited on Addgene: #133282). This tool meets the needs for in vitro PB evaluation and ZSD drug discovery.





Similar content being viewed by others
References
Argyriou C, D’Agostino MD, Braverman N (2016) Peroxisome biogenesis disorders. Transl Sci Rare Dis 1:111–144. https://doi.org/10.3233/trd-160003
Argyriou C, Polosa A, Cecyre B, Hsieh M, Di Pietro E, Cui W, Bouchard JF, Lachapelle P, Braverman N (2019) A longitudinal study of retinopathy in the PEX1-Gly844Asp mouse model for mild Zellweger Spectrum Disorder. Exp Eye Res. https://doi.org/10.1016/j.exer.2019.107713
Berendse K, Ebberink MS, Ijlst L, Poll-The BT, Wanders RJ, Waterham HR (2013) Arginine improves peroxisome functioning in cells from patients with a mild peroxisome biogenesis disorder. Orphanet J Rare Dis 8:138. https://doi.org/10.1186/1750-1172-8-138
Berendse K, Engelen M, Ferdinandusse S, Majoie CB, Waterham HR, Vaz FM, Koelman JH, Barth PG, Wanders RJ, Poll-The BT (2016) Zellweger spectrum disorders: clinical manifestations in patients surviving into adulthood. J Inherit Metab Dis 39:93–106. https://doi.org/10.1007/s10545-015-9880-2
Berendse K, Boek M, Gijbels M, Van der Wel NN, Klouwer FC, van den Bergh-Weerman MA, Shinde AB, Ofman R, Poll-The BT, Houten SM, Baes M, Wanders RJA, Waterham HR (2019) Liver disease predominates in a mouse model for mild human Zellweger spectrum disorder. Biochim Biophys Acta Mol Basis Dis. https://doi.org/10.1016/j.bbadis.2019.06.013
Brickner DG, Harada JJ, Olsen LJ (1997) Protein transport into higher plant peroxisomes. In vitro import assay provides evidence for receptor involvement. Plant Physiol 113:1213–1221. https://doi.org/10.1104/pp.113.4.1213
Demaret T, Varma S, Stephenne X, Smets F, Scheers I, Wanders R, Van Maldergem L, Reding R, Sokal E (2018) Living-donor liver transplantation for mild Zellweger spectrum disorder: up to 17 years follow-up. Pediatr Transplant 22:e13112
Deosaran E, Larsen KB, Hua R, Sargent G, Wang Y, Kim S, Lamark T, Jauregui M, Law K, Lippincott-Schwartz J, Brech A, Johansen T, Kim PK (2013) NBR1 acts as an autophagy receptor for peroxisomes. J Cell Sci 126:939–952. https://doi.org/10.1242/jcs.114819
Dias AF, Francisco T, Rodrigues TA, Grou CP, Azevedo JE (2016) The first minutes in the life of a peroxisomal matrix protein. Biochem Biophys Acta 1863:814–820. https://doi.org/10.1016/j.bbamcr.2015.09.025
Ebberink MS, Mooijer PA, Gootjes J, Koster J, Wanders RJ, Waterham HR (2011) Genetic classification and mutational spectrum of more than 600 patients with a Zellweger syndrome spectrum disorder. Hum Mutat 32:59–69. https://doi.org/10.1002/humu.21388
Emmanouilidis L, Gopalswamy M, Passon DM, Wilmanns M, Sattler M (2016) Structural biology of the import pathways of peroxisomal matrix proteins. Biochem Biophys Acta 1863:804–813. https://doi.org/10.1016/j.bbamcr.2015.09.034
Farre JC, Mahalingam SS, Proietto M, Subramani S (2019) Peroxisome biogenesis, membrane contact sites, and quality control. EMBO Rep. https://doi.org/10.15252/embr.201846864
Fujiki Y (2016) Peroxisome biogenesis and human peroxisome-deficiency disorders. Proc Jpn Acad Ser B Phys Biol Sci 92:463–477. https://doi.org/10.2183/pjab.92.463
Giros M, Roels F, Prats J, Ruiz M, Ribes A, Espeel M, Wanders RJ, Schutgens RB, Pampols T (1996) Long survival in a case of peroxisomal biogenesis disorder with peroxisome mosaicism in the liver. Ann N Y Acad Sci 804:747–749. https://doi.org/10.1111/j.1749-6632.1996.tb18689.x
Hiebler S, Masuda T, Hacia JG, Moser AB, Faust PL, Liu A, Chowdhury N, Huang N, Lauer A, Bennett J, Watkins PA, Zack DJ, Braverman NE, Raymond GV, Steinberg SJ (2014) The Pex1-G844D mouse: a model for mild human Zellweger spectrum disorder. Mol Genet Metab 111:522–532. https://doi.org/10.1016/j.ymgme.2014.01.008
Hunter M, Yuan P, Vavilala D, Fox M (2019) Optimization of protein expression in mammalian cells. Curr Protoc Protein Sci 95:e77. https://doi.org/10.1002/cpps.77
Klouwer FC, Berendse K, Ferdinandusse S, Wanders RJ, Engelen M, Poll-The BT (2015) Zellweger spectrum disorders: clinical overview and management approach. Orphanet J Rare Dis 10:151. https://doi.org/10.1186/s13023-015-0368-9
Koster J, Waterham HR (2017) Transfection of primary human skin fibroblasts for peroxisomal studies. Methods Mol Biol (Clifton, NJ) 1595:63–67. https://doi.org/10.1007/978-1-4939-6937-1_7
Law KB, Bronte-Tinkew D, Di Pietro E, Snowden A, Jones RO, Moser A, Brumell JH, Braverman N, Kim PK (2017) The peroxisomal AAA ATPase complex prevents pexophagy and development of peroxisome biogenesis disorders. Autophagy 13:868–884. https://doi.org/10.1080/15548627.2017.1291470
MacLean GE, Argyriou C, Di Pietro E, Sun X, Birjandian S, Saberian P, Hacia JG, Braverman NE (2019) Zellweger spectrum disorder patient-derived fibroblasts with the PEX1-Gly843Asp allele recover peroxisome functions in response to flavonoids. J Cell Biochem 120:3243–3258. https://doi.org/10.1002/jcb.27591
Mandel H, Espeel M, Roels F, Sofer N, Luder A, Iancu TC, Aizin A, Berant M, Wanders RJ, Schutgens RB (1994) A new type of peroxisomal disorder with variable expression in liver and fibroblasts. J Pediatr 125:549–555. https://doi.org/10.1016/s0022-3476(94)70006-0
Marcassa E, Kallinos A, Jardine J, Rusilowicz-Jones EV, Martinez A, Kuehl S, Islinger M, Clague MJ, Urbe S (2018) Dual role of USP30 in controlling basal pexophagy and mitophagy. EMBO Rep. https://doi.org/10.15252/embr.201745595
Mast FD, Jamakhandi A, Saleem RA, Dilworth DJ, Rogers RS, Rachubinski RA, Aitchison JD (2016) Peroxins Pex30 and Pex29 dynamically associate with reticulons to regulate peroxisome biogenesis from the endoplasmic reticulum. J Biol Chem 291:15408–15427. https://doi.org/10.1074/jbc.m116.728154
Matsunami M, Shimozawa N, Fukuda A, Kumagai T, Kubota M, Chong PF, Kasahara M (2016) Living-donor liver transplantation from a heterozygous parent for infantile refsum disease. Pediatrics. https://doi.org/10.1542/peds.2015-3102
Metz J, Castro IG, Schrader M (2017) Peroxisome motility measurement and quantification assay. Bio-protocol. https://doi.org/10.21769/bioprotoc.2536
Neuhaus A, Eggeling C, Erdmann R, Schliebs W (2016) Why do peroxisomes associate with the cytoskeleton? Biochem Biophys Acta 1863:1019–1026. https://doi.org/10.1016/j.bbamcr.2015.11.022
Santos MJ, Imanaka T, Shio H, Small GM, Lazarow PB (1988) Peroxisomal membrane ghosts in Zellweger syndrome—aberrant organelle assembly. Science 239:1536–1538. https://doi.org/10.1126/science.3281254
Schrader TA, Islinger M, Schrader M (2017) Detection and immunolabeling of peroxisomal proteins. Methods Mol Biol (Clifton, NJ) 1595:113–130. https://doi.org/10.1007/978-1-4939-6937-1_12
Sexton JZ, He Q, Forsberg LJ, Brenman JE (2010) High content screening for non-classical peroxisome proliferators. Int J High Throughput Screen 2010:127–140. https://doi.org/10.2147/ijhts.s10547
Sokal EM, Smets F, Bourgois A, Van Maldergem L, Buts JP, Reding R, Bernard Otte J, Evrard V, Latinne D, Vincent MF, Moser A, Soriano HE (2003) Hepatocyte transplantation in a 4-year-old girl with peroxisomal biogenesis disease: technique, safety, and metabolic follow-up. Transplantation 76:735–738. https://doi.org/10.1097/01.tp.0000077420.81365.53
Soliman K, Gottfert F, Rosewich H, Thoms S, Gartner J (2018) Super-resolution imaging reveals the sub-diffraction phenotype of Zellweger Syndrome ghosts and wild-type peroxisomes. Sci Rep 8:7809. https://doi.org/10.1038/s41598-018-24119-2
Tinevez JY, Perry N, Schindelin J, Hoopes GM, Reynolds GD, Laplantine E, Bednarek SY, Shorte SL, Eliceiri KW (2017) TrackMate: an open and extensible platform for single-particle tracking. Methods (San Diego, Calif) 115:80–90. https://doi.org/10.1016/j.ymeth.2016.09.016
Tymms MJ, Kola I (2001) Gene knockout protocols. Methods in molecular biology, vol 158. Humana Press Inc., Totowa
Van Maldergem L, Moser AB, Vincent MF, Roland D, Reding R, Otte JB, Wanders RJ, Sokal E (2005) Orthotopic liver transplantation from a living-related donor in an infant with a peroxisome biogenesis defect of the infantile Refsum disease type. J Inherit Metab Dis 28:593–600. https://doi.org/10.1007/s10545-005-0593-9
Wanders RJ (2014) Metabolic functions of peroxisomes in health and disease. Biochimie 98:36–44. https://doi.org/10.1016/j.biochi.2013.08.022
Wei H, Kemp S, McGuinness MC, Moser AB, Smith KD (2000) Pharmacological induction of peroxisomes in peroxisome biogenesis disorders. Ann Neurol 47:286–296
Zhang R, Chen L, Jiralerspong S, Snowden A, Steinberg S, Braverman N (2010) Recovery of PEX1-Gly843Asp peroxisome dysfunction by small-molecule compounds. Proc Natl Acad Sci U S A 107:5569–5574. https://doi.org/10.1073/pnas.0914960107
Acknowledgments
We acknowledge the patients for their implication in the study, Caroline Bouzin for the help with immunofluorescent staining, Caroline Daems for her great cloning tips, Valery Payen for the useful discussions about lentivirus production, and Julie Vandewalle for the proofreading. pEGFP-C1 + SKL was a gift from Jay Brenman (Addgene plasmid #53450), pRRLSIN.cPPT.PGK-GFP.WPRE, pMDLg/pRRE, and pRSV-Rev were a gift from Didier Trono (Addgene plasmid #12252, #12251, and #12253), and pCMV-VSV-G was a gift from Bob Weinberg (Addgene plasmid #8454); we thank them for their contributions. We are thankful to the reviewers for their relevant remarks and suggestions.
Funding
Tanguy DEMARET is FRIA Grant Holder of the Fonds de la Recherche Scientifique-FNRS.
Author information
Authors and Affiliations
Contributions
Conceptualization: TD, MN, and ES. Methodology: TD, GC, JR, and PVDS. Software: TD, GC, and PVDS. Validation: TD and JR. Formal analysis: TD. Data curation: TD. Writing—original draft preparation: TD. Writing—review and editing: TD, GC, JR, PVDS, MN, and ES. Supervision: MN and ES. Funding acquisition: TD and ES.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional research committee (Comission d’Ethique Hospitalo-Facultaire, Cliniques Universitaires Saint-Luc, F/2005/04) and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. All procedures performed in studies involving animals were in accordance with the ethical standards of the institution or practice at which the studies were conducted (Ethical Committee for Animal Experimentation at the Health Science Sector, UCLouvain, 2017/UCL/MD/006).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 2 (MP4 15358 kb)
Supplementary material 3 (MP4 11346 kb)
Rights and permissions
About this article
Cite this article
Demaret, T., Courtoy, G.E., Ravau, J. et al. Accurate and live peroxisome biogenesis evaluation achieved by lentiviral expression of a green fluorescent protein fused to a peroxisome targeting signal 1. Histochem Cell Biol 153, 295–306 (2020). https://doi.org/10.1007/s00418-020-01855-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00418-020-01855-z