Abstract
Introduction
Fish inhabiting different aquatic habitats adapts to the environment by metabolomic readjustments. Understanding the combined activities of all the metabolic pathways (metabolome) helps in better understanding the complex interactions between gene and environment.
Objectives
The anadromous migratory Tenualosa ilisha is a high value food fish comprising the dominant fishery of the rivers Padma and Hooghly. The present study aimed at understanding the influence of the two habitats on the nutritional composition of hilsa.
Methods
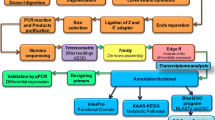
Metabolite profiling was carried out by GC/MS. De novo assembly of hilsa liver transcriptome was generated under Illumina HiSeq platform and multivariate analysis was employed for correlation and comparison.
Results
GC/MS fingerprinting showed C16:0, C18:1, C20:5 and C22:6 to be the predominant fatty acids present in hilsa liver, which were also found to be significantly higher in Hooghly hilsa. Comparative transcriptome analysis revealed that the differentially expressed genes were mainly associated with ‘lipid metabolism’ and ‘amino acid metabolism’ pathways. Multivariate analysis between the metabolites amino acid, fatty acid and corresponding gene expression showed that few genes of amino acid metabolism (EZH1, ALAS2 and ALDH4A1) positively correlated with individual amino acids (lysine, glycine and glutamate) in Hooghly hilsa. Similarly, the key genes for LC-PUFA biosynthesis (ELOVL5, FADS2, CPT1) showed positive correlation with individual LC-PUFAs (C18:3, C20:4, C20:5, C22:6), indicating higher LC-PUFA biosynthesis potential in Hooghly hilsa.
Conclusion
Comparative metabolomic study in hilsa from the two different habitats showed that the habitats influence the nutritional composition as evidenced by high abundance of amino acids lysine, leucine and arginine and LC-PUFAs C18:3, C20:4, C20:5, C22:6 in Hooghly hilsa.



Similar content being viewed by others
Data availability
The transcriptomic and metadata reported in this publication have been deposited in NCBI’s Sequence Read Archive and are accessible through SRA Accession Number SRR5936610 (https://www.ncbi.nlm.nih.gov/sra/?term=SRR5936610).
References
Agwu, D. E., McPhail, L. C., Sozzani, S., Bass, D. A., & McCall, C. E. (1991). Phosphatidic acid as a second messenger in human polymorphonuclear leukocytes. Effects on activation of NADPH oxidase. Journal of Clinical Investigation,88, 531–539.
Alam, A. K. M. N., Mohanty, B. P., Hoq, M. E., & Thilsted, S. H. (2012). Nutritional values, consumption and utilization of hilsa Tenualosa ilisha (Ham 1822). In Proceedings of regional workshop of hilsa: Potential for aquaculture (pp. 188–215). 16–17 September 2012, Dhaka, Bangladesh.
Albers, J. J., Vuletic, S., & Cheung, M. C. (2012). Role of plasma phospholipid transfer protein in lipid and lipoprotein metabolism. Biochimica et Biophysica Acta,1821, 345–357.
Altschul, S. F., Madden, T. L., SchaÈffer, A. A., Zhang, J., Zhang, Z., & Miller, W. (1997). Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Research,25, 3389–3402.
Anders, S., & Huber, W. (2015). Differential expression analysis for sequence count data. Genome Biology,11, 1–12.
Benjamini, Y., & Yekutieli, D. (2005). False discovery rate-adjusted multiple confidence intervals for selected parameters. Journal of the American Statistical Association,100, 71–81.
Betancor, M. B., Olsen, R. E., Solstorm, D., Skulstad, O. F., & Tocher, D. R. (2016). Assessment of a land-locked Atlantic salmon (Salmo salar L.) population as a potential genetic resource with a focus on long-chain polyunsaturated fatty acid biosynthesis. Biochimica et Biophysica Acta (BBA)—Molecular and Cell Biology of Lipids,1861, 227–238.
Bustin, S. A., Benes, V., Garson, J. A., Hellemans, J., Huggett, J., Kubista, M., et al. (2009). The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clinical Chemistry,55, 611–622.
Canovas, A., Quintanilla, R., Amills, M., & Pena, R. (2010). Muscle transcriptomics profiles in pigs with divergent phenotypes for fatness traits. BMC Genomics,11, 372.
Castro, L. F. C., Tocher, D. R., & Monroig, O. (2016). Long-chain polyunsaturated fatty acid biosynthesis in chordates: Insights into the evolution of Fads and Elovl gene repertoire. Progress in Lipid Research,62, 25–40.
Di Bartolomeo, F., Wagner, A., & Daum, G. (2017). Cell biology, physiology and enzymology of phosphatidylserine decarboxylase. Biochimica et Biophysica Acta (BBA)—Molecular and Cell Biology of Lipids,1862, 25–38.
Divya, B. K., Mohindra, V., Singh, R. K., & Yadav, P. (2019). Muscle transcriptome resource for growth, lipid metabolism and immune system in hilsa shad, Tenualosa ilisha. Genes Genomics,41(1), 1–15.
Du, F., Xu, G., Nie, Z., Xu, P., & Gu, R. (2014). Transcriptome analysis gene expression in the liver of Coila nasus during the stress response. BMC Genomics,15, 558.
Fang, C. H., Chang, Y. J., Chung, W. C., Hsieh, P. H., Lin, C. Y., & Ho, J. M. (2015). Subset selection of high-depth next generation sequencing reads for de novo genome assembly using MapReduce framework. BMC Genomics,16, S9.
FAO. (2011). Ganges-Brahmaputra-Meghna river basin. Irrigation in southern and eastern Asia in figures. AQUASTAT survey 2011. Retrieved from https://www.fao.org/nr/water/aquastat/basins/gbm/gbm-CP_eng.pdf.
Fergusan, L. R., Philpott, M., & Barnett, M. P. G. (2010). Nutrigenomics: Integrating genomic approaches into nutrition research. In G. P. Patrinos & W. J. Ansorge (Eds.), Molecular Diagnostics (2nd ed., pp. 347–363). New York: Academic Press.
Folch, J., Lees, M., & Sloane-Stanley, G. H. (1957). A simple method for the isolation and purification of total lipids from animal tissues. Journal of Biological Chemistry,226(1), 497–509.
Ganguly, S., Mahanty, A., Mitra, T., Mohanty, S., Das, B. K., & Mohanty, B. P. (2018). Nutrigenomic studies on hilsa to evaluate flesh quality attributes and genes associated with fatty acid metabolism from the rivers Hooghly and Padma. Food Research International,103, 21–29.
Gebrekiros, S. T. (2016). Factors affecting stream fish community composition and habitat suitability. Journal of Aquaculture and Marine Biology,4(2), 00076.
Grabherr, M. G., Haas, B. J., Yassour, M., Levin, J. Z., Thompson, D. A., & Amit, I. (2011). Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nature Biotechnology,29, 644–652.
Gregory, M. K., & James, M. J. (2014). Rainbow trout (Oncorhynchus mykiss) Elovl5 and Elovl2 differ in selectivity for elongation of omega-3 docosapentaenoic acid. Biochimica et Biophysica Acta (BBA)—Molecular and Cell Biology of Lipids,1841, 1656–1660.
Haque, M. A., Jewel, M. A. S., & Sultana, M. P. (2019). Assessment of physicochemical and bacteriological parameters in surface water of Padma River, Bangladesh. Applied Water Science, 9, 10.
Hu, C. A., Lin, W. W., & Valle, D. (1996). Cloning, characterization, and expression of cDNAs encoding human ∆1-pyrroline-5-carboxylate dehydrogenase. The Journal of Biological Chemistry,271(16), 9795–9800.
Huang, Q., Xu, M., Zhang, H., He, D., Kong, Y., Chen, L., et al. (2019). Transcriptome and proteome analyses of the molecular mechanisms associated with coix seed nutritional quality in the process of breeding. Food Chemistry,272, 549–558.
Jump, D. B. (2009). Mammalian fatty acid elongases. Methods in Molecular Biology,579, 375–389.
Kohjima, M., Enjoji, M., Higuchi, N., Kato, M., Kotoh, K., Yoshimoto, T., et al. (2007). Re-evaluation of fatty acid metabolism-related gene expression in nonalcoholic fatty liver disease. International Journal of Molecular Medicine,20, 351–358.
Lagace, T. A., & Ridgway, N. D. (2013). The role of phospholipids in the biological activity and structure of the endoplasmic reticulum. Biochimica et Biophysica Acta,1833, 2499–2510.
Leaver, M. J., Bautista, J. M., Björnsson, B. T., Jönsson, E., Krey, G., Tocher, D. R., et al. (2008). Towards fish lipid nutrigenomics: Current state and prospects for fin-fish aquaculture. Reviews in Fisheries Science,16(1), 73–94.
Liu, X., Strable, M. S., & Ntambi, J. M. (2011). Stearoyl CoA desaturase 1: Role in cellular inflammation and stress. Advances in Nutrition,2, 15–22.
Livak, K. J., & Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods,25(4), 402–408.
Marioni, J. C., Mason, C. E., Mane, S. M., Stephens, M., & Gilad, Y. (2008). RNA-seq: An assessment of technical reproducibility and comparison with gene expression arrays. Genome Research,18, 1509–1517.
Martinez-Botas, J., Anderson, J. B., Tessier, D., Lapillonne, A., Chang, B. H., Quast, M. J., et al. (2000). Absence of perilipin results in leanness and reverses obesity in Leprdb/db mice. Nature Genetics,26, 474–479.
Metcalfe, L. D., Schmitz, A. A., & Petha, J. R. (1966). Rapid preparation of fatty acid esters from lipids for gas chromatography analysis. Analytical Chemistry,38, 514–517.
Mitra, A. (2013). Sensitivity of mangrove ecosystem to changing climate (pp 33). New Delhi: Springer.
Mitra, T., Mahanty, A., Ganguly, S., Purohit, G. K., Mohanty, S., Parida, P. K., et al. (2019). Expression patterns of heat shock protein genes in Rita rita from natural riverine habitat as biomarker response against environmental pollution. Chemosphere,211, 535–546.
Mohanty, B. P., Ganguly, S., Mahanty, A., Sankar, T. V., Anandan, R., Chakrabarty, K., et al. (2016). DHA, EPA content and fatty acid profile of 39 food fishes from India. Biomed Research International. https://doi.org/10.1155/2016/4027437.
Mohanty, B. P., Mahanty, A., Ganguly, S., Mitra, T., Karunakaran, D., & Anandan, R. (2019). Nutritional composition of food fishes and their importance in providing food and nutritional security. Food Chemistry,293, 561–570.
Mohanty, B. P., Paria, P., Mahanty, A., Behera, B. K., Mathew, S., Sankar, T. V., et al. (2012). Fatty acid profile of Indian shad Tenualosa ilisha oil and its dietary significance. National Academy of Science Letter,35, 263–269.
Mohindra, V., Dangi, T., Tripathi, R. K., Kumar, R., Singh, R. K., Jena, J. K., et al. (2019). Draft genome assembly of Tenualosa ilisha, Hilsa shad, provides resource for osmoregulation studies. Scientific Reports,9, 16511.
Mollah, M. B. D., Khan, M. G. Q., Islam, M. S., & Alam, M. S. (2019). First draft genome assembly and identification of SNPs from hilsa shad (Tenualosa ilisha). F1000 Research,8, 320.
Monroig, Ó., Li, Y., & Tocher, D. R. (2011). Delta-8 desaturation activity varies among fatty acyl desaturases of teleost fish: High activity in delta-6 desaturases of marine species. Comparative Biochemistry and Physiology Part-B Biochemistry and Molecular Biology,159, 206–213.
Monroig, O., Lopes-Marques, M., Navarro, J. C., Hontoria, F., Ruivo, R., Santos, M. M., et al. (2016). Evolutionary functional elaboration of the Elovl2/5 gene family in chordates. Scientific Report,6, 20510.
Monroig, O., Rotllant, J., Sanchez, E., Cerda-Reverter, J. M., & Tocher, D. R. (2009). Expression of long-chain polyunsaturated fatty acid (LC-PUFA) biosynthesis genes during zebrafish Danio rerio early embryogenesis. Biochimica et Biophysica Acta (BBA)—Molecular and Cell Biology of Lipids,1791, 1093–1101.
Morais, S., Monroig, O., Zheng, X., Leaver, M. J., & Tocher, D. R. (2009). Highly unsaturated fatty acid synthesis in Atlantic salmon: characterization of ELOVL5- and ELOVL2-like elongases. Marine Biotechnology,11, 627–639.
Morais, S., Taggart, J. B., Guy, D. R., Bell, J. G., & Tocher, D. R. (2012). Hepatic transcriptome analysis of inter-family variability in flesh n-3 long-chain polyunsaturated fatty acid content in Atlantic salmon. BMC Genomics,13, 410.
Ohanian, J., & Ohanian, V. (2001). Lipid second messenger regulation: The role of diacylglycerol kinases and their relevance to hypertension. Journal of Human Hypertension,15, 93–98.
Overturf, K., Barrows, F. T., Hardy, R. W., Brezas, A., & Dumas, A. (2016). Energy composition of diet affects muscle fiber recruitment, body composition, and growth trajectory in rainbow trout (Oncorhynchus mykiss). Aquaculture,457, 1–14.
Ozsolak, F., & Milos, P. M. (2011). RNA sequencing: Advances, challenges and opportunities. Nature Review Genetics,12(2), 87–98.
Panserat, S., & Kaushik, S. J. (2010). Regulation of gene expression by nutritional factors in fish. Aquaculture Research,41, 751–762.
Putney, J. W., & Bird, G. S. J. (1993). The inositol phosphate-calcium signaling system in non excitable cells. Endocrinology Review,14, 610–631.
Rahman, M. J., Wahab, M. A., Amin, S. M. N., & Romano, M. N. N. (2018). Catch trend and stock assessment of hilsa Tenualosa ilisha using digital image measured length-frequency data. Marina and Coastal Fisheries,10(4), 386–401.
Roberts, L. D., Souza, A. L., Gerszten, R. E., & Clish, C. B. (2012). Targeted Metabolomics.,98(1), 1–24.
Rosith, C. M., Meena, D. K., Manna, R. K., Sahoo, A. K., Swain, H. S., Raman, R. K., et al. (2018). Phytoplankton community structure of the Gangetic (Hooghly-Matla) estuary: Status and ecological implications in relation to eco-climatic variability. Flora,240, 133–143.
Rozen, S., & Skaletsky, H. (2000). Primer 3 on the WWW for general users and for biologists programmers. In S. Krawetz & S. Misener (Eds.), Bioinformatics methods and protocols: Methods in molecular biology (pp. 365–386). Totowa, NJ: Humana Press.
Sahoo, A. K., Wahab, M. A., Phillips, M., Rahman, A., Padiyar, A., Puvanendran, V., et al. (2018). Breeding and culture status of Hilsa (Tenualosa ilisha, Ham. 1822) in South Asia: A review. Reviews in Aquaculture,10(1), 96–110.
Shetty, H. P. C., & Saha, S. B. (1971). On the significance of the occurrence of blooms of the diatom Hemidiscus hardmannianus (Greville) mann in relation to hilsa fishery in Bengal. Current Science,15, 410–411.
Shoolingin-Jordan, P. M., Al-Daihan, S., Alexeev, D., Baxter, R. L., Bottomley, S. S., Kahari, I. D., et al. (2003). 5-Aminolevulinic acid synthase: Mechanism, mutations and medicine. Biochimica et Biophysica Acta,1647, 361–366.
Sinclair, G. M., O’Brien, A. L., Keough, M., De Souza, D. P., Dayalan, S., Kanojia, K., et al. (2019). Using metabolomics to assess the sub-lethal effects of zinc and boscalid on an estuarine polychaete worm over time. Metabolomics,15, 108.
Tansey, J., Sztalryd, C., Hlavin, E., Kimmel, A., & Londos, C. (2004). The central role of perilipin A in lipid metabolism and adipocyte lipolysis. IUBMB Life,56, 379–385.
Venkatesan, J., Lowe, B., & Kim, S. K. (2015). Application of diatom biosilica in drug delivery. In S. K. Kim (Ed.), Handbook of marine microalgae (pp. 245–255). New York: Academic Press.
Wang, H., Falck, J. R., Hall, T. M., & Shears, S. B. (2011). Structural basis for an inositol pyrophosphate kinase surmounting phosphate crowding. Nature Chemical Biology,8, 111–116.
Wedow, J. M., Yendrek, C. R., Mello, T. R., Creste, S., Martinez, C. A., & Ainsworth, E. A. (2019). Metabolite and transcript profiling of Guinea grass (Panicum maximum Jacq) response to elevated [CO2] and temperature. Metabolomics,15, 51.
Wu, G., & Morris, S. M. (1998). Arginine metabolism: Nitric oxide and beyond. Biochemical Journal,336, 1–17.
Zydlewski, J., & Wilkie, M. P. (2012). Freshwater to seawater transitions in migratory fishes. In S. D. McCormick, A. P. Farrell, & C. J. Brauner (Eds.), Euryhaline Fishes Fish Physiology (Vol. 32, pp. 253–326). New York: Elsevier. https://doi.org/10.1016/B978-0-12-396951-4.00006-2
Acknowledgements
The present study was partly sponsored by Fisheries Science Division, Indian Council of Agricultural Research (ICAR) (ICAR-FSD-OA#3), Ministry of Agriculture and Farmer’s Welfare, Govt. of India, New Delhi, India under the consortium project Outreach Activity on Nutrient profiling and evaluation of fish as a dietary component. The authors are thankful to the Director, CIFRI, Barrackpore for the facilities and to Dr Md. Aftabuddin, Principal Scientist for the laboratory facilities for the qPCR analysis. The authors (SG, TM, AM) are thankful to ICAR for the research fellowship. Technical assistance received from Shri Ladu Ram Mahaver, Asim Kumar Jana, Samir Kumar Paul, Rabiul Sk and Sujoy Nath is thankfully acknowledged.
Author information
Authors and Affiliations
Contributions
BPM conceived, designed and guided the study, provided all laboratory facilities and edited the manuscript. SG collected the samples and performed the experiments. TM and SM analyzed the transcriptome data. SG and AM did the GC/MS experiments and analysed the metabolite data. SG prepared the manuscript and all authors read and provided critical feedback on the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
The study along with tissue collection and experimentation was carried out according to the ethical guidelines and adhering to the legal requirements of the study country. The study was approved by the Institute Animal Ethics Committee (IAEC) (CIFRI/IAEC-16-17/05).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ganguly, S., Mitra, T., Mahanty, A. et al. A comparative metabolomics study on anadromous clupeid Tenualosa ilisha for better understanding the influence of habitat on nutritional composition. Metabolomics 16, 30 (2020). https://doi.org/10.1007/s11306-020-01655-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-020-01655-5