Abstract
Introduction
Colorectal cancer (CRC) remains an incurable disease. Previous metabolomic studies show that metabolic signatures in plasma distinguish CRC patients from healthy controls. Chronic enteritis (CE) represents a risk factor for CRC, with a 20 fold greater incidence than in healthy individuals. However, no studies have performed metabolomic profiling to investigate CRC biomarkers in CE.
Objective
Our aims were to identify metabolomic signatures in CRC and CE and to search for blood-derived metabolite biomarkers distinguishing CRC from CE, especially early-stage biomarkers.
Methods
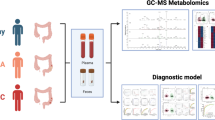
In this case–control study, 612 subjects were prospectively recruited between May 2015 and May 2016, and including 539 CRC patients (stage I, 102 cases; stage II, 259 cases; stage III, 178 cases) and 73 CE patients. Untargeted metabolomics was performed to identify CRC-related metabolic signatures in CE.
Results
Five pathways were significantly enriched based on 153 differential metabolites between CRC and CE. 16 biomarkers were identified for diagnosis of CRC from CE and for guiding CRC staging. The AUC value for CRC diagnosis in the external validation set was 0.85. Good diagnostic performances were also achieved for early-stage CRC (stage I and stage II), with an AUC value of 0.84. The biomarker panel could also stage CRC patients, with an AUC of 0.72 distinguishing stage I from stage II CRC and AUC of 0.74 distinguishing stage II from stage III CRC.
Conclusions
The identified metabolic biomarkers exhibit promising properties for CRC monitoring in CE patients and are superior to commonly used clinical biomarkers (CEA and CA19-9).




Similar content being viewed by others
References
Chan, E. C., et al. (2009). Metabolic profiling of human colorectal cancer using high-resolution magic angle spinning nuclear magnetic resonance (HR-MAS NMR) spectroscopy and gas chromatography mass spectrometry (GC/MS). Journal of Proteome Research,8, 352–361. https://doi.org/10.1021/pr8006232.
Cheng, Y., et al. (2012). Distinct urinary metabolic profile of human colorectal cancer. Journal of Proteome Research,11, 1354–1363. https://doi.org/10.1021/pr201001a.
Cross, A. J., et al. (2014). A prospective study of serum metabolites and colorectal cancer risk. Cancer,120, 3049–3057. https://doi.org/10.1002/cncr.28799.
Derdak, Z., Mark, N. M., Beldi, G., Robson, S. C., Wands, J. R., & Baffy, G. (2008). The mitochondrial uncoupling protein-2 promotes chemoresistance in cancer cells. Cancer Research,68, 2813.
Dowling, P., et al. (2015). Elevated levels of 14–3-3 proteins, serotonin, gamma enolase and pyruvate kinase identified in clinical samples from patients diagnosed with colorectal cancer. Clinica Chimica Acta,441, 133–141. https://doi.org/10.1016/j.cca.2014.12.005.
Edge, S. B., & Compton, C. C. (2010). The American Joint Committee on Cancer: the the AJCC cancer staging manual and the future of TNM. Annals of Surgical Oncology,17, 1471–1474. https://doi.org/10.1245/s10434-010-0985-4.
Farshidfar, F., et al. (2016). A validated metabolomic signature for colorectal cancer: Exploration of the clinical value of metabolomics. British Journal of Cancer,115, 848–857. https://doi.org/10.1038/bjc.2016.243.
Ferlay, J., et al. (2015). Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. International Journal of Cancer,136, E359–E386.
Ikeda, A., et al. (2012). Serum metabolomics as a novel diagnostic approach for gastrointestinal cancer. Biomedical Chromatography,26, 548–558. https://doi.org/10.1002/bmc.1671.
Jimenez, B., et al. (2013). 1H HR-MAS NMR spectroscopy of tumor-induced local metabolic "field-effects" enables colorectal cancer staging and prognostication. Journal of Proteome Research,12, 959–968. https://doi.org/10.1021/pr3010106.
Ke, C., et al. (2015). Large-scale profiling of metabolic dysregulation in ovarian cancer. International Journal of Cancer,136, 516–526.
Kuhl, C., Tautenhahn, R., Bottcher, C., Larson, T. R., & Neumann, S. (2011). CAMERA: An integrated strategy for compound spectra extraction and annotation of liquid chromatography/mass spectrometry data sets. Analytical Chemistry,84, 283–289.
Leichtle, A. B., et al. (2012). Serum amino acid profiles and their alterations in colorectal cancer. Metabolomics,8, 643–653. https://doi.org/10.1007/s11306-011-0357-5.
Li, F., et al. (2013). Lipid profiling for early diagnosis and progression of colorectal cancer using direct-infusion electrospray ionization Fourier transform ion cyclotron resonance mass spectrometry. Rapid Communications in Mass Spectrometry,27, 24–34. https://doi.org/10.1002/rcm.6420.
Liesenfeld, D. B., et al. (2015). Changes in urinary metabolic profiles of colorectal cancer patients enrolled in a prospective cohort study (ColoCare). Metabolomics,11, 998–1012. https://doi.org/10.1007/s11306-014-0758-3.
Liu, S., Zheng, R., Zhang, M., Zhang, S., Sun, X., & Chen, W. (2015). Incidence and mortality of colorectal cancer in China, 2011. Chinese Journal of Cancer Research,27, 22–28.
Ma, Y., Zhang, P., Wang, F., Liu, W., Yang, J., & Qin, H. (2012). An integrated proteomics and metabolomics approach for defining oncofetal biomarkers in the colorectal cancer. Annals of Surgery,255, 720–730. https://doi.org/10.1097/SLA.0b013e31824a9a8b.
Manna, S. K., et al. (2014). Biomarkers of coordinate metabolic reprogramming in colorectal tumors in mice and humans. Gastroenterology,146, 1313–1324. https://doi.org/10.1053/j.gastro.2014.01.017.
Mayerle, J., et al. (2018). Metabolic biomarker signature to differentiate pancreatic ductal adenocarcinoma from chronic pancreatitis. Gut,67, 128–137.
Mazzanti, R., et al. (2006). Differential expression proteomics of human colon cancer. American Journal of Physiology Gastrointestinal and Liver Physiology,290, G1329–G1338.
Meyerhardt, J. A., & Mayer, R. J. (2005). Systemic therapy for colorectal cancer. New England Journal of Medicine,352, 476–487.
Mirnezami, R., et al. (2014). Rapid diagnosis and staging of colorectal cancer via high-resolution magic angle spinning nuclear magnetic resonance (HR-MAS NMR) spectroscopy of intact tissue biopsies. Annals of Surgery,259, 1138–1149. https://doi.org/10.1097/SLA.0b013e31829d5c45.
Ni, Y., Xie, G., & Jia, W. (2014). Metabonomics of human colorectal cancer: New approaches for early diagnosis and biomarker discovery. Journal of Proteome Research,13, 3857–3870.
Nicholson, J. K., & Lindon, J. C. (2008). Systems biology: Metabonomics. Nature,455, 1054.
Nishiumi, S., et al. (2012). A novel serum metabolomics-based diagnostic approach for colorectal cancer. PLoS ONE,7, e40459. https://doi.org/10.1371/journal.pone.0040459.
Patel, D., et al. (2017). Unique and novel urinary metabolomic features in malignant versus benign adrenal neoplasms. Clinical Cancer Research,23, 5302–5310.
Phua, L. C., Chue, X. P., Koh, P. K., Cheah, P. Y., Ho, H. K., & Chan, E. C. (2014). Non-invasive fecal metabonomic detection of colorectal cancer. Cancer Biology & Therapy,15, 389–397. https://doi.org/10.4161/cbt.27625.
Platten, M., Wick, W., & Bj, V. D. E. (2012). Tryptophan catabolism in cancer: Beyond IDO and tryptophan depletion. Cancer Research,72, 5435.
Ritchie, S. A., et al. (2010). Reduced levels of hydroxylated, polyunsaturated ultra long-chain fatty acids in the serum of colorectal cancer patients: Implications for early screening and detection. BMC Medicine,8, 13. https://doi.org/10.1186/1741-7015-8-13.
Shen, X., et al. (2016). Normalization and integration of large-scale metabolomics data using support vector regression. Metabolomics,12, 89.
Silva, C. L., Passos, M., & Camara, J. S. (2011). Investigation of urinary volatile organic metabolites as potential cancer biomarkers by solid-phase microextraction in combination with gas chromatography-mass spectrometry. British Journal of Cancer,105, 1894–1904. https://doi.org/10.1038/bjc.2011.437.
Smith, C. A., Want, E. J., O'Maille, G., Abagyan, R., & Siuzdak, G. (2006). XCMS: Processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Analytical Chemistry,78, 779–787.
Tan, B., et al. (2013). Metabonomics identifies serum metabolite markers of colorectal cancer. Journal of Proteome Research,12, 3000–3009. https://doi.org/10.1021/pr400337b.
Trygg, J., Holmes, E., & Lundstedt, T. (2007). Chemometrics in metabonomics. Journal of Proteome Research,6, 469–479.
Wang, H., Tso, V. K., Slupsky, C. M., & Fedorak, R. N. (2010). Metabolomics and detection of colorectal cancer in humans: A systematic review. Future Oncology,6, 1395–1406.
Wang, H., et al. (2013). (1)H NMR-based metabolic profiling of human rectal cancer tissue. Molecular Cancer,12, 121. https://doi.org/10.1186/1476-4598-12-121.
Wang, C., et al. (2014). Noninvasive detection of colorectal cancer by analysis of exhaled breath. Analytical and Bioanalytical Chemistry,406, 4757–4763. https://doi.org/10.1007/s00216-014-7865-x.
Weedon, D. D., Shorter, R. G., Ilstrup, D. M., Huizenga, K. A., & Taylor, W. F. (1973). Crohn's disease and cancer. New England Journal of Medicine,289, 1099–1103.
Yue, H., et al. (2013). A metabonomics study of colorectal cancer by RRLC-QTOF/MS. Journal of Liquid Chromatography and Related Technologies,36, 428–438.
Zheng, Z.-X., Zheng, R.-S., Zhang, S.-W., & Chen, W.-Q. (2013). Colorectal cancer incidence and mortality in China, 2010. Asian Pacific journal of cancer prevention,15, 8455–8460.
Zhu, J., et al. (2014). Colorectal cancer detection using targeted serum metabolic profiling. Journal of Proteome Research,13, 4120–4130. https://doi.org/10.1021/pr500494u.
Acknowledgements
We thank Zhengjiang Zhu, Prof, from Interdisciplinary Research Center on Biology and Chemistry, Shanghai Institute of Organic Chemistry, Chinese Academy of Sciences, for UHPLC-QTOF/MS analyses. Funding for the project was provided by the National Natural Science Foundation of China (81773551), Harbin Medical University Innovation Fund (31041180075).
Author information
Authors and Affiliations
Contributions
KL, FZ and PH designed the research and wrote the manuscript. CL, WWZ, KY, CYY, ZWR, LC, YXL and YH collected the sample and data. KD analyzed the data. ZZW conducted the metabolomics research.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the Ethical Standards of the Institutional and/or National Research Committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, F., Li, C., Deng, K. et al. Metabolic phenotyping to monitor chronic enteritis canceration. Metabolomics 16, 29 (2020). https://doi.org/10.1007/s11306-020-1651-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-020-1651-x