Abstract
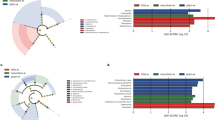
Bacterial contamination of corn-based ethanol biorefineries can reduce their efficiency and hence increase their carbon footprint. To enhance our understanding of these bacterial contaminants, we temporally sampled four biorefineries in the Midwestern USA that suffered from chronic contamination and characterized their microbiomes using both 16S rRNA sequencing and shotgun metagenomics. These microbiotas were determined to be relatively simple, with 13 operational taxonomic units (OTUs) accounting for 90% of the bacterial population. They were dominated by Firmicutes (89%), with Lactobacillus comprising 80% of the OTUs from this phylum. Shotgun metagenomics confirmed our 16S rRNA data and allowed us to characterize bacterial succession at the species level, with the results of this analysis being that Lb. helveticus was the dominant contaminant in this fermentation. Taken together, these results provide insights into the microbiome of ethanol biorefineries and identifies a species likely to be commonly responsible for chronic contamination of these facilities.









Similar content being viewed by others
References
Beckner M, Ivey ML, Phister TG (2011) Microbial contamination of fuel ethanol fermentations. Lett Appl Microbiol 53:387–394. https://doi.org/10.1111/j.1472-765X.2011.03124.x
Bischoff KM, Skinner-Nemec KA, Leathers TD (2007) Antimicrobial susceptibility of Lactobacillus species isolated from commercial ethanol plants. J Ind Microbiol Biotechnol 34(11):739–744. https://doi.org/10.1007/s10295-007-0250-4
Bischoff KM, Liu S, Leathers TD, Worthington RE, Rich JO (2009) Modeling bacterial contamination of fuel ethanol fermentation. Biotechnol Bioeng 103:117–122. https://doi.org/10.1002/bit.22244
Bokulich NA, Mills DA (2012) Next-generation approaches to the microbial ecology of food fermentations. BMB Rep 45:377–389. https://doi.org/10.5483/BMBRep.2012.45.7.148
Bokulich NA, Bamforth CW, Mills DA (2012) Brewhouse-resident microbiota are responsible for multi-stage fermentation of American coolship ale. PLoS One 7:e35507. https://doi.org/10.1371/journal.pone.0035507
Bokulich NA, Joseph CML, Allen G, Benson AK, Mills DA (2013) Correction: Next-generation sequencing reveals significant bacterial diversity of botrytized wine. PLoS One. https://doi.org/10.1371/annotation/4d347090-bca1-4184-a5d5-bc2d38baec7d
Bonatelli ML, Quecine MC, Silva MS, Labate CA (2017) Characterization of the contaminant bacterial communities in sugarcane first-generation industrial ethanol production. FEMS Microbiol Lett. https://doi.org/10.1093/femsle/fnx159
Carvalho-Netto OV, Carazzolle MF, Mofatto LS, Teixeira PJPL, Noronha MF, Calderón LAL, Mieczkowski PA, Argueso LL, Pereira GAG (2015) Saccharomyces cerevisiae transcriptional reprograming due to bacterial contamination during industrial scale bioethanol production. Microb Cell Fact 14:1–13. https://doi.org/10.1186/s12934-015-0196-6
Chao A (1984) Nonparametric estimation of the number of classes in a population. Scand J Stat 11:265–270
Cherubin RA (2003) Efeitos da viabilidade da levedura e da contaminação bacteriana na fermentação alcoólica. Universidade de São Paulo
Daniel JS, Dowe N, Ibsen KN, Riley CJ, Ruth MF, Lumpkin RE (2007) Contaminant occurrence, identification and control in a pilot-scale corn fiber to ethanol conversion process. Bioresour Technol 98:2942–2948. https://doi.org/10.1016/j.biortech.2006.10.002
De Filippis F, Parente E, Ercolini D (2017) Metagenomics insights into food fermentations. Microb Biotechnol 10:91–102. https://doi.org/10.1111/1751-7915.12421
Devantier R, Pedersen S, Olsson L (2005) Characterization of very high gravity ethanol fermentation of corn mash. Effect of glucoamylase dosage, pre-saccharification and yeast strain. Appl Microbiol Biotechnol 68:622–629. https://doi.org/10.1007/s00253-005-1902-9
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Flugge MJ, Lewandrowski J, Rosenfeld J, Boland C, Hendrickson T, Jaglo K, Kolansky S, Moffroid K, Riley-Gilbert M, Pape D (2017) A life-cycle analysis of the greenhouse gas emissions of corn-based ethanol. Report prepared by ICF under USDA Contract No. AG-34142-D-16-0243
Gibson BR, Lawrence SJ, Leclaire JPR, Powell CD, Smart KA (2007) Yeast responses to stresses associated with industrial brewery handling. FEMS Microbiol Rev 31:535–569. https://doi.org/10.1111/j.1574-6976.2007.00076.x
Hamady M, Knight R (2009) Microbial community profiling for human microbiome projects: Tools, techniques, and challenges. Genome Res 19:1141–1152. https://doi.org/10.1101/gr.085464.108
Illumina (2017) bcl2fastq2 Conversion v2.19. In: Illumina, User Guid. 15051736, v2. https://usermanual.wiki/Document/bcl2fastq2guide15051736v2.164202932/view. Accessed 10 Feb 2019
Illumina I (2017) Calculating percent passing filter for patterned and nonpatterned flow cells. In: Illumina, Tech. Note Informatics. chrome-extension://oemmndcbldboiebfnladdacbdfmadadm/https://www.illumina.com/content/dam/illumina-marketing/documents/products/technotes/hiseq-x-percent-pf-technical-note-770-2014-043.pdf. Accessed 10 Feb 2019
Illumina (2018) HiSeq X System Guide. In: Illumina, Hiseq x Syst. Guid. (15050091), v07. http://www.illumina.com/systems/hiseq-x-sequencing-system/performance-specifications.html. Accessed 10 Feb 2019
Kchouk M, Gibrat JF, Elloumi M (2017) Generations of sequencing technologies: From first to next generation. Biol Med 09:8. https://doi.org/10.4172/0974-8369.1000395
Khatibi PA, Roach DR, Donovan DM, Hughes SR, Bischoff KM (2014) Saccharomyces cerevisiae expressing bacteriophage endolysins reduce Lactobacillus contamination during fermentation. Biotechnol Biofuels 7:1–13. https://doi.org/10.1186/1754-6834-7-104
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the miseq illumina sequencing platform. Appl Environ Microbiol 79:5112–5120. https://doi.org/10.1128/AEM.01043-13
Kultima JR, Coelho LP, Forslund K, Huerta-Cepas J, Li SS, Driessen M, Voigt AY, Zeller G, Sunagawa S, Bork P (2016) MOCAT2: a metagenomic assembly, annotation and profiling framework. Bioinformatics 32:2520–2523. https://doi.org/10.1093/bioinformatics/btw183
Leathers TD, Bischoff KM, Rich JO, Price NPJ, Manitchotpisit P, Nunnally MS, Anderson AM (2014) Inhibitors of biofilm formation by biofuel fermentation contaminants. Bioresour Technol 169:45–51. https://doi.org/10.1016/j.biortech.2014.06.065
Leite IR, Faria JR, Marquez LDS, Reis MHM, De Resende MM, Ribeiro EJ, Cardoso VL (2013) Evaluation of hop extract as a natural antibacterial agent in contaminated fuel ethanol fermentations. Fuel Process Technol 106:611–618. https://doi.org/10.1016/j.fuproc.2012.09.050
Li Q, Heist EP, Moe LA (2016) Bacterial community structure and dynamics during corn-based bioethanol fermentation. Microb Ecol 71:409–421. https://doi.org/10.1007/s00248-015-0673-9
Lucena BTL, dos Santos BM, Moreira JL, Moreira APB, Nunes AC, Azevedo V, Miyoshi A, Thompson FL, de Morais M (2010) Diversity of lactic acid bacteria of the bioethanol process. BMC Microbiol. https://doi.org/10.1186/1471-2180-10-298
Montel MC, Buchin S, Mallet A, Delbes-Paus C, Vuitton DA, Desmasures N, Berthier F (2014) Traditional cheeses: rich and diverse microbiota with associated benefits. Int J Food Microbiol 177:136–154. https://doi.org/10.1016/j.ijfoodmicro.2014.02.019
Morris EK, Caruso T, Buscot F, Fischer M, Hancock C, Maier TS, Meiners T, Müller C, Obermaier E, Prati D, Socher SA, Sonnemann I, Wäschke N, Wubet T, Wurst S, Rillig MC (2014) Choosing and using diversity indices: Insights for ecological applications from the German biodiversity exploratories. Ecol Evol 4:3514–3524. https://doi.org/10.1002/ece3.1155
Murphree CA, Heist EP, Moe LA (2014) Antibiotic resistance among cultured bacterial isolates from bioethanol fermentation facilities across the United States. Curr Microbiol 69:277–285. https://doi.org/10.1007/s00284-014-0583-y
Narendranath NV, Thomas KC, Ingledew WM (2001) Effects of acetic acid and lactic acid on the growth of Saccharomyces cerevisiae in a minimal medium. J Ind Microbiol Biotechnol 26:171–177. https://doi.org/10.1038/sj.jim.7000090
Neto PDO, De Lima FA, Da Silva KC, Da Silva DF, Carvalho AFA, Dos Santos C (2014) Chemical inhibition of the contaminant Lactobacillus fermentum from distilleries producing fuel bioethanol. Braz Arch Biol Technol 57:441–447. https://doi.org/10.1590/S1516-8913201401214
Nurk S, Meleshko D, Korobeynikov A, Pevzner PA (2017) metaSPAdes: a new versatile metagenomic assembler. Genome Res 27:824–834. http://www.genome.org/cgi/doi/10.1101/gr.213959.116
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Szoecs E, Wagner H (2018) Vegan: Community ecology package. https://cran.r-project.org/. Accessed 10 Feb 2019
Olivier JGJ, Peters JAHW (2017) Trends in global CO2 and total greenhouses gas emissions. https://www.pbl.nl/en/publications/trends-in-global-co2-and-total-greenhouse-gas-emissions-2018-report. Accessed 10 Feb 2019
Pandya S, Ravi K, Srinivas V, Jadhav S, Khan A, Arun A, Riley LW, Madhivanan P (2017) Comparison of culture-dependent and culture-independent molecular methods for characterization of vaginal microflora. J Med Microbiol 66:149–153. https://doi.org/10.1099/jmm.0.000407
Paulus Compart DM, Carlson AM, Crawford GI, Fink RC, Diez-Gonzalez F, Dicostanzo A, Shurson GC (2013) Presence and biological activity of antibiotics used in fuel ethanol and corn co-product production. J Anim Sci 91:2395–2404. https://doi.org/10.2527/jas.2012-5714
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res 41:590–596. https://doi.org/10.1093/nar/gks1219
R Core Team (2018). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/. Accessed 10 Feb 2019
Rich JO, Leathers TD, Bischoff KM, Anderson AM, Nunnally MS (2015) Biofilm formation and ethanol inhibition by bacterial contaminants of biofuel fermentation. Bioresour Technol 196:347–354. https://doi.org/10.1016/j.biortech.2015.07.071
Rückle L, Senn T (2006) Hop acids can efficiently replace antibiotics in ethanol production. Int Sugar J 108:139–147
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/AEM.01541-09
Takai K, Koki H (2000) Rapid detection and quantification of members of the archael community by quantitative PCR using fluorogenic probes. Appl Environ Microbiol 66:5066–5072. https://doi.org/10.1128/AEM.66.11.5066-5072.2000
Thomas KC, Hynes SH, Ingledew WM (2001) Effect of lactobacilli on yeast growth, viability and batch and semi-continuous alcoholic fermentation of corn mash. Appl Microbiol 90:819–828. https://doi.org/10.1007/s10295-004-0159-0
Tran B, Brown AMK, Bedard PL, Winquist E, Goss GD, Hotte SJ, Welch SA, Hirte HW, Zhang T, Stein LD, Ferretti V, Watt S, Jiao W, Ng K, Ghai S, Shaw P, Petrocelli T, Hudson TJ, Neel BG, Onetto N, Siu LL, McPherson JD, Kamel-Reid S, Dancey JE (2013) Feasibility of real time next generation sequencing of cancer genes linked to drug response: results from a clinical trial. Int J Cancer 132:1547–1555. https://doi.org/10.1002/ijc.27817
Truong DT, Franzosa EA, Tickle TL, Scholz M, Weingart G, Pasolli E, Tett A, Huttenhower C, Segata N (2015) MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat Methods 12:902–903. https://doi.org/10.1002/ijc.27817
Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, Schleifer KH, Whitman WB, Euzéby J, Amann R, Rosselló-Móra R (2014) Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol 12:635–645. https://doi.org/10.1038/nrmicro3330
Zheng J, Ruan L, Sun M, Gänzle M (2015) A genomic view of lactobacilli and pediococci demonstrates that phylogeny matches ecology and physiology. Appl Environ Microbiol 81:7233–7243. https://doi.org/10.1128/AEM.02116-15
Zheng J, Zhao X, Lin XB, Gänzle M (2015) Comparative genomics Lactobacillus reuteri from sourdough reveals adaptation of an intestinal symbiont to food fermentations. Sci Rep 5:18234. https://doi.org/10.1038/srep18234
Acknowledgements
This research was supported by grants from Wisconsin Alumni Research Foundation (WARF) and by Lallemand Inc. We acknowledge the ethanol biorefineries for providing samples, Professor Garret Suen and his team for helping with data analysis, and the analytical team from Mascoma, LLC for all their support during this project. Fernanda Firmino gratefully acknowledges the scholarship from CAPES (Brazil) to pursue her postgraduate studies.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Firmino, F.C., Porcellato, D., Cox, M. et al. Characterization of microbial communities in ethanol biorefineries. J Ind Microbiol Biotechnol 47, 183–195 (2020). https://doi.org/10.1007/s10295-019-02254-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-019-02254-7