Abstract
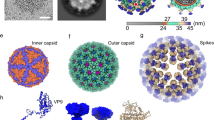
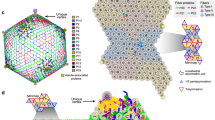
Nervous necrosis virus (NNV) is a non-enveloped virus that causes massive mortality in aquaculture fish production worldwide. Recently X-ray crystallography and single particle cryo-EM have independently determined the icosahedral capsid of NNV to near-atomic resolutions to show the capsid protein is composed of a S-domain (shell) and a P-domain (protrusion) connected by a linker. However, the structure of the spike on NNV capsid made of trimeric P-domains was poorly resolved by cryo-EM. In addition, comparing the spike in the cryo-EM with that by X-ray suggests that the P-domain can move drastically relative to the shell, implicating an underlying structural mechanism during the infectious process. Yet, it remains unclear that such structural re-arrangement is ascribed to the change of the conformation of individual P-domain or in the association among P-domains. Given that molecular structure of the P-domain in solution phase is still lacking, we aim to determine the structure of the P-domain by solution NMR spectroscopy. In this communication, we report backbone and side chain 1H, 13C and 15N chemical shifts of the P-domain (residues 221–338) together with the linker region (residues 214–220), revealing ten β-strands via chemical shift propensity analysis. Our findings are consistent with the X-ray crystal structure of the P-domain reported elsewhere. The current study provides a framework towards further structural analyses of the P-domain in various solution conditions.


Similar content being viewed by others
References
Chen NC, Yoshimura M, Guan HH, Wang TY, Misumi Y, Lin CC, Chuankhayan P, Nakagawa A, Chan SI, Tsukihara T, Chen TY, Chen CJ (2015) Crystal structures of a piscine betanodavirus: mechanisms of capsid assembly and viral infection. PLoS Pathog 11(10):e1005203. https://doi.org/10.1371/journal.ppat.1005203
Doan QK, Vandeputte M, Chatain B, Morin T, Allal F (2017) Viral encephalopathy and retinopathy in aquaculture: a review. J Fish Dis 40(5):717–742. https://doi.org/10.1111/jfd.12541
Ito Y, Okinaka Y, Mori K, Sugaya T, Nishioka T, Oka M, Nakai T (2008) Variable region of betanodavirus RNA2 is sufficient to determine host specificity. Dis Aquat Organ 79(3):199–205. https://doi.org/10.3354/dao01906
Iwamoto T, Okinaka Y, Mise K, Mori K, Arimoto M, Okuno T, Nakai T (2004) Identification of host-specificity determinants in betanodaviruses by using reassortants between striped jack nervous necrosis virus and sevenband grouper nervous necrosis virus. J Virol 78(3):1256–1262. https://doi.org/10.1128/jvi.78.3.1256-1262.2004
Keller R (2004) The computer aided resonance assignment, 1st edn. CANTINA Verlag, Germany
Reverter D, Lima CD (2009) Preparation of SUMO proteases and kinetic analysis using endogenous substrates. Methods Mol Biol (Clifton, NJ) 497:225–239. https://doi.org/10.1007/978-1-59745-566-4_15
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44(4):213–223. https://doi.org/10.1007/s10858-009-9333-z
Souto S, Merour E, Biacchesi S, Bremont M, Olveira JG, Bandin I (2015) In vitro and in vivo characterization of molecular determinants of virulence in reassortant betanodavirus. J Gen Virol 96(Pt 6):1287–1296. https://doi.org/10.1099/vir.0.000064
Wishart DS, Bigam CG, Yao J, Abildgaard F, Dyson HJ, Oldfield E, Markley JL, Sykes BD (1995) 1H, 13C and 15N chemical shift referencing in biomolecular NMR. J Biomol NMR 6(2):135–140. https://doi.org/10.1007/BF00211777
Xie J, Li K, Gao Y, Huang R, Lai Y, Shi Y, Yang S, Zhu G, Zhang Q, He J (2016) Structural analysis and insertion study reveal the ideal sites for surface displaying foreign peptides on a betanodavirus-like particle. Vet Res 47:16. https://doi.org/10.1186/s13567-015-0294-9
Acknowledgements
We gratefully thank Claire Yang (Institute of Chemistry, Academia Sinica) for preparing the manuscript. NMR spectra were collected at the High-field NMR Center (HFNMRC), Academia Sinica, supported by Academia Sinica Core Facility and Innovative Instrument Project (AS-CFII-108-112). This research is funded by Grant MOST 103-2321-B-001-048, 104-2321-B-001-019, 105-2321-B-001-009 (WHC) and 107-2113-M-001-017 (DLT) from the Ministry of Science and Technology of Taiwan.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Štěrbová, P., Wu, D., Lou, YC. et al. NMR assignments of protrusion domain of capsid protein from dragon grouper nervous necrosis virus. Biomol NMR Assign 14, 63–66 (2020). https://doi.org/10.1007/s12104-019-09921-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-019-09921-x