Abstract
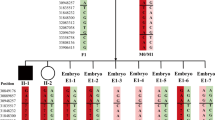
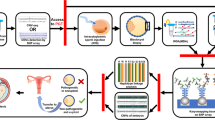
Here, we present the successful application of two different preimplantation genetic testing for monogenic diseases (PGT-M) methods for a couple facing the genetic risk of Achondroplasia (ACH). The first preimplantation genetic haplotyping (PGH) cycle was based on short tandem repeats (STRs) and 8 STRs were chosen. The multiple displacement amplification (MDA) products were analyzed using the informative STR loci and PCR-restriction enzyme digestion of FGFR3. A healthy girl was delivered. Two years later, we performed the second PGT-M cycle for this couple with a newly established PGT-M platform based on next generation sequencing (NGS). Haplotype analysis was established by a selection of several informative single nucleotide polymorphisms (SNPs). Preimplantation genetic testing for aneuploidy (PGT-A) was also performed on embryos with normal FGFR3 genotype. Another healthy girl was born. PGH system could be established using STRs or NGS-SNP systems. The NGS-SNP system could detect more sites and simultaneously performs PGT-A with an automated operation.







Similar content being viewed by others
References
Rousseau, F., Bonaventure, J., Legeai-Mallet, L., Pelet, A., Rozet, J.M., Maroteaux, P., Le Merrer, M. & Munnich, A. Mutations in the gene encoding fibroblast growth factor receptor-3 in achondroplasia. Nature 371, 252–4 (1994).
Shiang, R., Thompson, L.M., Zhu, Y.Z., Church, D.M., Fielder, T.J., Bocian, M., Winokur, S.T. & Wasmuth, J.J. Mutations in the transmembrane domain of FGFR3 cause the most common genetic form of dwarfism, achondroplasia. Cell 78, 335–42 (1994).
Moutou, C., Rongieres, C., Bettahar-Lebugle, K., Gardes, N., Philippe, C. & Viville, S. Preimplantation genetic diagnosis for achondroplasia: genetics and gynaecological limits and difficulties. Hum. Reprod. 18, 509–514 (2003).
Di Rocco, F., Biosse Duplan, M., Heuzé, Y., Kaci, N., Komla-Ebri, D., Munnich, A., Mugniery, E., Benoist-Lasselin, C. & Legeai-Mallet, L. FGFR3 mutation causes abnormal membranous ossification in achondroplasia. Hum. Mol. Genet. 23, 2914–25 (2014).
Altarescu, G., Renbaum, P., Brooks, P.B., Margalioth, E.J., Ben Chetrit, A., Munter, G., Levy-Lahad, E. & Eldar-Geva, T. Successful polar body-based preimplantation genetic diagnosis for achondroplasia. Reprod. BioMed. Online 16, 276–82 (2008).
Dreesen, M., Foulon, V., Vanhaecht, K., De Pourcq, L., Hiele, M., Willems, L. Identifying patient-centered quality indicators for the care of adult home parenteral nutrition (HPN) patients. JPEN J. Parenter Enteral Nutr. 38, 840–6 (2014).
Renwick, P.J., Trussler, J., Ostad-Saffari, E., Fassihi, H., Black, C., Braude, P., Ogilvie, C.M. & Abbs, S. Proof of principle and first cases using preimplantation genetic haplotyping — a paradigm shift for embryo diagnosis. Reprod. BioMed. Online 13, 110–119 (2006).
Altarescu, G., Zeevi, D.A., Zeligson, S., Perlberg, S., Eldar-Geva, T., Margalioth, E.J., Levy-Lahad, E. & Renbaum, P. Familial haplotyping and embryo analysis for Preimplantation genetic diagnosis (PGD) using DNA microarrays: a proof of principle study. J. Assist. Reprod. Genet. 30, 1595–603 (2013).
Yan, L., Huang, L., Xu, L., Huang, J., Ma, F., Zhu, X., Tang, Y., Liu, M., Lian, Y., Liu, P., Li, R., Lu, S., Tang, F., Qiao, J., & Xie, X.S. Live births after simultaneous avoidance of monogenic diseases and chromosome abnormality by next-generation sequencing with linkage analyses. Proc. Natl. Acad. Sci. U.S.A. 112, 15964–9 (2015).
Ren, Y., Zhi, X., Zhu, X., Huang, J., Lian, Y., Li, R., Jin, H., Zhang, Y., Zhang, W., Nie, Y., Wei, Y., Liu, Z., Song, D., Liu, P., Qiao, J. & Yan, L. Clinical applications of MARSALA for preimplantation genetic diagnosis of spinal muscular atrophy. J. Genet. Genomics 43, 541–547 (2016).
Wang, Y., Liu Z., Liu, Z., Zhao, H., Zhou, X., Cui, Y. & Han J. Advances in research on and diagnosis and treatment of achondroplasia in China. Intractable Rare Dis. Res. 2, 45–50 (2013).
Harton, G.L., De Rycke, M., Fiorentino, F., Moutou, C., SenGupta, S., Traeger-Synodinos, J. & Harper, J. C. ESHRE PGD consortium best practice guidelines for amplification-based PGD. Hum. Reprod. 26, 33–40 (2011).
Van der Aa, N., Zamani Esteki, M., Vermeesch, J.R. & Voet, T. Preimplantation genetic diagnosis guided by single-cell genomics. Genome Med. 5, 71 (2013).
Zhou, X., Ren, L., Meng, Q., Li, Y., Yu, Y. & Yu, J. The next-generation sequencing technology and application. Protein Cell 1, 520–36 (2010).
Treff, N.R., Fedick, A., Tao, X., Devkota, B., Taylor, D. & Scott, R.T.Jr. Evaluation of targeted next-generation sequencing-based preimplantation genetic diagnosis of monogenic disease. Fertil. Steril. 99, 1377–1384.e6 (2013).
Chen, L., Diao, Z., Xu, Z., Zhou, J., Yan, G. & Sun, H. The clinical application of NGS-based SNP haplotyping for PGD of Hb H disease. Syst. Biol. Reprod. Med. 63, 212–217 (2017).
Chen, S.C., Xu, X.L., Zhang, J.Y., Ding, G.L., Jin, L., Liu, B., Sun, D.M., Mei, C.L., Yang, X.N., Huang, H.F. & Xu, C.M. Identification of PKD2 mutations in human preimplantation embryos in vitro using a combination of targeted next-generation sequencing and targeted haplotyping. Sci. Rep. 6, 25488 (2016).
Bar-El, L., Kalma, Y., Malcov, M., Schwartz, T., Raviv, S., Cohen, T., Amir, H., Cohen, Y., Reches, A., Amit, A. & Ben-Yosef D. Blastomere biopsy for PGD delays embryo compaction and blastulation: a time-lapse microscopic analysis. J. Assist. Reprod. Genet. 33, 1449–1457 (2016).
Kokkali, G., Traeger-Synodinos, J., Vrettou, C., Stavrou, D., Jones, G.M., Cram, D.S., Makrakis, E., Trounson, A.O., Kanavakis, E. & Pantos, K. Blastocyst biopsy versus cleavage stage biopsy and blastocyst transfer for preimplantation genetic diagnosis of beta-thalassaemia: a pilot study. Hum. Reprod. 22, 1443–9 (2007).
Ling, J.W., Deng, Y., Long, X., Liu, J., Du, H., Cao, B. & Xu, K. Single-nucleotide polymorphism array coupled with multiple displacement amplification: Accuracy and spatial resolution for analysis of chromosome copy numbers in few cells. Biotechnol. Appl. Biochem. 59, 35–44 (2012).
Capalbo, A., Romanelli, V., Cimadomo, D., Girardi, L., Stoppa, M., Dovere, L., Dell’Edera, D., Ubaldi, F.M. & Rienzi, L. Implementing PGD/PGD-A in IVF clinics: considerations for the best laboratory approach and management. J. Assist. Reprod. Genet. 33, 1279–1286 (2016).
Scott, R.T.Jr., Upham, K.M., Forman, E.J., Zhao, T. & Treff, N.R. Cleavage-stage biopsy significantly impairs human embryonic implantation potential while blastocyst biopsy does not: a randomized and paired clinical trial. Fertil. Steril. 100, 624–30 (2013).
Liu, X., Xu, Y., Sun, J., Zhang, Z., Wang, J., Ding, C., Zheng, S.L., Xu, J. & Zhou, C. Preimplantation genetic haplotyping for six Chinese pedigrees with thalassemia using a single nucleotide polymorphism microarray. Prenatal Diagn. 37, 460–468 (2017).
Shen, X., Xu, Y., Zhong, Y., Zhou, C., Zeng, Y., Zhuang, G., Ding, C. & Li, T. Preimplantation genetic diagnosis for alpha-and beta-double thalassemia. J. Assist. Reprod. Genet. 28, 957–64 (2011).
Acknowledgements
The study was supported by National Key Research and Development Program (20 16YFC1000205) and Guangdong Provincial Key Laboratory of Reproductive Medicine (2012A0614000 03).
Author information
Authors and Affiliations
Corresponding author
Additional information
Xiaoting Shen, Dongjia Chen and Yan Xu contributed equally to this work and all should be considered as first authors
Rights and permissions
About this article
Cite this article
Shen, X., Chen, D., Xu, Y. et al. Preimplantation Genetic Testing of Achondroplasia by Two Haplotyping Systems: Short Tandem Repeats and Single Nucleotide Polymorphism. BioChip J 13, 165–173 (2019). https://doi.org/10.1007/s13206-018-3207-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13206-018-3207-y