Abstract
A mixture of multiple scaled generalized hyperbolic distributions (MMSGHDs) is introduced. Then, a coalesced generalized hyperbolic distribution (CGHD) is developed by joining a generalized hyperbolic distribution with a multiple scaled generalized hyperbolic distribution. After detailing the development of the MMSGHDs, which arises via implementation of a multi-dimensional weight function, the density of the mixture of CGHDs is developed. A parameter estimation scheme is developed using the ever-expanding class of MM algorithms and the Bayesian information criterion is used for model selection. The issue of cluster convexity is examined and a special case of the MMSGHDs is developed that is guaranteed to have convex clusters. These approaches are illustrated and compared using simulated and real data. The identifiability of the MMSGHDs and the mixture of CGHDs are discussed in an appendix.
Similar content being viewed by others
1 Introduction
Finite mixture models have been linked with clustering since the idea of defining a cluster in terms of a component in finite mixture model was put forth more than 60 years ago (see McNicholas 2016a, Section 2.1). Nowadays, mixture model–based clustering is a popular approach to clustering. A random vector X arises from a finite mixture model if, for all x ⊂ X, its density can be written
where πg > 0 such that \({\sum }_{g = 1}^{G} \pi _{g} = 1\) are the mixing proportions, fg(x∣θg) is the g th component density, and 𝜗 = (π, θ1, … , θG) denotes the vector of parameters with π = (π1, … , πG). The component densities f1(x∣θ1), … , fG(x∣θG) are typically taken to be of the same type, most commonly multivariate Gaussian. In fact, until a few years after the turn of the century, almost all work on clustering and classification using mixture models had been based on Gaussian mixture models (e.g., Banfield and Raftery 1993; Celeux and Govaert 1995; Celeux and Govaert 1995; Ghahramani and Hinton 1997; Tipping and Bishop 1999; McLachlan and Peel 2000; Fraley and Raftery2002).
Early work on non-Gaussian mixtures was on mixtures of multivariate t-distributions (e.g., Peel and McLachlan 2000). A little beyond the turn of the century, work on t-mixtures burgeoned into a substantial subfield of mixture model-based classification (e.g., McLachlan et al. 2007; Andrews and McNicholas2011a, b, 2012; Baek and McLachlan 2011; Steane et al. 2012; Lin et al. 2014; Pesevski et al. 2018). Around the same time, work on mixtures of skewed distributions took off, including work on skew-normal mixtures (e.g., Lin 2009), skew-t mixtures (e.g., Lin 2010; Vrbik and McNicholas 2012, 2014; Lee and McLachlan 2013a, b; Murray et al. 2014), Laplace mixtures (e.g., Franczak et al.2014), variance-gamma mixtures (McNicholas et al. 2017), generalized hyperbolic mixtures (Browne and McNicholas 2015), and other non-elliptically contoured distributions (e.g., Karlis and Santourian 2009; Murray et al. 2017; Tang et al. 2018). A thorough review of work on model-based clustering is given by McNicholas (2016b).
More recently, mixtures of multiple scaled distributions have been considered (Section 2). In the present manuscript, a multiple scaled generalized hyperbolic distribution is introduced (Section 3). Because the (multivariate) generalized hyperbolic distribution is not a special case of the multiple scaled generalized hyperbolic distribution, a coalesced generalized hyperbolic distribution is also developed (Section 4). The issue of cluster convexity, which has essentially been ignored in work to date on multiple scaled distributions is discussed and a special case of the multiple scaled generalized hyperbolic distribution is developed to guarantee that the components, i.e., the clusters, are convex (Section 5).
2 Multiple Scaled Distributions
The distribution of a p-dimensional random variable X is said to be a normal variance-mean mixture if its density can be written in the form
where ϕp (x∣μ + wα, wΣ) is the density of a p-dimensional Gaussian distribution with mean μ + wα and covariance matrix wΣ, and h (w∣θ) is the density of a univariate random variable W > 0 that has the role of a weight function (see Barndorff-Nielsen et al. 1982; Gneiting1997). This weight function can take on many forms, some of which lead to density representations for well-known non-Gaussian distributions, e.g., if h (w∣θ) is the density of an inverse-gamma random variable with parameters (ν/2, ν/2), then (1) is a representation of the skew-t distribution with ν degrees of freedom (see Demarta and McNeil 2005; Murray et al. 2014). Further details on, and examples of, normal variance-mean mixtures are given by Barndorff-Nielsen (1978), Kotz et al. (2001), and Kotz and Nadarajah (2004), among others.
Now, the density of W > 0 from an inverse-gamma distribution with parameters (α, β) is given by
where Γ(⋅) is the Gamma function. Setting α = β = ν/2 in (2) gives
Setting α = 0 in (1), and using (3) for h (w∣θ), it follows that the density of the multivariate t-distribution with ν degrees of freedom can be written
where δ (x, μ∣Σ) is the squared Mahalanobis distance between x and μ. Forbes and Wraith (2014) show that a multi-dimensional weight variable
can be incorporated into Eq. 1 via an eigen-decomposition of the symmetric positive-definite matrix Σ. Specifically, they set Σ = ΓΦΓ′, where Γ is a p × p matrix of eigenvectors and Φ is a p × p diagonal matrix containing the eigenvalues of Σ. It follows that the density of X becomes
where
is a p-dimensional density such that the random variables W1, … , Wp are independent, i.e., the weights are independent. The density given in Eq. 5 adds flexibility to normal variance-mean mixtures because the parameters θ1, … , θp are free to vary in each dimension. Using the density in Eq. 5, Forbes and Wraith (2014) derive the density of a multiple scaled multivariate-t distribution, Wraith and Forbes (2015) derive a multiple scaled normal-inverse Gaussian distribution, and Franczak et al. (2015) develop a multiple scaled shifted asymmetric Laplace distribution.
Setting Σ = ΓΦΓ′, it follows from Eq. 5 that the density of a multiple scaled analogue of Eq. 4 can be written
where \(\boldsymbol {\Delta }_{\mathbf {W}} = \text {diag}\left (w_{1}^{-1},\dots ,w_{p}^{-1}\right )\) and the weight function
is a p-dimensional Gamma density, where h (wj∣νj/2, νj/2) is given by Eq. 3. Note that the scaled Gaussian density in Eq. 6 can be written
where \(\phi _{1}\left ([\boldsymbol {\Gamma }^{\prime }(\mathbf {x}-\boldsymbol {\mu })]_{j}\mid 0,{\Phi }_{j}w_{j}^{-1} \right ) \) is the density of a univariate Gaussian distribution with mean 0 and variance \({\Phi }_{j}w_{j}^{-1}\), [Γ′ (x − μ)]j is the j th element of Γ′(x −μ), and Φj is the j th eigenvalue of Φ, i.e., the j th diagonal element of the matrix Φ. It follows that Eq. 6 can be written
Solving the integral in Eq. 8 gives the density of a multiple scaled multivariate-t distribution,
where Φj is the j th eigenvalue of Φ, Γ is a matrix of eigenvectors, μ is a location parameter, and \([\boldsymbol {\Gamma }^{\prime }(\mathbf {x}-\boldsymbol {\mu })]_{j}^{2}/{\Phi }_{j}\) can be regarded as the squared Mahalanobis distance between x and μ.
The main difference between the traditional multivariate-t density given in Eq. 4 and the multiple scaled multivariate-t density given in Eq. 9 is that the degrees of freedom can now be parameterized separately in each dimension j. Therefore, unlike the standard multivariate-t distribution, the multiple scaled density in Eq. 9 can account for different tail weight in each dimension (Forbes and Wraith 2014).
3 Mixture of Multiple Scaled Generalized Hyperbolic Distributions
There are different ways to formulate the density of a generalized hyperbolic distribution (GHD; see McNeil et al. 2005, for example). Browne and McNicholas (2015) use a mixture of GHDs (MGHDs) for clustering and they use the following formulation for the density of a p-dimensional random vector X from a GHD:
where \(\boldsymbol {\mu }\in \mathbb {R}^{p}\) is the location parameter, \(\boldsymbol {\alpha }\in \mathbb {R}^{p}\) is the skewness parameter, \(\boldsymbol {\Sigma }\in \mathbb {R}^{p\times p}\) is the scale matrix, \(\lambda \in \mathbb {R}\) is the index parameter, \(\omega \in \mathbb {R}^{+}\) is the concentration parameter, and Kλ is the modified Bessel function of the third kind with index λ. Now, X∣w ∽N(μ + wα, wΣ) and W∣x ∼GIG(ω + α′Σ− 1α, ω + δ (x, μ|Σ), λ − p/2),where W ∼GIG(a, b, λ) denotes that W follows a generalized inverse Gaussian (GIG) distribution with density formulated as
for w > 0, where \(a,b\in \mathbb {R}^{+}\), and \(\lambda \in \mathbb {R}\). The GIG distribution has some attractive properties including the tractability of the following expected values:
Write X ∼GHD(μ, Σ, α, ω, λ) to denote that the p-dimensional random variable X has the density in Eq. 10.
Now, note that the formulation of the GHD in Eq. 10 can be written as a normal variance-mean mixture where the univariate density is GIG, i.e.,
where V ∼N(0, Σ) and W has density
for w > 0, where ω and λ are as previously defined. Note that Eq. 15 is just an alternative parameterization of the GIG distribution. From Eqs. 14 and 15, it follows that the generalized hyperbolic density can be written
We can use Eqs. 5 and 16 and an alternative parameterization to write the density of a multiple scaled generalized hyperbolic distribution (MSGHD) as
where ω = (ω1, … , ωp)′, λ = (λ1, … , λp)′, 1 is a p-vector of 1s, and
From Eqs. 6 and 7, it follows that Eq. 17 can be written
where [Γ′x]j is the j th element of the vector Γ′x, μj is the j th element of the location parameter μ, αj is the j th element of the skewness parameter α, Γ is a p × p matrix of eigenvectors, Φj is the j th eigenvalue of the diagonal matrix Φ, ω = (ω1, … , ωp)′ controls the concentration in each dimension p, and λ = (λ1, … , λp)′ is a p-dimensional index parameter. Write X ∽MSGHD(μ, Γ, Φ, α, ω, λ) to indicate that the random vector X follows an MSGHD with density fMSGHD(x∣μ, Γ, Φ, α, ω, λ). Then, a mixture of MSGHDs (MMSGHDs) has density
The identifiability of the MMSGHD is discussed in Appendix 3.
4 Mixture of Coalesced Generalized Hyperbolic Distributions
Note that the generalized hyperbolic distribution is not a special or limiting case of the MSGHD under any parameterization with p > 1. Motivated by this, consider a coalesced generalized hyperbolic distribution (CGHD) that contains both the generalized hyperbolic distribution and MSGHD as limiting cases. The CGHD arises through the introduction of a random vector
where X = ΓY, Y ∽GHD (μ, Σ, α, ω0, λ0), Σ = ΓΦΓ′, S ∽MSGHD(μ, Γ, Φ, α, ω, λ), and U is an indicator variable such that
It follows that \(\mathbf {X} = \boldsymbol {\Gamma }\boldsymbol {\mu } + W\boldsymbol {\Gamma }\boldsymbol {\alpha } + \sqrt {W}\boldsymbol {\Gamma }\mathbf {V}, \)where ΓV ∽Np (0, ΓΦΓ′), S = Γμ + ΓαΔw + ΓA,where ΓA ∽Np (0, ΓΔwΦΓ′), and the density of R can be written
where fGHD(⋅) is the density of a generalized hyperbolic random variable, fMSGHD(⋅) is the density of a MSGHD random variable, and ϖ ∈ (0, 1) is a mixing proportion. Note that the random vector R would be distributed generalized hyperbolic if ϖ = 1 and would be distributed MSGHD if ϖ = 0. However, we must restrict ϖ ∈ (0, 1) for identifiability reasons. Specifically, if ϖ = 0 then the value of fCGHD(r∣μ, Γ, Φ, α, ω, λ, ω0, λ0, ϖ) will be the same for any ω0 and λ0 whereas, if ϖ = 1, then the value of fCGHD(r∣μ, Γ, Φ, α, ω, λ, ω0, λ0, ϖ) will be the same for any ω and λ. The parameters μ, α, Γ, and Φ are the same for both densities, the parameters ω0 and λ0 are univariate values peculiar to the generalized hyperbolic distribution, and the p-dimensional parameters ω and λ are peculiar to the MSGHD. Write R ∽CGHD(μ, Γ, Φ, α, ω, λ, ω0, λ0, ϖ) to indicate that the random vector R follows a CGHD with density in Eq. 20.
A mixture of CGHDs (MCGHDs) has density
Parameter estimation can be carried out via a generalized expectation-maximization (GEM) algorithm (Dempster et al. 1977). There are four sources of missing data: the latent w0ig, the multi-dimensional weights Δwig = diag(w1ig, … , wpig), the component membership labels zig, and the inner component labels uig, for i = 1, … , n and g = 1, … , G. As usual, zig = 1 if observation i belongs to component g and zig = 0 otherwise. Similarly, uig = 1 if observation i, in component g, is distributed generalized hyperbolic and uig = 0 if observation i, in component g, is distributed MSGHD. It follows that the complete-data log-likelihood for the MCGHD is
Further details on parameter estimation (Appendix 1) and the identifiability of the MCGHD (Appendix 3) are discussed in appendices.
5 Cluster Convexity
The definition of clusters has been discussed quite extensively in the literature. Recently, Hennig (2015) provides a very interesting discussion of some potential characteristics that clusters may have. Even though they cannot always be observed—and, in some situations, desired characteristics may even conflict—such characteristics provide important links and contrasts between different definitions that are used for a cluster. In fact, the idea of desirable characteristics is not a new one, e.g., Cormack (1971) gives internal cohesion and external isolation as two “basic ideas” in this direction. In this paper, we follow the definition given by McNicholas (2016a): “a cluster is a unimodal component within an appropriate finite mixture model.” The term appropriate means that the model has the flexibility to fit the data and, in many cases, this also means that each cluster is convex (see McNicholas 2016a, Section 9.1). The development of flexible models outlined herein may make it a little easier to find an “appropriate” component density.
The MSGHD is more flexible than the GHD; however, similar to the multiple scaled multivariate t-distribution of Forbes and Wraith (2014), the MSGHD can have contours that are not convex. Accordingly, the MMSGHD can have components that are non-convex, leading to non-convex clusters. Consider the data in Fig. 1. How many clusters are there? The most plausible answer to this question is two overlapping clusters: one with positive correlation between the variables and another with negative correlation between the variables. There may also be an argument for four or five. Note that the data are generated from a G = 2 component mixture of multivariate t-distributions.
The MMSGHD is fitted to these data for G = 1, … , 5 and, as is common in model-based clustering applications, the Bayesian information criterion (BIC; Schwarz 1978) is used to select G. Note that the BIC is given by
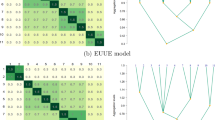
where \(l(\hat {\boldsymbol {\vartheta }})\) is the maximized log-likelihood, ρ is the number of free parameters, and n is the number of observations. For the MMSGHD, the BIC selects a G = 1 component model (Fig. 1). Furthermore, looking at the G = 2 component MSGHD solution (Fig. 1) confirms that the problem is not just one of model selection; the G = 2 component MSGHD solution selects one component that is roughly elliptical and another that is not convex. On the other hand, forcing the MSGHD to be convex—which can be done by imposing the constraint λj > 1, for j = 1, … , p—leads to what we call the convex MSGHD (cMSGHD). Fitting the corresponding mixture of cMSGHDs (McMSGHDs) ensures that, if each component is associated with a cluster, then convex clusters are guaranteed. Results in a G = 2 component model being selected (Fig. 1). Formally, this amounts to insuring that the MSGHD is quasi-convex (see Appendix 2). The general point here is that if convexity is not enforced, then the MSGHD can give components that contain multiple clusters. While it is easy to spot this in two dimensions, e.g., Fig. 1, this phenomenon may go unrecognized in higher dimensions, possibly resulting in greatly misleading results. Of course, the issue of non-convex clusters does not arise with most model-based approaches; however, when multiple scaled mixtures are considered, the issue can crop up. Another example in a similar vein is given in Fig. 2, where data are generated from a G = 3 component mixture of multivariate t-distributions. The selected MMSGHD has G = 2 components, including one clearly non-convex cluster, while the McMSGHD gives sensible clustering results.
The intention behind the introduction of the McMSGHD is not that it should supplant the MMSGHD, but rather that it provides a convenient check on the number of components. In a higher dimensional application, where visualization is difficult or impossible, situations where the selected MMSGHD has fewer components than the selected McMSGHD will deserve special attention. Of course, this is not to say that the selected McMSGHD will always have more components in situations where the MSGHD has too few, but rather that it will help to avoid the sort of situations depicted in Figs. 1 and 2. Note that parameter estimation for the McMSGHD is analogous to the algorithm for the MMSGHD algorithm but λkj (k = 0, 1, … , G, j = 1, … , p) is updated only if the value of its update exceeds 1.
6 Illustrations
6.1 Implementation and Evaluation
In the following illustrations, we fit the MGHDs, the MMSGHDs, the McMSGHDs, and the MCGHDs using the corresponding functions available in the MixGHD package (Tortora et al. 2017) for R (R Core Team 2017). We use k-means and k-medoids clustering to initialize the \(\hat {z}_{ig}\). The adjusted Rand index (ARI; Hubert and Arabie 1985) is used to compare predicted classifications with true classes. The ARI corrects the Rand index (Rand 1971) for chance, its expected value under random classification is 0, and it takes a value of 1 when there is perfect class agreement. Steinley (2004) gives guidelines for interpreting ARI values.
Several mixtures of skewed distributions have been proposed for model-based clustering and classification. Among them, three different formulations of the multivariate skew-t distribution have been used for clustering in the mixture setting. The formulation used by Murray et al. (2014) is a special case of the generalized hyperbolic distributions. Azzalini et al. (2016) refers to the other two formulations as the classical and SDB formulations (for the initials of the authors’ names), respectively. Lee and McLachlan (2013b, 2014) compare the mixture of classical skew-t distributions to the mixture of SDB skew-t distributions (MSDBST) and to skew-normal analogues of both formulations; their results lead one to believe that the MSDBST is generally preferable (see Azzalini et al. 2016, for an alternative viewpoint). The MSDBST approach is implemented using the EMMIXuskew package (Lee and McLachlan 2013a) for R and it is used for comparison herein.
6.2 Simulation Study
We use a simulation study to measure the performance of the proposed methods. The interest is to observe the ARI under different circumstances, the data are generated using two-component mixtures of Gaussian (Scenario 1), generalized hyperbolic (Scenario 2), and multiple scaled generalized hyperbolic distributions (Scenario 3), using the R function mvrnorm and the stochastic relationships given in Eq. 14 and Section 4. For each scenario, a three-factor full factorial design was used, where the factors are medium (M) or high (H) correlation, 25% or 50% overlapping, and same (S), n1 = n2 = 100, or different (D), n1 = 150 and n2 = 50, number of elements per component. In all the scenarios, μ1 = 1. Table 1 shows the average ARI, with standard deviation, obtained on 10 data sets generated from a Gaussian distribution with p = 10, G = 2, correlation equal to 0.33 for M and 0.66 for H, μ2 = 2 for 25% overlapping and μ2 = 3 for 50% overlapping. Table 2 shows the average ARI, with standard deviation, obtained on 10 data sets generated from a MGHD distribution with p = 10, G = 2, α = 1, λ = 0.5, ω = 1 and correlation equal to 0.25 for M and 0.5 for H, μ2 = 10 for 25% overlapping and μ2 = 15 for 50% overlapping. Table 3 shows the average ARI, with 1 standard deviation, obtained on 10 data sets generated from a MMSGHD distribution with p = 10, G = 2, α1 = 1, α2 = -1, λ = 0.5, ω = 1 and correlation equal to 0.25 for M and 0.35 for H, μ2 = 19 for 25% overlapping and μ2 = 23 for 50% overlapping. For the data sets generated using the MGHD and the MMSGHD, the values of the correlation had to be reduced in order to maintain 25% and 50% overlap, higher correlation lead to higher overlap. Figure 3 shows the scatterplots of the data simulated using the two-component MGDs, MGHDs, and MMSGHDs with high correlation, 50% overlapping and cluster of different size, where plotting symbol and color represent the true classifications.
The MCGHDs perform at least comparably to the best of the MMSGHDs and MGHDs. As expected, the MGHDs over performs the MMSGHDs when the data are generated using a MGHDs model and vice versa when the data are generated from a MMSGHDs. The two methods perform similarly on normally distributed clusters. The generated clusters are all convex and that explains the similarity of the results obtained using MMSGHDs and McMSGHDs. The level of overlapping has a notable impact on the ARI, as expected. The performance of all the methods is slightly better when the clusters are of the same size and when there is a lower correlation.
6.3 Real Data Analysis
To assess the classification performance of the MGHDs, the MMSGHDs, the McMSGHDs, the MCGHDs, and the MSDBST, we consider four real data sets that are commonly used within the model-based clustering literature (Table 4).
To compare the performance of the methods, we set the number of components, G, equal to the true number of classes. In general, the BIC is used to select the best starting criterion between k-means and k-medoids; however, because of the presence of outliers, only k-medoids starts are used for the bankruptcy data set. Table 5 displays the classification performance. Note that MSDBST is not used on the AIS data set because of the prohibitively high dimensionality (p = 11). The MCGHD generally performs comparably to the best of the other four approaches. Specifically, the MCGHD gives the best classification performance—either outright or jointly—for the four data sets. Interestingly, the MGHD gives very good classification performance on three of the four data sets; however, this must be taken in context with its very poor classification performance on the bankruptcy data set (ARI ≈ 0). It is also interesting to compare the classification performance of the McMSGHD to the MMSGHD as well as that of the MGHD to the MMSGHD. The McMSGHD and the MGHD approaches both outperform MMSGHD for one data set, and give a similar performance on two of the other three data sets. This highlights the fact that a mixture of multiple scaled distributions may well not outperform its single scaled analogue, and underlines the need for an approach with both the MSGHD and MGHD models as special cases. The results for the bankruptcy data illustrate that the MCGHD approach can give very good classification performance in situations where neither the MGHD nor the MMSGHD perform well. Finally, the MCGHD outperforms the MSDBST on two data sets and gives the same result on the third (recall that the MSDBST could not be fitted to the AIS data).
In this analysis, we took the number of components to be known. However, in a true clustering scenario, we would not have a priori information about the number of groups. Therefore, the analysis was repeated without this assumption and, for all but the bankruptcy data set, all approaches were run for G = 1, … , 5 components. Because of the small number of observations, the bankruptcy data were run for G = 1, 2, 3. Table 6 gives the number of components selected by the BIC.
For the bank note data, the BIC selects the correct number of components for every method; however, for the bankruptcy data, it picks the right number of components only for McMSGHDs. For the seed and AIS data sets, the BIC does not select the correct number of components for any approach. Recall that the MSDBST could not be fitted to the AIS data; also, it could not be fitted to the seed data for G = 4. These results illustrate that the BIC is not necessarily reliable for selecting the number of components in real data examples for any of the five approaches.
6.4 Computation Time
For the data sets listed in Table 4, we measured the elapsed user times, in seconds, to perform 100 (G)EM iterations. Note that all code was run in R version 3.0.2 on a 32-core Intel Xeon E5 server with 256GB RAM running 64-bit CentOS. Figure 4 displays the average elapsed time for two replications of each algorithm using a k-means and a k-medoids starting partition for G = 1, … , 5 components. The EM algorithm for the MSDBST is significantly slower than that of the hyperbolic-based approaches (Fig. 4). In fact, on the banknote data set, it takes the MSDBST more than 11 h to perform the required number of EM iterations when G = 2 and about 63 h when G = 5. For the seeds data set, it takes the MSDBST more than 5 h when G = 2 and more than 10 h when G = 5, whereas the G = 5 component MMSGHD, McMSGHD, and MCGHD approaches need less than 25 s. For the bankruptcy data set, the MSDBST requires approximately 80 s when G = 3, whereas the hyperbolic distributions need less than 20 s each.
7 Discussion
Novel MCGHDs, MMSGHDs, and McMSGHDs models have been introduced and applied for model-based clustering. The GHD is a flexible distribution, capable of handling skewness and heavy tails, and has many well known distributions as special or limiting cases. Furthermore, it is a normal variance-mean mixture, arising via a relationship between a multivariate Gaussian and an univariate GIG distribution. The MSGHD extends the GHD to include a multivariate GIG distribution, increasing the flexibility of the model. However, the GHD is not a special case of the MSGHD; hence, we created MCGHDs, which has both the GHD and MSGHD as special cases. The McMSGHD approach was introduced as a convex version of the MMSGHD, and this point deserves some further discussion. The extension of the multivariate-t distribution to multiple scale was carried out by Forbes and Wraith (2014); as discussed in the Appendix 2, the multiple scaled multivariate t-distribution cannot be quasi-concave, i.e., the clusters associated with a mixture of multiple scaled multivariate t-distributions cannot be convex. We have seen examples where the MMSGHD can put multiple clusters into one component, and the McMSGHD has an important role in helping to prevent this; if both approaches are fitted and lead to different numbers of components, then further attention is warranted.
Comparing the MGHD, MMSGHD, McMSGHD, and MCGHD approaches yielded some interesting results. Among them, we see that the MMSGHD does not necessarily outperform the MGHD; far from it, in fact, only the MGHD approach gave better clustering performance than the MMSGHD approach on one of the four real data sets we considered, as well as identical performance on the other two. This underlines the fact that a mixture of multiple scaled distributions may well not outperform its single scaled analogue, and highlights the benefit of approaches with both a multiple scaled distribution and its single scaled analogue as special cases. The MCGHDs represent one such approach, with the MSGHD and MGHD models as special cases. The approaches introduced herein, as well as the MGHDs, have been made freely available via the MixGHD package for R.
Future work will focus in several directions. For one, it will be interesting to study the performance of the approaches introduced herein within the fractionally supervised classification framework (Vrbik and McNicholas 2015; Gallaugher and McNicholas 2019a). The extension of these approaches to the matrix-variate paradigm is also of interest and may proceed in an analogous fashion to the work of Gallaugher and McNicholas (2018, 2019b). Finally, the models introduced herein can be extended to account for missing data (see Wei et al.2019).
References
Aitken, A.C. (1926). A series formula for the roots of algebraic and transcendental equations. Proceedings of the Royal Society of Edinburgh, 45, 14–22.
Altman, E. (1968). Financial ratios, discriminant analysis and the prediction of corporate bankruptcy. Journal of Finance, 23(4), 589–609.
Andrews, J.L., & McNicholas, P.D. (2011a). Extending mixtures of multivariate t-factor analyzers. Statistics and Computing, 21(3), 361–373.
Andrews, J.L., & McNicholas, P.D. (2011b). Mixtures of modified t-factor analyzers for model-based clustering, classification, and discriminant analysis. Journal of Statistical Planning and Inference, 141(4), 1479–1486.
Andrews, J.L., & McNicholas, P. (2012). Model-based clustering, classification, and discriminant analysis via mixtures of multivariate t-distributions. Statistics and Computing, 22(5), 1021–1029.
Azzalini, A., Browne, R.P., Genton, M.G., McNicholas, P.D. (2016). On nomenclature for, and the relative merits of, two formulations of skew distributions. Statistics and Probability Letters, 110, 201–206.
Baek, J., & McLachlan, G.J. (2011). Mixtures of common t-factor analyzers for clustering high-dimensional microarray data. Bioinformatics, 27, 1269–1276.
Banfield, J.D., & Raftery, A.E. (1993). Model-based Gaussian and non-Gaussian clustering. Biometrics, 49(3), 803–821.
Barndorff-Nielsen, O. (1978). Hyperbolic distributions and distributions on hyperbolae. Scandinavian Journal of Statistics, 5(3), 151–157.
Barndorff-Nielsen, O., Kent, J., Sørensen, M. (1982). Normal variance-mean mixtures and z distributions. International Statistical Review / Revue Internationale de Statistique, 50(2), 145–159.
Böhning, D., Dietz, E., Schaub, R., Schlattmann, P., Lindsay, B. (1994). The distribution of the likelihood ratio for mixtures of densities from the one-parameter exponential family. Annals of the Institute of Statistical Mathematics, 46, 373–388.
Browne, R.P., & McNicholas, P.D. (2014). Estimating common principal components in high dimensions. Advances in Data Analysis and Classification, 8(2), 217–226.
Browne, R.P., & McNicholas, P.D. (2015). A mixture of generalized hyperbolic distributions. Canadian Journal of Statistics, 43(2), 176–198.
Celeux, G., & Govaert, G. (1995). Gaussian parsimonious clustering models. Pattern Recognition, 28(5), 781–793.
Charytanowicz, M., Niewczas, J., Kulczycki, P., Kowalski, P.A., Łukasik, S., żak, S. (2010). A complete gradient clustering algorithm for features analysis of x-ray images. In Piȩtka, E., & Kawa, J. (Eds.) Information Technologies in Biomedicine, (Vol. 2 pp. 15–24). Berlin: Springer.
Cook, R.D., & Weisberg, S. (1994). An Introduction to Regression Graphics. New York: Wiley.
Cormack, R.M. (1971). A review of classification (with discussion). Journal of the Royal Statistical Society: Series A, 34, 321–367.
Debreu, G., & Koopmans, T.C. (1982). Additively decomposed quasiconvex functions. Mathematical Programming, 24(1), 1–38.
Demarta, S., & McNeil, A.J. (2005). The t copula and related copulas. International Statistical Review, 73(1), 111–129.
Dempster, A.P., Laird, N.M., Rubin, D.B. (1977). Maximum likelihood from incomplete data via the EM algorithm. Journal of the Royal Statistical Society: Series B, 39(1), 1–38.
Flury, B., & Riedwyl, H. (1988). Multivariate Statistics: A Practical Approach. London: Chapman & Hall.
Forbes, F., & Wraith, D. (2014). A new family of multivariate heavy-tailed distributions with variable marginal amounts of tailweights: Application to robust clustering. Statistics and Computing, 24(6), 971–984.
Fraley, C., & Raftery, A.E. (2002). Model-based clustering, discriminant analysis, and density estimation. Journal of the American Statistical Association, 97(458), 611–631.
Franczak, B.C., Browne, R.P., McNicholas, P.D. (2014). Mixtures of shifted asymmetric Laplace distributions. IEEE Transactions on Pattern Analysis and Machine Intelligence, 36(6), 1149–1157.
Franczak, B.C., Tortora, C., Browne, R.P., McNicholas, P.D. (2015). Unsupervised learning via mixtures of skewed distributions with hypercube contours. Pattern Recognition Letters, 58(1), 69–76.
Gallaugher, M.P.B., & McNicholas, P.D. (2018). Finite mixtures of skewed matrix variate distributions. Pattern Recognition, 80, 83–93.
Gallaugher, M.P.B., & McNicholas, P.D. (2019a). On fractionally-supervised classification: weight selection and extension to the multivariate t-distribution. Journal of Classification 36. In press.
Gallaugher, M.P.B., & McNicholas, P.D. (2019b). Three skewed matrix variate distributions. Statistics and Probability Letters, 145, 103–109.
Ghahramani, Z., & Hinton, G.E. (1997). The EM algorithm for factor analyzers Technical Report CRG-TR-96-1. Toronto: University Of Toronto.
Gneiting, T. (1997). Normal scale mixtures and dual probability densities. Journal of Statistical Computation and Simulation, 59(4), 375–384.
Hennig, C. (2015). What are the true clusters? Pattern Recognition Letters, 63, 53–62.
Holzmann, H., Munk, A., Gneiting, T. (2006). Identifiability of finite mixtures of elliptical distributions. Scandinavian Journal of Statistics, 33, 753–763.
Hubert, L., & Arabie, P. (1985). Comparing partitions. Journal of Classification, 2(1), 193–218.
Hunter, D.R., & Lange, K. (2000). Quantile regression via an MM algorithm. Journal of Computational and Graphical Statistics, 9(1), 60–77.
Karlis, D., & Santourian, A. (2009). Model-based clustering with non-elliptically contoured distributions. Statistics and Computing, 19(1), 73–83.
Kent, J.T. (1983). Identifiability of finite mixtures for directional data. The Annals of Statistics, 11, 984–988.
Kiers, H.A. (2002). Setting up alternating least squares and iterative majorization algorithms for solving various matrix optimization problems. Computational Statistics and Data Analysis, 41(1), 157–170.
Kotz, S., Kozubowski, T.J., Podgorski, K. (2001). The Laplace Distribution and Generalizations: A Revisit with Applications to Communications, Economics, Engineering, and Finance 1st edn: Burkhauser Boston.
Kotz, S., & Nadarajah, S. (2004). Multivariate t-distributions and their applications. Cambridge: Cambridge University Press.
Lee, S.X., & McLachlan, G.J. (2013a). EMMIXuskew: fitting unrestricted multivariate skew t Mixture Models. R package version 0.11–5.
Lee, S.X., & McLachlan, G.J. (2013b). On mixtures of skew normal and skew t-distributions. Advances in Data Analysis and Classification, 7(3), 241–266.
Lee, S.X., & McLachlan, G.J. (2014). Finite mixtures of multivariate skew t-distributions: some recent and new results. Statistics and Computing, 24(2), 181–202.
Lin, T.I. (2009). Maximum likelihood estimation for multivariate skew normal mixture models. Journal of Multivariate Analysis, 100(2), 257–265.
Lin, T.I. (2010). Robust mixture modeling using multivariate skew t distributions. Statistics and Computing, 20(3), 343–356.
Lin, T.-I., McNicholas, P.D., Hsiu, J.H. (2014). Capturing patterns via parsimonious t mixture models. Statistics and Probability Letters, 88, 80–87.
Lindsay, B. (1995). Mixture models: Theory, geometry and applications. In NSF-CBMS Regional Conference Series in Probability and Statistics, Vol. 5. California: Institute of Mathematical Statistics: Hayward.
McLachlan, G.J., & Peel, D. (2000). Mixtures of factor analyzers. In Proceedings of the Seventh International Conference on Machine Learning (pp. 599–606). San Francisco: Morgan Kaufmann.
McLachlan, G.J., Bean, R.W., Jones, L. B. -T. (2007). Extension of the mixture of factor analyzers model to incorporate the multivariate t-distribution. Computational Statistics and Data Analysis, 51(11), 5327–5338.
McLachlan, G.J., & Krishnan, T. (2008). The EM Algorithm and Extensions. New York: Wiley.
McNeil, A.J., Frey, R., Embrechts, P. (2005). Quantitative risk management: concepts, techniques and tools. Princeton: Princeton University Press.
McNicholas, P.D. (2016a). Mixture Model-Based Classification. Boca-Raton: Chapman & Hall/CRC press.
McNicholas, P.D. (2016b). Model-based clustering. Journal of Classification, 33 (3), 331–373.
McNicholas, P.D., Murphy, T.B., McDaid, A.F., Frost, D. (2010). Serial and parallel implementations of model-based clustering via parsimonious Gaussian mixture models. Computational Statistics and Data Analysis, 54(3), 711–723.
McNicholas, S.M., McNicholas, P.D., Browne, R.P. (2017). A mixture of variance-gamma factor analyzers. In Ahmed, S. E. (Ed.) Big and Complex Data Analysis: Methodologies and Applications (pp. 369–385). Cham: Springer International Publishing.
Murray, P.M., Browne, R.B., McNicholas, P.D. (2014). Mixtures of skew-t factor analyzers. Computational Statistics and Data Analysis, 77, 326–335.
Murray, P.M., Browne, R.B., McNicholas, P.D. (2017). Hidden truncation hyperbolic distributions, finite mixtures thereof, and their application for clustering. Journal of Multivariate Analysis, 161, 141–156.
Niculescu, C., & Persson, L. (2006). Convex Functions and Their Applications. New York: Springer.
Ortega, J.M., & Rheinboldt, W.C. (1970). Iterative Solutions of Nonlinear Equations in Several Variables. New York: Academic Press.
Peel, D., & McLachlan, G.J. (2000). Robust mixture modelling using the t distribution. Statistics and Computing, 10(4), 339–348.
Pesevski, A., Franczak, B.C., McNicholas, P.D. (2018). Subspace clustering with the multivariate-t distribution. Pattern Recognition Letters, 112(1), 297–302.
R Core Team. (2017). R: A Language and Environment for Statistical Computing Vienna. Austria: R Foundation for Statistical Computing.
Rand, W.M. (1971). Objective criteria for the evaluation of clustering methods. Journal of the American Statistical Association, 66(336), 846–850.
Rockafellar, R.T., & Wets, R.J.B. (2009). Variational Analysis. New York: Springer.
Schwarz, G. (1978). Estimating the dimension of a model. Annals of Statistics, 6(2), 461–464.
Steane, M.A., McNicholas, P.D., Yada, R. (2012). Model-based classification via mixtures of multivariate t-factor analyzers. Communications in Statistics – Simulation and Computation, 41(4), 510–523.
Steinley, D. (2004). Properties of the Hubert-Arable adjusted Rand index. Psychological methods, 9(3), 386.
Tang, Y., Browne, R.P., McNicholas, P.D. (2018). Flexible clustering of high-dimensional data via mixtures of joint generalized hyperbolic distributions. Stat, 7 (1), e177.
Tipping, M.E., & Bishop, C.M. (1999). Mixtures of probabilistic principal component analysers. Neural Computation, 11(2), 443–482.
Tortora, C., Franczak, B.C., Browne, R.P., McNicholas, P.D. (2014). Mixtures of multiple scaled generalized hyperbolic distributions. arXiv:1403.2332v1.
Tortora, C., Browne, R.P., Franczak, B.C., McNicholas, P.D. (2017). MixGHD: model based clustering, classification and discriminant analysis using the mixture of generalized hyperbolic distributions. R package version 2.1.
Vrbik, I., & McNicholas, P.D. (2012). Analytic calculations for the EM algorithm for multivariate skew-mixture models. Statistics and Probability Letters, 82(6), 1169–1174.
Vrbik, I., & McNicholas, P.D. (2014). Parsimonious skew mixture models for model-based clustering and classification. Computational Statistics and Data Analysis, 71, 196–210.
Vrbik, I., & McNicholas, P.D. (2015). Fractionally-supervised classification. Journal of Classification, 32(3), 359–381.
Wei, Y., Tang, Y., McNicholas, P.D. (2019). Mixtures of generalized hyperbolic distributions and mixtures of skew-t distributions for model-based clustering with incomplete data. Computational Statistics and Data Analysis, 130, 18–41.
Wraith, D., & Forbes, F. (2015). Clustering using skewed multivariate heavy tailed distributions with flexible tail behaviour. arXiv:http://arXiv.org/abs/1408.0711.
Yakowitz, S.J., & Spragins, J. (1968). On the identifiability of finite mixtures. Annals of Mathematical Statistics, 39, 209–214.
Acknowledgements
The authors are grateful to two anonymous reviewers and the Editor for helpful comments that have improved this manuscript. This work was supported by a grant-in-aid from Compusense Inc., a Collaborative Research and Development Grant from the Natural Sciences and Engineering Research Council of Canada, and the Canada Research Chairs program. The work on the introduction of the MMSGHD presented herein was first made publicly available as an arXiv preprint (Tortora et al. 2014).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendices
Appendix 1: Parameter Estimation
We use the EM algorithm to estimate the parameters of the MCGHDs. The EM algorithm belongs to a larger class of algorithms known as MM algorithms (Ortega and Rheinboldt 1970; Hunter and Lange 2000) and is well-suited for problems involving missing data. “MM” stands for “minorize-maximize” or “majorize-minimize,” depending on the purpose of the algorithm; in the EM context, the minorizing function is the expected value of the complete-data log-likelihood. The EM algorithm iterates between two steps, an E-step and a M-step, and has been used to estimate the parameters of mixture models in many experiments (McLachlan and Krishnan 2008). On each E-step, the expected value of the complete-data log-likelihood, \(\mathcal {Q}\), is calculated and on each M-step is maximized with respect to πg, μg, Φg, αg, ωg, λg, ω0g, λ0g, ϖg. However, in each M-step \(\mathcal {Q}\) increases with respect to Γg rather than maximize; accordingly, the algorithm is formally a generalized EM (GEM) algorithm. For our MCGHDs, there are four sources of missing data: the latent variable W0ig, the multi-dimensional weight variable Δwig, the group component indicator labels zig, and inner component labels uig, for i = 1, … , n and g = 1, … , G. For each observation i, zig = 1 if observation i is in component g and zig = 0 otherwise. Similarly, for each observation i, uig = 1 if observation i, in component g, is distributed generalized hyperbolic and uig = 0 if observation i, in component g, is distributed multiple scaled generalized hyperbolic. It follows that the complete-data log-likelihood for the MCGHDs is given by
where ϕp(⋅) represents a p-dimensional Gaussian density function, ϕ1(⋅) is a unidimensional Gaussian density function, and h(⋅) is the density of a GIG distribution given in Eq. 15.
We are now prepared to outline the calculations for our GEM algorithm for the MCGHDs. On the E-step, the expected value of the complete-data log-likelihood,\( \mathcal {Q}\), is computed by replacing the sufficient statistics of the missing data by their expected values. For each component indicator label zig and inner component label uig, for i = 1, … , n and g = 1, … , G, we require the expectations
and
where fCGHD is given in Eq. 20, fGHD is given in Eq. 10 and fMSGHD is given in Eq. 17. For the latent variable W0ig, we use the expected value given in Browne and McNicholas (2015). The authors show that, given the density in Eq. 15, the following is true
For the MCGHDs, the maximization of \(\mathcal {Q}\) requires the expected values of W0ig, \(W^{-1}_{0ig}\), and log W0ig, i.e.,
where \(d_{g}= \omega _{0g}+\boldsymbol {\alpha }_{g}^{\prime } (\boldsymbol {\Gamma }_{g}\boldsymbol {\Phi }_{g}\boldsymbol {\Gamma }^{\prime }_{g})^{-1} \boldsymbol {\alpha }_{g}\) and \(e_{ig}= \omega _{0g}+\delta (\mathbf {x}_{i}, \boldsymbol {\mu }_{g} \mid \boldsymbol {\Gamma }_{g}\boldsymbol {\Phi }_{g}\boldsymbol {\Gamma }^{\prime }_{g})\).
The maximization of \(\mathcal {Q}\) also requires the expected values of the multidimensional weight variables Δwig, \(\boldsymbol {\Delta }_{\mathbf {w} ig}^{-1}\), and log Δwig. Given the density in Eq. 17, it follows that
Each multidimensional weight variable is replaced by its expected value and so we need to compute E1ig = diag{E1i1g, … , E1ipg}, E2ig = diag{E2i1g, … , E2ipg}, and E3ig = diag{E3i1g, … , E3ipg}, where
\(\bar {d}_{jg} = \omega _{jg} + \alpha _{jg}^{2}{\Phi }_{jg}^{-1}\) and \(\bar {e}_{ijg} =\omega _{jg} + ([\mathbf {x}_{i} -\boldsymbol {\mu }_{g}]_{j} )^{2}/\phi _{jg}\). Let \(n_{g}={\sum }_{i = 1}^{n} \hat z_{ig}\), \(A_{g}=(1/n_{g}){\sum }_{i = 1}^{n} \hat z_{ig}a_{ig}\), \(B_{g}=(1/n_{g}){\sum }_{i = 1}^{n} \hat z_{ig}b_{ig}\), \(C_{g}=(1/n_{g}){\sum }_{i = 1}^{n} \hat z_{ig}c_{ig}\), \({\bar {E}}_{1jg}=(1/n_{g}){\sum }_{i = 1}^{n} \hat z_{ig}{ E}_{1ijg}\), \({\bar {E}}_{2jg}=(1/n_{g}){\sum }_{i = 1}^{n} \hat z_{ig}{ E}_{2ijg}\), and \({\bar {E}}_{3jg}=(1/n_{g}){\sum }_{i = 1}^{n} \hat z_{ig} { E}_{3ijg}\).
In the M-step, we maximize the expected value of the complete-data log-likelihood with respect to the model parameters. The mixing proportions and inner mixing proportions are updated via \(\hat {\pi }_{g}=n_{g}/n\) and \(\hat {\varpi }_{g}={{\sum }_{i = 1}^{n} \hat {u}_{ig} \hat {z}_{ig}}/{n_{g}}\), respectively. The elements of the location parameter μg and skewness parameter αg are replaced with
respectively, where \([\boldsymbol {\Gamma }_{g}^{\prime } \mathbf {x}_{i}]_{j}\) is the j th element of the matrix \(\boldsymbol {\Gamma }_{g}^{\prime } \mathbf {x}_{i}\), \(s_{1ijg}= \hat {u}_{ig}a_{ig}+\left (1- \hat {u}_{ig} \right ) { E}_{1ijg}\), \(s_{2ijg} =\hat {u}_{ig}b_{ig}+\left (1- \hat {u}_{ig} \right ) { E}_{2ijg}\), \(\bar {s}_{1jg}= 1/n_{g}{\sum }_{i = 1}^{n} \hat {z}_{ig}s_{1ijg},\bar {s}_{2jg}= 1/n_{g}{\sum }_{i = 1}^{n} \hat {z}_{ig}s_{2ijg}\). The diagonal elements of the matrix Φg are updated using
To update the component eigenvector matrices Γg, we wish to minimize the objective function
with respect to Γg, where \(\mathbf V_{ig} = \hat {u}_{ig} b_{ig}\mathbf {I}_{p} + (1-\hat {u}_{ig})\textbf {E}_{2ig}\). We employ an optimization routine that uses two simpler majorization-minimization algorithms. Our optimization routine exploits the convexity of the objective function in Eq. 24, providing a computationally stable algorithm for estimating Γg. Specifically, we follow Kiers (2002) and Browne and McNicholas (2014) and use the surrogate function
where C is a constant that does not depend on Γg, r ∈ {1, 2} is an index, and the matrices Frg are defined in Eqs. 26 and 27.
Therefore, on each M-step, we calculate either
or
where α1ig is the largest eigenvalue of the diagonal matrix \( \boldsymbol {\Phi }_{g}^{-1}\mathbf V_{ig}\), and α2ig is equal to \(\hat {z}_{ig}\mathbf {x}_{i}^{\prime } \mathbf {x}_{i}\), which is the largest eigenvalue of the rank-1 matrix \(\hat {z}_{ig}\mathbf {x}_{i} \mathbf {x}_{i}^{\prime }\). Following this, we compute the singular value decomposition of Frg given by
It follows that our update for Γg is given by
The p-dimensional concentration and index parameters, i.e., ωg and λg, are estimated by maximizing the function
This leads to
and
where the superscript “prev” denotes that the estimate from the previous iteration is used. The univariate parameters ω0g and λ0g are estimated by maximizing the function
giving
and
Our GEM algorithm is iterated until convergence, which is determined using the Aitken acceleration (Aitken 1926). Formally, the Aitken acceleration is given by
where l(k) is the value of the log-likelihood at the iteration k and
is an asymptotic estimate of the log-likelihood on iteration k + 1. The algorithm can be considered to have converged when \(l^{(k + 1)}_{\infty }-l^{(k)}< \epsilon \), provided this difference is positive (Böhning et al. 1994; Lindsay 1995; McNicholas et al. 2010). Herein, we set 𝜖 = 0.01. When the algorithm converges, we compute the maximum a posteriori (MAP) classification values using the posterior \(\hat {z}_{ig}\), where \(\text {MAP}\left \{\hat {z}_{ig}\right \}= 1\) if \(g=\arg \max _{h}\left \{\hat {z}_{ih}\right \}\), and \(\text {MAP}\left \{\hat {z}_{ig}\right \}= 0\) otherwise.
Appendix 2: Quasi-Concavity of the cMSGHD
In essence, we might want to consider only densities whose contours contain a set of points that are convex. Formally, such densities are quasi-concave. Extensive details on quasi-concavity, quasi-convexity, and related notions are given by Niculescu and Persson (2006) and Rockafellar and Wets (2009).
Definition 1
A function f(x) is quasi-concave if each upper-level set Uα(f) = {x|f(x) ≥ α} is convex, for \(\alpha \in \mathbb {R}\).
Definition 2
A function f(x) is quasi-convex if each sub-level set Sα(f) = {x|f(x) ≤ α} is convex, for \(\alpha \in \mathbb {R}\).
Lemma 1
The class of elliptical distributions, whose density functions have theform
are quasi-concave if the generator function, g, is monotonic non-increasing.
Proof
Result follows from the fact that if the function δ (x, μ|Σ) is convex since Σ is positive definite and the function g is monotonic non-increasing, then the function f(x) is quasi-concave. □
Theorem 1
The generalized hyperbolic distribution (GHD) is quasi-concave.
Proof
It is straightforward to show that the function
is convex, where a and b are positive constants, and δ (x, μ|Σ) is the Malahanobis distance between x and μ. Let τ = λ − p/2. Then, the function
where \(z\in \mathbb {R}^{+}\), and Kτ is the modified Bessel function of the third kind with index τ, is monotonic decreasing (or non-increasing) because the first derivative
is negative for all \(\tau \in \mathbb {R}\) and z > 0. In addition to being monotonic decreasing, k(z) is convex for τ < 1/2, concave and convex (linear) for τ = 1/2, and concave for τ > 1/2. Because k(z) is a monotonic function, it satisfies the criteria for quasi-convexity and quasi-concavity, so it is simultaneously quasi-convex and quasi-concave. In this context, monotone functions are also known as quasi-linear or quasi-montone.
Recall that if the function U is quasi-convex and the function g is decreasing, then the function f(x) = g(U(x)) is quasi-concave. It follows that the composition k(h(x)) is quasi-concave. Consider the skewness part of the GHD density function, i.e., a(x) = −(x −μ)′Σ− 1α, which is a linear function. It follows that the function
is also quasi-concave, and the result follows from the fact that Eq. 30 is proportional to the density of the GHD. □
Theorem 2
The convex multiple scaled generalized hyperbolic distribution(cMSGHD) is quasi-concave. In other words, the multiple scaledgeneralized hyperbolic distribution (MSGHD) is quasi-concave provided thatλj > 1 for allj = 1, … , p.
Proof
A p-dimensional multiple scaled distribution is a product of p independent univariate densities. The density of the MSGHD has form
where g1(xj|θj) is the density of the univariate hyperbolic distribution with parameters θj, j = 1, … , p. From Theorem 1, log g1(xj|θj) is a concave function for τj > 1/2, i.e., for λj > 1 (because p = 1). Therefore, the function
is concave provided that λj > 1 for all j = 1, … , p. Therefore, the function
is quasi-concave provided that λj > 1 for all j = 1, … , p. □
Note that addition does not preserve quasi-convexity or quasi-concavity. The sum of two quasi-convex functions defined on different domains will be quasi-concave if they are additively decomposed (see Debreu and Koopmans 1982). Debreu and Koopmans (1982) give necessary and sufficient conditions for the sum f of a set of functions f1, … , fm to be additively decomposed. These conditions depend on the convexity index c(f) in which f is quasi convex if and only if either of the following hold: (i) c(fi) ≥ 0 for every i, or (ii) c(fj) < 0 for some j, c(fi) > 0 for every i ≠ j, and \({\sum }_{i = 1}^{m} \frac {1}{c(f_{i})} \le 0\). For differentiable functions, the convexity index satisfies the inequality f″(x)/[f′(x)]2 ≥ c(f).
We have that a sufficient condition for the MSGHD to be quasi-concave is that all λj > 1. Furthermore, a sufficient condition for the MSGHD not to be quasi-concave is that all λj < 1 and finite. Interestingly, this means the multiple scaled t-distribution cannot provide convex level sets for any finite degrees of freedom. For large degrees of freedom, the multiple scaled t-distribution will behave similarly to a normal distribution near the mode; however, as one moves away from the mode, non-convex contours will be encountered. Finally, note that Debreu and Koopmans (1982) give necessary and sufficient conditions that suggest a quasi-concave MSGHD with some λj positive and others negative is possible, but going this route would greatly complicate the estimation procedure.
Appendix 3: Finite Mixture Identifiability
In this section, we consider the notion of identifiability for finite mixtures of MSGHDs and coalesced generalized hyperbolic distributions (CGHDs). Herein, we take the term identifiability to mean finite mixture identifiability.
3.1 3.1 Background
Holzmann et al. (2006) prove identifiability of finite mixtures of elliptical distributions. They state that “finite mixtures are said to be identifiable if distinct mixing distributions with finite support correspond to distinct mixtures.” A finite mixture of the densities fp(x|Ψ1), … , fp(x|ΨG) is identifiable if the family \(\left \{ f_{p}(\mathbf {x}|\boldsymbol {\Psi }) : \boldsymbol {\Psi } \in \mathcal {A}^{p} \right \}\) is linearly independent. The founding work on finite mixture identifiability is by Yakowitz and Spragins (1968), who state that this linear independence is a necessary and sufficient condition for identifiability.
The GHD can be expressed as a normal variance-mean mixture. The stochastic relationship of the normal variance-mean mixture is given by
where \(\mathbf {U} \backsim \mathcal {N}_{p}(\mathbf {0}, \boldsymbol {\Sigma })\) and W, independent of U, is a positive univariate random variable with density h(w|θ). Browne and McNicholas (2015) proved identifiability for finite mixtures of GHDs through additivity of disjoint sets of identifiable distributions.
Definition 3
In the present context, a finite mixture of the multiple scale distributions f(x|θ1), … , f(x|θG) is identifiable if
for \(\mathbf {x} \in \mathbb {R}^{p}\), where G is a positive integer, \({\sum }_{g = 1}^{G} \pi _{g} = {\sum }_{g = 1}^{G} \pi _{g}^{\star } = 1\) and \(\pi _{g}, \pi _{g}^{\star } > 0\) for g = 1, … , G, implies that there exists a permutation σ such that (πg, θg) = (πσ(g), θσ(g)) for all g.
Browne and McNicholas (2015) prove identifiability for normal variance-mean mixtures, which includes the generalized hyperbolic. Here, we view the results from a different vantage point to illustrate the concepts required for the identifiability of the multiple scaled distributions. We begin by noting the characteristic function for the generalized hyperbolic arises from the characteristic function of the normal variance-mean mixture,
where
The characteristic function for the generalized hyperbolic is
In the context of a coalesced distribution, with a eigen-decomposed scale matrix, the characteristic function is
Now, we let v = tz and obtain
To prove identifiability of the generalized hyperbolic, we could now use the results from Browne and McNicholas (2015) and Yakowitz and Spragins (1968, p. 211) that implies there exists z such that the tuple \((\mathbf {z}^{\prime } \boldsymbol {\Sigma }_{g} \mathbf {z}, \boldsymbol {\beta }_{g}^{\prime } \mathbf {z}, \mathbf {z}^{\prime }\boldsymbol {\mu }_{g})\), where \(\boldsymbol {\Sigma }_{g}= \boldsymbol {\Gamma }_{g} \boldsymbol {\Phi }_{g} \boldsymbol {\Gamma }_{g}^{\prime }\) is unique for all g = 1, … , G, allows a reduction to the univariate case. Now, we rewrite the term z′Σgz as
which implies the tuple
is unique for all g = 1, … , G. A similar argument indicates there exists a z such that the tuple
is unique. In fact, a more general statement indicates that there exists a z such that the tuple
is unique for monotonic \(\varphi : \mathbb {R}^{+} \mapsto \mathbb {R}^{+} \). Deriving this unique set of tuples facilitates the reduction to the univariate case. This is useful because the univariate generalized hyperbolic density is identifiable (see Browne and McNicholas2015).
3.2 3.2 Identifiability of a Finite Mixture of Multiple Scaled Distributions
For a multiple scaled distribution, we only need to find a single direction where the distribution is finite mixture identifiable because, as noted in Remark 2 of Kent (1983), a distribution might be non-identifiable on a subset of \(\mathbb {R}^{p}\) but identifiability can endure over \(\mathbb {R}^{p}\). In other words, for a distribution to be non-identifiable, a linear combination has to be equal to zero for all \(x \in \mathbb {R}^{p}\). This is illustrated by the example given in Kent (1983):
“the polynomials P(x1, x2) = 1 and \(P(x_{1}, x_{2}) = ({x_{1}^{2}}+ x_{2})^{3}\), \(x\in \mathbb {R}^{2}\), are equal on the unit circle, but are not the same on all of \(\mathbb {R}^{2}\).”
As a consequence, if a multivariate distribution is identifiable in some direction then it is identifiable over \(\mathbb {R}^{p}\).
To begin, consider that if there is at one least direction or column of Γg that is equal across g = 1, … , G, then the identifiability of a multiple scaled distribution follows from the identifiability of the univariate distribution. Whereas if one column of Γg is unequal, that implies, by the nature of orthonormal matrices, that two columns of Γg are unequal. We will now illustrate how the bivariate multiple scaled distribution is identifiable, which implies identifiability for finite p.
When Γg differ, the identifiability of the multiple scaled distribution depends on the behavior of the multiple scaled distribution’s density and moment generating functions when we consider moving along directions other than the columns of Γg. For example, a bivariate multiple scaled t-distribution behaves (by definition) like a t-distribution with ν1 and ν2 degrees of freedom along each of it’s principal axes, but along any other direction, a bivariate multiple scaled t-distribution behaves asymptotically like a t-distribution with ν1 + ν2 degrees of freedom.
Consider the following three orthonormal matrices in the context of an eigen-decomposition of a matrix;
If we have equal eigenvalues then we cannot distinguish between Γ1 and Γ2. In the same way, if we have the same distribution along the first and second axis, we cannot distinguish between them. However, if we have eigenvalue ordering we can distinguish between Γ1 and Γ2, but eigenvalue ordering will not allow us to distinguish between Γ1 and Γ3, since they yield the same basis or set of directions. Therefore, in general, Γ is unique up to multiplication by
One way to establish uniqueness is to require the largest value of each column of Γ to be positive. An equivalent requirement is for Γ1 ≠ Γ2 which requires that
for j = 1, … , p, \(\mathbf {z} \in \mathbb {R}^{P}\), z ≠ 0p and R is a set of diagonal matrices such that diag(R) = (± 1, … , ± 1) excluding the identity matrix. Note that [a]j denotes the j th element of the vector a. However, if we had two orthonormal matrices such that \(\boldsymbol {\Gamma }_{1}^{\prime } \boldsymbol {\Gamma }_{2} = \mathbf {I}\), then Γ1 = Γ2. If \(\boldsymbol {\Gamma }_{1}^{\prime } \boldsymbol {\Gamma }_{2} = \mathbf {R}\), then our orthonormal condition amounts to \( \boldsymbol {\Gamma }_{1}^{\prime } = \boldsymbol {\Gamma }_{2}\) or equivalently, for all directions \(\mathbf {z} \in \mathbb {R}^{P}\) and z ≠ 0p
This prevents the j th column of Γ2 from being in the opposite direction of the j th column of Γ1. This form of the condition is easier to incorporate into the identifiability illustration.
In the MSGHD, if we consider moving the amount t in a direction z, which entails setting x = tz, we can write the density as
Note, if z is equal to the k th eigenvector, which is the k th column of Γ, then the density reduces to
where
Therefore, the density is simply proportional to
First, note that if the parameterizations are one-to-one, then if one parameterization is shown to be identifiable, the others are identifiable as well. Similar to Browne and McNicholas (2015), we let δj = βj/Φj, \(\alpha _{j} = \sqrt { \omega _{j}/{\Phi }_{j} + {\beta _{j}^{2}}/{{\Phi }_{j}^{2}}} \) and \(\kappa _{j} = \sqrt {{\Phi }_{j} \omega _{j}} \), where αj ≥|δj|. Under this reparameterization, we now have
For large z, the Bessel function can approximated by
which yields, using the alternative parameterization,
If z is not equal to the k th eigenvector, than, using the reparameterization given in Eq. 38, we have
The characteristic function for a multiple scaled distribution can be written as
which, under the alternative parameterization from Eq. 38, becomes
Now if we consider moving t in the direction z
and, for large t, the characteristic function is
Therefore, from the condition given in Eq. 34, there exists z such that the tuple \(({\sum }_{j = 1}^{P} \kappa _{j} \left | [ \boldsymbol {\Gamma }^{\prime } \mathbf {z} ]_{j} \right |, \mathbf {z}^{\prime } \boldsymbol {\Gamma } \boldsymbol {\mu } )\) is unique for all g = 1, … , G and reduces to the univariate hyperbolic distribution, which is identifiable.
3.3 3.3 Identifiability of the Coalesced Generalized Hyperbolic Distribution
To prove the identifiability of the CGHD, we only need to show that two sets of distributions, the multiple scaled and the generalized hyperbolic distribution are disjoint. Consider moving along the k th eigenvalue such that (λk, κk) is distinct from (λ0, κ0) and the proof easily follows from the identifiability of the univariate generalized hyperbolic distribution.
Rights and permissions
About this article
Cite this article
Tortora, C., Franczak, B.C., Browne, R.P. et al. A Mixture of Coalesced Generalized Hyperbolic Distributions. J Classif 36, 26–57 (2019). https://doi.org/10.1007/s00357-019-09319-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00357-019-09319-3