Abstract
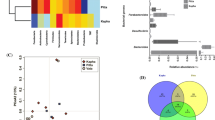
Sweat secretion keeps feet and insoles humid and rich in nutrients, which are the conditions needed to maintain abundant microbial growth. Analyzing the diversity and function of microorganisms in the insole is of great significance in the development of functional insoles and prediction of human foot hygiene condition. In this study, pure culture method, MiSeq high-throughput sequencing technology, and PICRUSt gene function prediction were used to analyze the diversity and function of the bacterial community from insoles of healthy population of different sexes and age groups. Staphylococcus, Micrococcus, and Brevibacterium are present in all insole samples, and there is no significant difference between sexes of the same age group. However, a significant difference in insole microbial population was obtained among age groups. For community function, all six samples expressed similarity in the preliminary metabolism, but in samples from the elderly, many specific catabolic genes were associated with human disease and drug resistance. This study provides a reference for the development of multi-function insoles and other sanitary products for disease prediction in a healthy population.




Similar content being viewed by others
References
Grice EA, Segre JA (2011) The skin microbiome. Nat Rev Microbiol 9:244–253. https://doi.org/10.1038/nrmicro2537
Tian M, Ding ZW, Liu XL, Wang CL, Wang ZX (2006) Research on improving the comfort of footwear by using single-guide wet technology and antibacterial technology. China Leather 18:119–121. https://doi.org/10.13536/j.cnki.issn1001-6813.2006.18.007
Li H, Zhao CQ, Zhou J, Shao HH, Chen WY (2011) Isolation, purification and identification of bacteria from the shoes worn by children. Afr J Biotechnol 10:4133–4137
Aly R (1991) Cutaneous microbiology. In: Orkin M, Maibach HI, Dahl MH (eds) Dermatology. Appleton and Lange, Los Altos, pp 22–25
Fierer N, Hamady M, Lauber CL, Knight R (2008) The influence of sex, handedness, and washing on the diversity of hand surface bacteria. Proc Natl Acad Sci USA 105:17994–17999. https://doi.org/10.1073/pnas.0807920105
Gao Z, Tseng CH, Pei Z, Blaser MJ (2007) Molecular analysis of human forearm superficial skin bacterial biota. Proc Natl Acad Sci USA 104:2927–2932. https://doi.org/10.1073/pnas.0607077104
Grice EA, Kong HH, Renaud G, Young AC, Bouffard GG, Blakesley RW et al (2008) A diversity profile of the human skin microbiota. Genome Res 18:1043–1050. https://doi.org/10.1101/gr.075549.107
Giacomoni P, Mammone T, Teri M (2009) Gender-linked differences in human skin. J Dermatol Sci 55:144–149. https://doi.org/10.1016/j.jdermsci.2009.06.001
Chen W, Thiboutot D, Zouloulis C (2002) Cutaneos androgens metabolism: basic research and clinical perspectives. Invest Dermatol 119:992–1007. https://doi.org/10.1046/j.1523-1747.2002.00613.x
James J, Leyden MD, Kenneth J, Mcginley OH, Mills MA, Albert M (1975) Age-related changes in the resident bacterial flora of the human face. J Investig Dermatol 65:379–381
Costello EK, Lauber CL, Hamady M, Fierer N, Gordon JI, Knight R (2009) Bacterial community variation in human body habitats across space and time. Science 326:1694–1697
Kong HH, Segre JA (2012) Skin microbiome: looking back to move forward. J Investig Dermatol 132:933–939. https://doi.org/10.1038/jid.2011.417
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821. https://doi.org/10.1038/nbt.2676
Koo H, Hakim JA, Morrow CD, Eipers PG, Davila A, Andersen DT et al (2017) Comparison of two bioinformatics tools used to characterize the microbial diversity and predictive functional attributes of microbial mats from Lake Obersee, Antarctica. J Microbiol Methods 140:15–22. https://doi.org/10.1016/j.mimet.2017.06.017
Kong HH (2011) Skin microbiome: genomics-based insights into the diversity and role of skin microbes. Trends Mol Med 17:320–328. https://doi.org/10.1016/j.molmed.2011.01.013
Grice EA, Kong HH, Conlan S, Deming CB, Davis J, Young AC et al (2009) Topographical and temporal diversity of the human skin microbiome. Science 324:1190–1192. https://doi.org/10.1126/science.1171700
Oh J, Byrd AL, Deming C, Conlan S, Program NCS, Kong HH, Segre JA (2014) Biogeography and individuality shape function in the human skin metagenome. Nature 514:59–64. https://doi.org/10.1038/nature13786
Myles IA, Reckhow JD, Williams KW, Sastalla I, Frank KM, Datta SK (2016) A method for culturing Gram-negative skin microbiota. BMC Microbiol. https://doi.org/10.1186/s12866-016-0684-9
Meneghetti KL, Canabarro MC, Otton LM, Hain TS, Geimba MP, Corção G (2018) Bacterial contamination of human skin allografts and antimicrobial resistance: a skin bank problem. BMC Microbiol 18:121. https://doi.org/10.1186/s12866-018-1261-1
Cogen AL, Nizet V, Gallo RL (2008) Skin microbiota: a source of disease or defence? Br J Dermatol 158:442–455. https://doi.org/10.1111/j.1365-2133.2008.08437.x
Hornef MW, Wick MJ, Rhen M, Normark S (2002) Bacterial strategies for overcoming host innate and adaptive immune responses. Nat Immunol 3:1033–1040. https://doi.org/10.1038/ni1102-1033
Gao Z, Perez-Perez GI, Chen Y, Blaser MJ (2010) Quantitation of major human cutaneous bacterial and fungal populations. J Clin Microbiol 48:3575–3581. https://doi.org/10.1128/jcm.00597-10
Somerville DA (2010) The normal flora of the skin in different age groups. Br J Dermatol 81:248–258. https://doi.org/10.1111/j.1365-2133.1969.tb13976.x
Petersen PK, Venhoet J, Sabath LD, Quie PG (1976) Extracellular and bacterial factors influencing staphylococcal phagocytosis and killing by human polymorphonuclear leukocytes. Infect Immun 14:496–501
Lai Y, Gallo RL (2010) Commensal skin bacteria as the probiotic of the cutaneous immune response. Expert Rev Dermatol 5:251–253. https://doi.org/10.1586/edm.10.24
Acknowledgements
Not applicable.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hu, X., Chang, L., Wang, Z. et al. Age- and Sex-Linked Bacterial Community Variation and Function Prediction from Insoles of Healthy Chinese Population. Indian J Microbiol 60, 222–229 (2020). https://doi.org/10.1007/s12088-020-00855-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12088-020-00855-w