Abstract
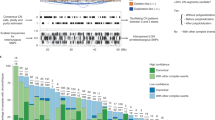
Cervical cancer is a common gynecological malignancy with high incidence and mortality. Somatic copy number alterations (CNAs) play an important role in identifying tumor suppressor genes and oncogenes and are a useful diagnostic indicator for many cancer types. However, the genomic landscape of CNAs in cervical cancer has not yet been comprehensively characterized. In the present study, we collected 974 cervical cancer samples from different data sources. All samples were analyzed by genomic arrays to obtain high-resolution CNAs. Focal genomic regions with CNA events and potential cancer driver genes were identified by GISTIC2.0. Meanwhile, we constructed a comprehensive cervical cancer database by PHP and self-written Perl and R scripts. In total, 54 recurrent regions of amplification and deletion were detected. Frequently altered tumor suppressor genes were found in these regions, including PIK3CA, ERBB2, EP300 and FBXW7. CNA hotspots and related enriched functional categories were also identified. The incidence of chromothripsis in cervical cancer was estimated to be 6.06%, and the chromosome pulverization hotspot regions were detected. Based on the curated data, we developed CNAdbCC (http://cailab.labshare.cn/CNAdbCC/), a comprehensive database for copy number alterations in cervical cancer. We provide a user-friendly Web interface for data mining and visualization. It is the most comprehensive public database devoted exclusively to genomic alterations in cervical cancer. These results extend our molecular understanding of cervical cancer. The database will enable researchers to explore specific CNA patterns in this lethal cancer and facilitate the discovery of therapeutic candidates.





Similar content being viewed by others
References
Almal SH, Padh H (2012) Implications of gene copy-number variation in health and diseases. J Hum Genet 57:6–13
Bae DS, Cho SB, Kim YJ, Whang JD, Song SY, Park CS, Kim DS, Lee JH (2001) Aberrant expression of cyclin D1 is associated with poor prognosis in early stage cervical cancer of the uterus. Gynecol Oncol 81:341–347
Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, Marshall KA, Phillippy KH, Sherman PM, Holko M, Yefanov A, Lee H, Zhang N, Robertson CL, Serova N, Davis S, Soboleva A (2013) NCBI GEO: archive for functional genomics data sets–update. Nucleic Acids Res 41:D991–995
Bengtsson H, Irizarry R, Carvalho B, Speed TP (2008) Estimation and assessment of raw copy numbers at the single locus level. Bioinformatics 24:759–767
Bengtsson H, Wirapati P, Speed TP (2009) A single-array preprocessing method for estimating full-resolution raw copy numbers from all Affymetrix genotyping arrays including GenomeWideSNP 5 and 6. Bioinformatics 25:2149–2156
Beroukhim R, Getz G, Nghiemphu L, Barretina J, Hsueh T, Linhart D, Vivanco I, Lee JC, Huang JH, Alexander S, Du J, Kau T, Thomas RK, Shah K, Soto H, Perner S, Prensner J, Debiasi RM, Demichelis F, Hatton C, Rubin MA, Garraway LA, Nelson SF, Liau L, Mischel PS, Cloughesy TF, Meyerson M, Golub TA, Lander ES, Mellinghoff IK, Sellers WR (2007) Assessing the significance of chromosomal aberrations in cancer: methodology and application to glioma. Proc Natl Acad Sci USA 104:20007–20012
Beroukhim R, Mermel CH, Porter D, Wei G, Raychaudhuri S, Donovan J, Barretina J, Boehm JS, Dobson J, Urashima M, Mc Henry KT, Pinchback RM, Ligon AH, Cho YJ, Haery L, Greulich H, Reich M, Winckler W, Lawrence MS, Weir BA, Tanaka KE, Chiang DY, Bass AJ, Loo A, Hoffman C, Prensner J, Liefeld T, Gao Q, Yecies D, Signoretti S, Maher E, Kaye FJ, Sasaki H, Tepper JE, Fletcher JA, Tabernero J, Baselga J, Tsao MS, Demichelis F, Rubin MA, Janne PA, Daly MJ, Nucera C, Levine RL, Ebert BL, Gabriel S, Rustgi AK, Antonescu CR, Ladanyi M, Letai A, Garraway LA, Loda M, Beer DG, True LD, Okamoto A, Pomeroy SL, Singer S, Golub TR, Lander ES, Getz G, Sellers WR, Meyerson M (2010) The landscape of somatic copy-number alteration across human cancers. Nature 463:899–905
Bodelon C, Vinokurova S, Sampson JN, den Boon JA, Walker JL, Horswill MA, Korthauer K, Schiffman M, Sherman ME, Zuna RE, Mitchell J, Zhang X, Boland JF, Chaturvedi AK, Dunn ST, Newton MA, Ahlquist P, Wang SS, Wentzensen N (2016) Chromosomal copy number alterations and HPV integration in cervical precancer and invasive cancer. Carcinogenesis 37:188–196
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68:394–424
Burki TK (2017) Novel mutations in cervical cancer. Lancet Oncol 18:e137
Cai H, Kumar N, Bagheri HC, von Mering C, Robinson MD, Baudis M (2014) Chromothripsis-like patterns are recurring but heterogeneously distributed features in a survey of 22,347 cancer genome screens. BMC Genom 15:82
Catarino R, Matos A, Pinto D, Pereira D, Craveiro R, Vasconcelos A, Lopes C, Medeiros R (2005) Increased risk of cervical cancer associated with cyclin D1 gene A870G polymorphism. Cancer Genet Cytogenet 160:49–54
Chiang DY, Villanueva A, Hoshida Y, Peix J, Newell P, Minguez B, LeBlanc AC, Donovan DJ, Thung SN, Solé M, Tovar V, Alsinet C, Ramos AH, Barretina J, Roayaie S, Schwartz M, Waxman S, Bruix J, Mazzaferro V, Ligon AH, Najfeld V, Friedman SL, Sellers WR, Meyerson M, Llovet JM (2008) Focal gains of VEGFA and molecular classification of hepatocellular carcinoma. Cancer Res 68:6779–6788
Crosbie EJ, Einstein MH, Franceschi S, Kitchener HC (2013) Human papillomavirus and cervical cancer. Lancet 382:889–899
Deng M, Brägelmann J, Kryukov I, Saraiva-Agostinho N, Perner S (2017) FirebrowseR: an R client to the Broad Institute’s Firehose Pipeline. Database (Oxf) 2017:2017
Dutta P, Bui T, Bauckman KA, Keyomarsi K, Mills GB, Nanjundan M (2013) EVI1 splice variants modulate functional responses in ovarian cancer cells. Mol Oncol 7:647–668
Femi OF (2018) Genetic alterations and PIK3CA gene mutations and amplifications analysis in cervical cancer by racial groups in the US. Int J Health Sci (Qassim) 12:28–32
Forbes SA, Beare D, Boutselakis H, Bamford S, Bindal N, Tate J, Cole CG, Ward S, Dawson E, Ponting L, Stefancsik R, Harsha B, Kok CY, Jia M, Jubb H, Sondka Z, Thompson S, De T, Campbell PJ (2017) COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res 45:D777–D783
Forment JV, Kaidi A, Jackson SP (2012) Chromothripsis and cancer: causes and consequences of chromosome shattering. Nat Rev Cancer 12:663–670
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, Cerami E, Sander C, Schultz N (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6:pl1
Goodman A (2015) HPV testing as a screen for cervical cancer. BMJ 350:h2372
Guan Z, Zhang J, Wang J, Wang H, Zheng F, Peng J, Xu Y, Yan M, Liu B, Cui B, Huang Y, Liu Q1 (2014) SOX1 down-regulates β-catenin and reverses malignant phenotype in nasopharyngeal carcinoma. Mol Cancer 13:257
Haeussler M, Zweig AS, Tyner C, Speir ML, Rosenbloom KR, Raney BJ, Lee CM, Lee BT, Hinrichs AS, Gonzalez JN, Gibson D, Diekhans M, Clawson H, Casper J, Barber GP, Haussler D, Kuhn RM, Kent WJ (2019) The UCSC genome browser database: 2019 update. Nucleic Acids Res 47:D853–D858
Heselmeyer K, Macville M, Schröck E, Blegen H, Hellström AC, Shah K, Auer G, Ried T (1997) Advanced-stage cervical carcinomas are defined by a recurrent pattern of chromosomal aberrations revealing high genetic instability and a consistent gain of chromosome arm 3q. Genes Chromosomes Cancer 19:233–240
Heselmeyer-Haddad K, Janz V, Castle PE, Chaudhri N, White N, Wilber K, Morrison LE, Auer G, Burroughs FH, Sherman ME, Ried T (2003) Detection of genomic amplification of the human telomerase gene (TERC) in cytologic specimens as a genetic test for the diagnosis of cervical dysplasia. Am J Pathol 163:1405–1416
Hsu F-H, Chen H-IH, Tsai M-H, Lai L-C, Huang C-C, Tu S-H, Chuang EY, Chen Y (2011) A model-based circular binary segmentation algorithm for the analysis of array CGH data. BMC Res Notes 4:394
Hu Z, Zhu D, Wang W, Li W, Jia W, Zeng X, Ding W, Yu L, Wang X, Wang L, Shen H, Zhang C, Liu H, Liu X, Zhao Y, Fang X, Li S, Chen W, Tang T, Fu A, Wang Z, Chen G, Gao Q, Li S, Xi L, Wang C, Liao S, Ma X, Wu P, Li K, Wang S, Zhou J, Wang J, Xu X, Wang H, Ma D (2015) Genome-wide profiling of HPV integration in cervical cancer identifies clustered genomic hot spots and a potential microhomology-mediated integration mechanism. Nat Genet 47:158–163
Kim T-M, Xi R, Luquette LJ, Park RW, Johnson MD, Park PJ (2013) Functional genomic analysis of chromosomal aberrations in a compendium of 8000 cancer genomes. Genome Res 23:217–227
Lin D-C, Wang M-R, Koeffler HP (2018) Genomic and epigenomic aberrations in esophageal squamous cell carcinoma and implications for patients. Gastroenterology 154:374–389
Maher EA, Furnari FB, Bachoo RM, Rowitch DH, Louis DN, Cavenee WK, DePinho RA (2001) Malignant glioma: genetics and biology of a grave matter. Genes Dev 15:1311–1333
Marioni JC, Thorne NP, Valsesia A, Fitzgerald T, Redon R, Fiegler H, Andrews TD, Stranger BE, Lynch AG, Dermitzakis ET, Carter NP, Tavaré S, Hurles ME (2007) Breaking the waves: improved detection of copy number variation from microarray-based comparative genomic hybridization. Genome Biol 8:R228
McIntyre JB, Wu JS, Craighead PS, Phan T, Köbel M, Lees-Miller SP, Ghatage P, Magliocco AM, Doll CM (2013) PIK3CA mutational status and overall survival in patients with cervical cancer treated with radical chemoradiotherapy. Gynecol Oncol 128:409–414
Mermel CH, Schumacher SE, Hill B, Meyerson ML, Beroukhim R, Getz G (2011) GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol 12:R41
Mikhail FM (2014) Copy number variations and human genetic disease. Curr Opin Pediatr 26:646–652
Ojesina AI, Lichtenstein L, Freeman SS, Pedamallu CS, Imaz-Rosshandler I, Pugh TJ, Cherniack AD, Ambrogio L, Cibulskis K, Bertelsen B, Romero-Cordoba S, Treviño V, Vazquez-Santillan K, Guadarrama AS, Wright AA, Rosenberg MW, Duke F, Kaplan B, Wang R, Nickerson E, Walline HM, Lawrence MS, Stewart C, Carter SL, McKenna A, Rodriguez-Sanchez IP, Espinosa-Castilla M, Woie K, Bjorge L, Wik E, Halle MK, Hoivik EA, Krakstad C, Gabiño NB, Gómez-Macías GS, Valdez-Chapa LD, Garza-Rodríguez ML, Maytorena G, Vazquez J, Rodea C, Cravioto A, Cortes ML, Greulich H, Crum CP, Neuberg DS, Hidalgo-Miranda A, Escareno CR, Akslen LA, Carey TE, Vintermyr OK, Gabriel SB, Barrera-Saldaña HA, Melendez-Zajgla J, Getz G, Salvesen HB, Meyerson M (2014) Landscape of genomic alterations in cervical carcinomas. Nature 506:371–375
Olshen AB, Venkatraman ES, Lucito R, Wigler M (2004) Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5:557–572
Poolkerd S, Leelahakorn S, Manusirivithaya S, Tangjitgamol S, Thavaramara T, Sukwattana P, Pataradule K (2006) Survival rate of recurrent cervical cancer patients. J Med Assoc Thail 89:275–282
Rosa-Rosa JM, Pita G, González-Neira A, Milne RL, Fernandez V, Ruivenkamp C, van Asperen CJ, Devilee P, Benitez J (2009) A 7 Mb region within 11q13 may contain a high penetrance gene for breast cancer. Breast Cancer Res Treat 118:151–159
Scotto L, Narayan G, Nandula SV, Subramaniyam S, Kaufmann AM, Wright JD, Pothuri B, Mansukhani M, Schneider A, Arias-Pulido H, Murty VV (2008) Integrative genomics analysis of chromosome 5p gain in cervical cancer reveals target over-expressed genes, including Drosha. Mol Cancer 7:58
Smith M, Marioni J, Hardcastle T, Thorne N (2006) snapCGH: segmentation, normalization and processing of aCGH data users’ guide. Bioconductor
Stephens PJ, Greenman CD, Fu B, Yang F, Bignell GR, Mudie LJ, Pleasance ED, Lau KW, Beare D, Stebbings LA, McLaren S, Lin ML, McBride DJ, Varela I, Nik-Zainal S, Leroy C, Jia M, Menzies A, Butler AP, Teague JW, Quail MA, Burton J, Swerdlow H, Carter NP, Morsberger LA, Iacobuzio-Donahue C, Follows GA, Green AR, Flanagan AM, Stratton MR, Futreal PA, Campbell PJ (2011) Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144:27–40
Tomczak K, Czerwińska P, Wiznerowicz M (2015) The cancer genome atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol (Pozn) 19:A68–77
Wang H, Schaefer T, Konantz M, Braun M, Varga Z, Paczulla AM, Reich S, Jacob F, Perner S, Moch H, Fehm TN, Kanz L, Schulze-Osthoff K, Lengerke C (2017a) Prominent oncogenic roles of EVI1 in breast carcinoma. Cancer Res 77:2148–2160
Wang Z, Sun P, Gao C, Chen J, Li J, Chen Z, Xu M, Shao J, Zhang Y, Xie J (2017b) Down-regulation of LRP1B in colon cancer promoted the growth and migration of cancer cells. Exp Cell Res 357:1–8
Yang J, Liu J, Ouyang L, Chen Y, Liu B, Cai H (2016) CTLPScanner: a web server for chromothripsis-like pattern detection. Nucleic Acids Res 44:W252–258
Yu G, Wang L-G, Han Y, He Q-Y (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16:284–287
Yu L, Jiang M, Qu P, Wu Z, Sun P, Xi M, Qin Y, Liu X, Liao G, Lei X, Sun L, Zhang Y, Li Z, Chen W, Qiao Y (2018) Clinical evaluation of human papillomavirus 16/18 oncoprotein test for cervical cancer screening and HPV positive women triage. Int J Cancer 143:813–822
Zarrei M, MacDonald JR, Merico D, Scherer SW (2015) A copy number variation map of the human genome. Nat Rev Genet 16:172–183
Funding
This work was supported by a grant from the National Natural Science Foundation of China (Grant Nos. U1603120, 31571314, 31771394).
Author information
Authors and Affiliations
Contributions
HC and PY conceived the study. HL, XX, JY, KW and CW contributed to data collection. HL and XX carried out the analysis. JY, KW and CW assisted with all the statistical analysis. HC and PY wrote the draft of the manuscript. All authors discussed the results and contributed to the final manuscript.
Corresponding authors
Ethics declarations
Conflicts of interest
There is no conflict of interest in this study.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
438_2019_1636_MOESM4_ESM.tiff
Fig. S4 An example of data series browsing. a The detailed information of queried series. b The copy number alteration frequencies for each chromosome across the data series (TIFF 473 kb)
Rights and permissions
About this article
Cite this article
Luo, H., Xu, X., Yang, J. et al. Genome-wide somatic copy number alteration analysis and database construction for cervical cancer. Mol Genet Genomics 295, 765–773 (2020). https://doi.org/10.1007/s00438-019-01636-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-019-01636-x