Abstract
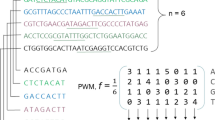
The paper discusses a hypergraph-based polynomial time algorithm for constructing a DNA sequence corresponding to a given spectrum with no errors. The presence of n-ary relations in the expression of DNA sequencing led to the advent of hypergraphs in DNA sequencing. An efficient algorithm, unimodular hypergraph (UMHG), has been proposed for constructing unimodular hypergraph that represents the short-read DNA sequence of the given spectrum. The performance of the proposed algorithm is evaluated in comparison with prominent algorithms such as greedy, greedy (lag), ant colony optimization(ACO), multi-level ACO, enhanced genetic algorithm (GA), hybrid GA and tabu search against 40 instances. The proposed UMHG algorithm is found to outperform the other algorithms in terms of average similarity score. UMHG is significant in terms of minimum computing time, especially as the spectrum size increases computing time decreases considerably, owing to the unimodularity property.


Similar content being viewed by others
References
Watson JD, Crick FHC (1953) A structure for deoxyribose nucleic acid. Nature 173:737–738
Khorlin Khrapko AA, Shik Yu VV, Lysov P, Florent’ev VL, Mirzabekov AD (1988) Determination of the nucleotide sequence of dna using hybridization with oligonucleotides. a new method. Doklady Acad Nauk Sci. USSR 303:1508–1511
Bains W, Smith GC (1988) A novel method for nucleic acid squence determination. J Theor Biol 135:303–307
Southern EM (1988) United kindom pattend application gb 8810400
Drukner I, Drmanac R, Lahat I, Crkvenjakov R (1989) Sequencing of mega base plus dna by hybridization: theory of the method. Genomics 4:114–128
Markiewicz M, Zebrowska A, Markiewicz WT, Andrych-rozek K, Astriab A (1994) Synthesis of oligonucleotides permenetly linked with solid supports for use as synthetic oligonucleotide combinatorial libraries—innovations in solid phase synthesis, biological and biomedical applications. Birmingham, Mayflower worldwide, pp 339–346
Pevzner Pavel A (1989) \(l\)-tuple dna sequencing: computer analysis. J Biomol Struct Dyn 7(1):63–73
Pevzner PA, Lipshutz RJ (1994) Towards dna sequencing chips. In: 19th conf. MAthematical foundations of computer science vol 841, pp 143–158
Berge C (1973) Graphs and hypergraphs. North-Holland Publishing Company, Amsterdam
Berge C (1989) Hypergraphs–combinatorics of finite sets. North-Holland Publishing Company, Amsterdam
Swaminathan V, Rajaram G, Abhishek V, Reddy BS, Kannan K (2017) A novel hypergraph-based genetic algorithm (hgga) built on unimodular and anti-homomorphism properties for dna sequencing by hybridization. Interdiscip Sci Comput Life Sci. https://doi.org/10.1007/s12539-017-0267-y
Uetz M (2010) Discrete Optimization 2010, Lecture 6, Total Unimodularity and Matching
Doerr Benjamin (2001) Lattice approximation and linear discrepancy of totally unimodular matrices. In: SODA ’01 Proceedings of the twelfth annual ACM-SIAM symposium on Discrete algorithms, pp 119–125
Kalita P, Kalita B (2013) A graph theoritical approach for dna sequencing. IOSR J Math 5(Issue I):40–46
Dharmarajan R, Kannan K (2010) A hypergraph based algorithm for image restoration from salt-and-pepper noise. Int J Electron Commun 64:1114–1122
Dharmarajan R, Kannan K (2012) On minimal tranversals in simple hypergraphs. Int J Comput Appl Math 7:117–122
Dharmarajan R, Kannan K (2012) Hypergraph based segmentation and edge deduction in gray images. Far East J Math Sci 68(2):185–230
Cherifi H, Bretto A, Azema J, Laget B (1997) Combinatorics and image processing. Graph Models Image Process 59:265–277
Cherifi H, Bretto A, Aboutajdine D (2002) Hypergraph imaging: an overview. Pattern Recongnition 35:651–658
Bretto A, Gillibert L (2005) Hypergraph based image representations. GbRPR, LNCS 3434:1–11
Swaminathan V, Chandrasekhara Rao K (2012) Diaspora operators. Int J Math Anal 6(6):1671–1778
Blesa MJ, Blum C, Vallès MY (2008) An ant colony optimization algorithm for dna sequencing by hybridization. Comput Oper Res 35:3620–3635
Web link www.bisc.cs.berkely.edu/flint/flintcib/presentations/wednesday/zhang_flint_cibi.pdf. (1996)
Acknowledgements
The authors thank SASTRA University for their financial and research support. Also, the authors thank the Department of Science and Technology—Fund for Improvement of S&T Infrastructure in Universities and Higher Educational Institutions Government of India (SR/FST/MSI-107/2015) for their financial support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Venkatraman, S., Rajaram, G. & Krithivasan, K. Unimodular Hypergraph for DNA Sequencing: A Polynomial Time Algorithm. Proc. Natl. Acad. Sci., India, Sect. A Phys. Sci. 90, 49–56 (2020). https://doi.org/10.1007/s40010-018-0561-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40010-018-0561-z