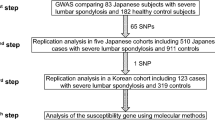
Abstract
Ankylosing spondylitis (AS) is a common complex inflammatory disease; however, up to now distinct genes with monogenic pattern have not been reported for this disease. In the present study, we report a large Iranian family with several affected members with AS. DNAs of the three affected and two healthy cases were chosen for performing whole-exome sequencing (WES). After several filtering steps, candidate variants in the following genes were detected: RELN, DNMT1, TAF4β, MUC16, DLG2, and FAM208. However, segregation analysis confirmed the association of only one variant, c.7456A>G; p.(Ser2486Gly) in the RELN gene with AS in this family. In addition, in silico predictions supported the probable pathogenicity of this variant. In this study, for the first time, we report a novel variant in the RELN gene, c.7456A>G; p.(Ser2486Gly), which completely co-segregates with AS. This association suggests potential insights into the pathophysiological bases of AS and it could broaden horizons toward new therapeutic strategies.
Similar content being viewed by others
Introduction
Ankylosing spondylitis (AS) is a chronic inflammatory disease that primarily influences the axial skeleton and sacroiliac joints. Extra-articular involvements including eyes (acute anterior uveitis), heart, and gastrointestinal tract (IBD; either Crohn’s disease or ulcerative colitis) can be manifested in AS patients [1]. For example, the pooled prevalence of acute anterior uveitis, psoriasis, and IBD in patients with AS has been reported to be 25.8%, 9.3%, and 6.8%, respectively [2]. AS is one of the famous prototypes of spondyloarthropathies, which include psoriatic arthritis, reactive arthritis, and enteropathic arthritis [3]. AS prevalence is 0.1–0.5% in Asian and European populations [4]. The hallmark characteristic of AS is new bone formation in affected tissues, which results from inflammatory mediators, but the precise etiopathogenesis of the disease has not yet been completely discovered [5].
The major genetic causes of AS remain to be identified because various detected loci, together with HLA-B*27, could explain only 24.4% of the heritability of AS [6]. To date, most of the reported variants for AS have been identified based on association studies targeting variants with mean minor allele frequency (MAF) more than 5%; nonetheless, association of AS with rare variants (MAF < 1%) has not been identified yet due to the low power of utilized preceding methods and the lack of large families with multiple affected members [7].
AS can be implied as a highly inheritable disease. The recurrence risk in monozygous twins is 63%, while it is around 8.2% in first-degree relatives [8]. The human leukocyte antigen B27 (HLA-B*27) genotype has been shown to have a strong association with AS [9]. For instance, HLA-B*27 carriers have an around 20-fold increased risk of showing spondyloarthropathy-related diseases. Although, it has been postulated that the presence of the HLA-B*27 genotype alone is not sufficient for AS development because only 1–5% HLA-B*27 carriers eventually develop AS [10].
Whole-exome sequencing (WES) is a powerful tool to identify variants inside the coding regions of the human genome, which are estimated to include about 85% of disease-causing mutations [11, 12]. Identifying new causative variants for AS can eventually give us hints about its pathogenesis, diagnosis, prevention, and therapy.
Here, we report a novel variant in the RELN gene, c.7456A>G; p.(Ser2486Gly), which completely co-segregates with AS in a large Iranian family with several affected members.
Methods
Study subjects
A large Iranian Kurdish family including seven patients with diagnosis of AS according to the 1984 Modified New York Criteria [13] referred to Rheumatology Research Center, Shariati Hospital, Tehran University of Medical Sciences. This study was approved by the Human Research Ethics Committee of Tehran University of Medical Sciences and was registered in NCBI Bioproject (Accession number: PRJNA558631; https://www.ncbi.nlm.nih.gov/bioproject/PRJNA558631). Written informed consent was obtained from all participants. Patients’ demographics, laboratory findings, physical examinations, duration of morning stiffness, pain or swelling in peripheral joints and back pain, specific AS indexes, such as Bath Ankylosing Spondylitis Disease Activity Index and Metrology Index (BASDAI and BASMI) were collected before the test (Table 1).
DNA extraction
About 10 ml peripheral blood was collected from each of the II.6, III.5, III.6, III.7, III.8, III.10, III.12, III.13, III.15, IV.5, and IV.8 individuals and genomic DNA was extracted using the standard phenol–chloroform method [14]. DNAs derived from II.6, III.6, III.7, IV.5, and IV.8 samples were selected for WES. The family’s pedigree is depicted in Fig. 1.
The DNAs from the following samples were available: II.6, III.5, III.6, III.7, III.8, III.10, III.12, III.13, III.15, IV.5, and IV.8. The III.12 and III.15 were additional affected members who were remotely related to this family. Because there were inconclusive data about the status (Healthy or Affected) of parents and former generations of III.12 and III.15, we did not directly depict the consanguineous relationships in this pedigree. The index case is indicated by an arrow. A dagger (†) indicates suspected patients; I.2 and II.5 were deceased but, according to the provided information by the family about common AS symptoms such as morning stiffness, low back pain, good response to NSAID, and peripheral arthritis, they were suspected to be affected with AS as well. An asterisk (*) indicates the samples that were selected for performing WES. The II.6 and III.15 individuals deceased during the study. The statue of HLA-B27 is shown as B27* for positive and B27(−) for negative antigen test. Besides, the genotypes of the RELN variants are indicated for all subjects.
Prediction of single-point variation on protein stability
The I-Mutant V2.0, which computes the ΔΔG values of protein variant, was utilized to predict and annotate the effect of the single nonsynonymous variant, p.(Ser2486Gly), on the protein stability from its sequence. I-Mutant V2.0 (https://www.folding.biofold.org/i-mutant/i-mutant2.0.html) is based on Support Vector Machine and ProTherm database [15] and is trained to predict the thermodynamic free energy change upon single-point variations in protein sequences.
Whole-exome sequencing (WES)
For exome enrichment, we used the TargetSeq Exome Enrichment System (Life Technologies, Carlsbad, CA, USA). Library preparation and sequencing using a SOLiD 5500xl sequencer was carried out according to the manufacturer’s instructions (Life Technologies, Carlsbad, CA, USA), generating 75 bp reads covering the enriched sequences.
We aligned the reads obtained from the sequencing runs with the human reference genome (GRCh37/hg19) using BFAST-0.6.55 [16]. Variant calling was performed with the SAM tools software [17] using reads that mapped unambiguously to the target region. The only variant calls supported by at least ten nonidentical reads, a Phred-like quality score ≥1 and an allelic percentage between 20 and 80% were considered.
The pattern of disease in the pedigree was compatible with an autosomal-dominant mode of inheritance, therefore, only variants heterozygous in all affected individuals and homozygous for the reference allele in healthy controls were considered. To exclude neutral polymorphisms, variants with an MAF greater than 0.01 using dbSNP132 (https://www.ncbi.nlm.nih.gov/SNP/), 1K genome (www.internationalgenome.org/), Exome Variant Server (http://www.evs.gs.washington.edu/EVS/), Exome Aggregation Consortium version 0.3 database (ExAC; http://www.exac.broadinstitute.org/), Genome Aggregation Database (gnomAD; https://gnomad.broadinstitute.org/) and Iranome (http://www.iranome.ir/) were filtered out.
Pathogenicity predictions were performed using at least four online databases namely SIFT (http://www.sift.bii.astar.edu.sg/), Polyphen2 (http://www.genetics.bwh.harvard.edu/pph2/), Provean (http://www.provean.jcvi.org/) and MutationTaster (http://www.mutationtaster.org/). Also, ConSurf (http://www.consurf.tau.ac.il) and Residual Variation Intolerance Score (RVIS) (http://genic-intolerance.org/) were applied to provide an evolutionary conservation profile for the reelin protein (Fig. 2).
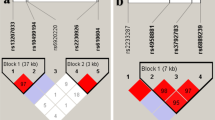
a Schematic physical and genetic maps of the RELN gene (NM_173054.2) and the c.7456A>G variant found in the AS family members. c.7456A>G is located in exon 47. b c.7456A>G is located between the R5 and R6 domain. c The amino acid sequences of RELN (p.Ser2486) is colored based on conservation scores by the ConSurf database. ConSurf demonstrates evolutionary conservation profiles for proteins of known structure in the PDB according to the phylogenetic relations between homologous sequences as well as amino acid’s structural and functional importance. d Comparison of normal and altered reelin predicted structure. Structure modeling of the normal protein and superimposed structure modeling of the variated protein; the normal and the variation sites of p.(Ser2486Gly) are emphasized by a yellow highlight and locally zoomed (Color figure online).
Sanger sequencing
Confirmation and family segregation for the final candidate variants, as well as checking out their presence in a cohort of ethnicity-matched controls was performed by Sanger sequencing. The primers listed in Supplementary Table 1 were used for PCR amplification of the variant regions. The PCRs were carried out in 20 μl volumes, containing an aqueous solution of standard 10× PCR buffer, 200 ng DNA, 0.2 μM of each primer, 1.5 mM of MgCl2 and 200 μM of dNTP mix and 0.5 U Taq polymerase. Cycling conditions were including the initial denaturation (94 °C, 3 min) followed by 40 cycles of 94 °C for 30 s, 64 °C for 30 s, 72 °C for 30 s, and a final extension at 72 °C for 10 min. Amplified DNA fragments were separated on 2% (w/v) agarose gel and viewed after staining with ethidium bromide. Consequently, the samples were sequenced using the ABI 3130xl Genetic Analyzer and sequencing chromatograms were analyzed using the CodonCode Aligner v. 5.1.5 software.
In silico structural modeling
The largest open reading frame of the RELN encodes a protein of 3460 amino acid residues. The identified c.7456A>G; p.(Ser2486Gly) variant affects the R5–R6 domain of the reelin protein. For further analysis, Phyre2 (http://www.sbg.bio.ic.ac.uk/phyre2/) and SWISS-PROT (https://www.ebi.ac.uk/swissprot/) were utilized to design the PDB file and then the variant’s impact on protein flexibility and stability were analyzed by PyMOL (https://pymol.org/) and I-Mutant V2.0. Figure 2d describes the amino acid substitution in the reelin R5/R6 domain. The RAMPAGE online tool (https://www.mordred.bioc.cam.ac.uk/~rapper/rampage.php) was applied to check out the detailed residue-by-residue stereochemical quality on the basis of Ramachandran plot.
Results
Whole-exome sequencing
Five samples, three patients and two healthy samples, were sequenced by WES with an average depth of 100×. Consequently, 9675 common heterozygote SNV and indel variants among all the three genotyped patients were detected, out of which, 2129 variants were also absent in the two healthy genotyped samples. By excluding the variants with MAF greater than 1% in publicly available databases, e.g: dbSNP, 1000 Genomes Project, ExAC and GnomAD, 260 variants were achieved. Sixteen variants from them were affecting the coding regions and only eight variants changed the amino acid codons. These eight variants reside inside the RELN, DNMT1, TAF4β, MUC16, DLG2, and FAM208 genes (Table 2). Consequently, they were selected to be verified by Sanger sequencing as well as segregation study in additional family members. Only the c.7456A>G; p.(Ser2486Gly) variant in the RELN gene co-segregated completely with the disease in the family and was absent in the studied healthy controls (Fig. 3a). The schematic presentation of the applied steps is depicted in Fig. 3b, c.
a Chromatograms showing nucleotide sequences of RELN in the region of c.7456A>G found in the family. Black arrow indicates the heterozygous nucleotide substitution. The substituted amino acid is indicated in red, whereas Ser as the original amino acid is shown in green. b Overlapping filter strategy. Red color indicates the remaining variants for further analysis that are present in three affected members (orange circles) but not among the shared variants in two controls (black circle). c Schematic representation of filtering strategies applied in this research. The filtering process was applied according to several strategies as demonstrated in the schematic representation. For further investigation, we reevaluated the filtering steps with regard to the possibility that the disease could be autosomal recessive, however, we could not detect any relevant variants according to this assumption.
All WES-derived data are available in Sequence Read Archive [18] (SRA; Accession numbers: from SAMN12565062 to SAMN12565066; https://dataview.ncbi.nlm.nih.gov/object/PRJNA558631). Besides, the novel variant, c.7456A>G, was registered in Leiden Open Variation Database [19] (LOVD; Individual ID: 00250194; https://databases.lovd.nl/shared/individuals/00250194) and ClinVar [20] (Accession number: SCV000930629: www.ncbi.nlm.nih.gov/clinvar) databases.
In silico prediction
The novel variant, p.(Ser2486Gly), was absent in 100 alleles from a cohort of ethnicity-matched controls that we checked by Sanger sequencing. In addition, the allele frequencies in different databases and in silico pathogenicity prediction using different tools are included in Table 2. Finally, a conservation study by Consurf software showed a high conservation score (score∼8) for the region of this variant (Fig. 2d).
The RAMPAGE online tool was used to elucidate detailed residue-by-residue stereochemical quality based on a Ramachandran plot by which the modeled structure of reelin R5–R6 indicated almost 98% of residues in the most favored regions, around 2% of residues in allowed regions, and only 0.3% of residues in the outlier regions, which suggested that the modeled structure of reelin R5–R6 was acceptable.
Discussion
AS is a complex inflammatory disease [21] which usually is manifested in the third decade of life and affects men more severely and frequently than women [21]. Among 32 identified loci associated with AS [22], HLA-B*27 is the most significant one, however, it does not completely segregate with the disease in the studied family here (Table 1), as there were both male and female affected members who were negative for HLA-B*27 (Fig. 1 and Table 1).
In the present study, WES was performed in order to detect the associated variant/variants with AS in a consanguineous Iranian family with multiple affected members. The study led to the identification of a novel heterozygous c.7456A>G variant in the RELN gene.
RELN gene comprises 65 exons and encodes reelin which is a large secreted extracellular matrix protein with around 3460 amino acids. It has been shown that reelin is circulating in the blood and the liver appears to be an important source [23]. This protein, controlling predominantly the cell–cell interactions, is an essential factor for neuronal migration during cortical development, neuronal maturation, and cell positioning during brain development in the prenatal period [24, 25]. Reelin is composed of a distinctive N-terminal domain followed by eight tandem repeats (Reelin Repeats; RRs) involving an extracellular growth factor (EGF)-like domain, which is surrounded by two sub-repeats (A and B) (Fig. 2) [26]. The C-terminal region comprises less than 1% of amino acids in reelin, however, it plays an important role in reelin secretion [27]. The eight RRs are structurally similar and have great conservation with an average sequence identity of 83.5–87.9% among 167 species (Fig. 2d) [26].
Mutations in RELN have been shown to be associated with autosomal recessive lissencephaly with cerebellar hypoplasia (MIM: 257320) [28], a phenotype similar to homozygous reeler mice [29], autosomal-dominant lateral temporal epilepsy and many neuropsychiatric disorders such as Alzheimer’s disease [30], autism spectrum disorders [31], and schizophrenia [32].
Based on the conservational analysis performed in this study by ConSurf and RVIS, the RELN gene is conserved, which explains the low rate of mutations in RELN and the severity of its related diseases.
The c.7456A>G or p.(Ser2486Gly) variant affects a highly conserved residue (Fig. 2c) in the R5–R6 repeat fragments of the EGF-like domain. The results of analysis by I-mutant v.2.0 [33], showed that the p.(Ser2486Gly) substitution decreases protein stability (Fig. 2c, d).
The R3–R6 repeat fragments of the EGF-like domain play a central role in reelin’s binding to the ApoER2 and VLDLR [34, 35] which they are involved in neuron signal transduction and also they have been detected in the surface of macrophages. Moreover, complementary studies have indicated that the R5–R6 region is sufficient to induce adapter protein Disabled‐1 (Dab1) phosphorylation which can promote and transfer signals to the downstream pathways [36].
The pathogenic role of immune cells, especially macrophages has repeatedly been implicated in AS. It is interesting to speculate that reelin may play an important role in AS pathogenesis through stimulating VLDLR on the surface of macrophages in AS patients [36]. The Human Protein Atlas (http://www.proteinatlas.org) shows that the reelin protein is expressed in liver, cerebellum, distinct cells of the lymph node and tonsil, in addition to HAP1, HEL, PC3, and RH30. Furthermore, the RELN transcript has been detected in multiple human tissues such as lymph node and spleen (http://www.genecards.org). In adult mice, this gene is highly expressed in immune cells as well as the nervous system (http://www.informatics.jax.org).
Reelin’s role in inflammation could be verified in autoinflammatory disorders such as rheumatoid arthritis (RA) [37]. Increased levels of reelin in both synovial fluids and sera of RA patients have been shown and reelin is being proposed as a novel diagnostic marker and therapeutic target in this disease [38]. Moreover, it has been found that RA patients express RELN and its related receptors in fibroblast-like synoviocytes (FLS) [37]. Although a probable involvement of FLS in AS has been described [39], further investigations need to be done in order to demonstrate the role of RELN in the context of autoimmune and autoinflammatory disorders.
Reeler mice have provided extra evidence on reelin involvement in inflammation. For example, it has been shown that both reeler mice and AS patients express a lower amount of IFN-γ [40, 41]. Interleukins’ vital roles in autoimmune diseases, especially AS, have been indicated [42, 43]; for instance, induced expression of COX-2 and IL-6, has been reported in patients with AS [44]. Furthermore, reeler mice exhibit higher inflammatory scores than wild-type mice, showing increased susceptibility to inflammation [45]. Similarly, Herz et al. showed that reelin increases the activity of NF-kB as an important inflammatory mediator [46]. Thus, a link between reelin and inflammatory mediators can be suggested, while further studies should be applied to shed light on its role in inflammation.
The second foremost feature of patients with AS is excess bone formation and syndesmophytes. It has been found that the Wnt pathway plays a major role in bone formation in these patients [47]. Reelin binds to other receptors such as LRP8 (low-density lipoprotein receptor-related protein 8) [48] and triggers the interaction between LRP8 and Dab1, which results in the recruitment of phosphatidylinositol‐3‐kinase and RAS [49, 50]. Increasing evidence shows that Wnt/β‐catenin signaling plays a critical role in osteoblast differentiation and bone formation [51]. Also, LRP8 mediates Wnt/β‐catenin signaling and controls osteoblast differentiation [52]. Hence, it can be inferred that reelin could play an important role in osteogenesis [52], but the precise role of the reelin signaling complex (Reelin/VLDLR/ApoER2/Dab1) in osteoblasts remains to be revealed.
The precise mechanism of reelin in AS pathogenesis is not discovered yet, but based on the clues presented here, it seems that inflammation and osteogenesis take center stage in AS development. Altogether, we hypothesize that the identified c.7456A>G; p.(Ser2486Gly) variant alters the signal transduction in macrophages, which are the essential cells in AS, to trigger inflammation.
In the present study, we utilized WES to identify gene causality in a familial case of AS. For the first time, we found the c.7456A>G; p.(Ser2486Gly) variant in the RELN gene which co-segregated completely with AS in a large family with multiple affected members. This suggests potential insights into the pathophysiological involvement of reelin via inflammation and osteogenesis pathways in AS, and it could broaden our horizon toward new therapeutic strategies.
Data availability
Human variant and phenotypes have been reported to ClinVar (Accession number: SCV000930629; www.ncbi.nlm.nih.gov/clinvar) and LOVD (Individual ID: 00250194). Whole-exome sequencing data produced in the current study have been deposited in the NCBI Sequence Read Archive (SRA) with accession numbers SAMN12565062, SAMN12565063, SAMN12565064, SAMN12565065, and SAMN12565066 and URL: https://dataview.ncbi.nlm.nih.gov/object/PRJNA558631. Also, bioproject is accessible by PRJNA558631 as an accession number and the following URL: https://www.ncbi.nlm.nih.gov/bioproject/PRJNA558631.
References
El Maghraoui A. Extra-articular manifestations of ankylosing spondylitis: prevalence, characteristics and therapeutic implications. Eur J Intern Med. 2011;22:554–60.
Stolwijk C, van Tubergen A, Castillo-Ortiz JD, Boonen A. Prevalence of extra-articular manifestations in patients with ankylosing spondylitis: a systematic review and meta-analysis. Ann Rheum Dis. 2015;74:65–73.
Schett G. Bone formation versus bone resorption in ankylosing spondylitis: molecular mechanisms of spondyloarthropathies. Springer; New York, NY. 2009;114–21.
Robinson PC, Brown MA. Genetics of ankylosing spondylitis. Mol Immunol. 2014;57:2–11.
Maksymowych WP, Chiowchanwisawakit P, Clare T, Pedersen SJ, Østergaard M, Lambert RG. Inflammatory lesions of the spine on magnetic resonance imaging predict the development of new syndesmophytes in ankylosing spondylitis: evidence of a relationship between inflammation and new bone formation. Arthritis Rheum. 2009;60:93–102.
Cortes A, Hadler J, Pointon JP, Robinson PC, Karaderi T, Leo P, et al. Identification of multiple risk variants for ankylosing spondylitis through high-density genotyping of immune-related loci. Nat Genet. 2013;45:730.
Bamshad MJ, Ng SB, Bigham AW, Tabor HK, Emond MJ, Nickerson DA, et al. Exome sequencing as a tool for Mendelian disease gene discovery. Nat Rev Genet. 2011;12:745.
Brown MA, Kennedy LG, Macgregor AJ, Darke C, Duncan E, Shatford JL, et al. Susceptibility to ankylosing spondylitis in twins the role of genes, HLA, and the environment. Arthritis Rheum. 1997;40:1823–8.
Ji X, Sun K, Hu Z, Zhang Y, Ma Y, Sun Z, et al. Comparison of clinical manifestations according to HLA-B (27) genotype in ankylosing spondylitis patients: real-world evidence from smart management system for spondyloarthritis. Zhonghua nei ke za zhi. 2018;57:179–84.
Timms AE, Crane AM, Sims A-M, Cordell HJ, Bradbury LA, Abbott A, et al. The interleukin 1 gene cluster contains a major susceptibility locus for ankylosing spondylitis. Am J Hum Genet. 2004;75:587–95.
Rabbani B, Tekin M, Mahdieh N. The promise of whole-exome sequencing in medical genetics. J Hum Genet. 2014;59:5.
Robinson PC, Leo PJ, Pointon JJ, Harris J, Cremin K, Bradbury LA, et al. Exome-wide study of ankylosing spondylitis demonstrates additional shared genetic background with inflammatory bowel disease. NPJ Genom Med. 2016;1:16008.
Moll J, Wright V. New York clinical criteria for ankylosing spondylitis. A statistical evaluation. Ann Rheum Dis. 1973;32:354.
Chomczynski P, Sacchi N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem. 1987;162:156–9.
Bava KA, Gromiha MM, Uedaira H, Kitajima K, Sarai A. ProTherm, version 4.0: thermodynamic database for proteins and mutants. Nucleic Acids Res. 2004;32:D120–1.
Homer N, Merriman B, Nelson SF. BFAST: an alignment tool for large scale genome resequencing. PloS one. 2009;4:e7767.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Leinonen R, Sugawara H, Shumway M. Collaboration INSD The sequence read archive. Nucleic Acids Res. 2010;39:D19–21.
Fokkema IF, Taschner PE, Schaafsma GC, Celli J, Laros JF, den Dunnen JT. LOVD v. 2.0: the next generation in gene variant databases. Hum Mutat. 2011;32:557–63.
Landrum MJ, Lee JM, Benson M, Brown G, Chao C, Chitipiralla S, et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 2015;44:D862–8.
Braun J, Sieper J. Ankylosing spondylitis. Lancet. 2007;369:1379–90.
O’rielly DD, Uddin M, Rahman P. Ankylosing spondylitis: beyond genome-wide association studies. Curr Opin Rheumatol. 2016;28:337–45.
Smalheiser NR, Costa E, Guidotti A, Impagnatiello F, Auta J, Lacor P, et al. Expression of reelin in adult mammalian blood, liver, pituitary pars intermedia, and adrenal chromaffin cells. Proc Natl Acad Sci. 2000;97:1281–6.
D'Arcangelo G, Miao GG, Chen S-C, Scares HD, Morgan JI, Curran T: A protein related to extracellular matrix proteins deleted in the mouse mutant reeler. Nature. 1995;374:719–23.
D’Arcangelo G, Nakajima K, Miyata T, Ogawa M, Mikoshiba K, Curran T. Reelin is a secreted glycoprotein recognized by the CR-50 monoclonal antibody. J Neurosci. 1997;17:23–31.
Ranaivoson FM, von Daake S, Comoletti D. Structural insights into reelin function: present and future. Front Cell Neurosci. 2016;10:137.
Esmaeilzadeh-Gharehdaghi E, Razmara E, Bitaraf A, Mahmoudi M, Garshasbi M. S3440P substitution in C-terminal region of human reelin dramatically impairs secretion of reelin from HEK 293T cells. Cell Mol Biol. 2019;65:12–6.
Hong SE, Shugart YY, Huang DT, Al Shahwan S, Grant PE, Hourihane JOB, et al. Autosomal recessive lissencephaly with cerebellar hypoplasia is associated with human RELN mutations. Nat Genet. 2000;26:93.
Nishibe M, Katsuyama Y, Yamashita T. Developmental abnormality contributes to cortex-dependent motor impairments and higher intracortical current requirement in the reeler homozygous mutants. Brain Struct Funct. 2018;1–13.
Seripa D, Matera MG, Franceschi M, Daniele A, Bizzarro A, Rinaldi M, et al. The RELN locus in Alzheimer’s disease. J Alzheimers Dis. 2008;14:335–44.
Lammert DB, Howell BW. RELN mutations in autism spectrum disorder. Front Cell Neurosci. 2016;10:84.
Guidotti A, Grayson DR, Caruncho HJ. Epigenetic RELN dysfunction in schizophrenia and related neuropsychiatric disorders. Front Cell Neurosci. 2016;10:89.
Capriotti E, Fariselli P, Casadio R. I-Mutant2. 0: predicting stability changes upon mutation from the protein sequence or structure. Nucleic Acids Res. 2005;33:W306–10.
Arnaud L, Ballif BA, Förster E, Cooper JA. Fyn tyrosine kinase is a critical regulator of disabled-1 during brain development. Curr Biol. 2003;13:9–17.
Suetsugu S, Tezuka T, Morimura T, Hattori M, Mikoshiba K, Yamamoto T, et al. Regulation of actin cytoskeleton by mDab1 through N-WASP and ubiquitination of mDab1. Biochem J. 2004;384:1–8.
Du Y, Yang M, Wei W, Huynh H, Herz J, Saghatelian A, et al. Macrophage VLDL receptor promotes PAFAH secretion in mother’s milk and suppresses systemic inflammation in nursing neonates. Nat Commun. 2012;3:1008.
Magnani A, Pattacini L, Boiardi L, Casali B, Salvarani C. Reelin levels are increased in synovial fluid of patients with rheumatoid arthritis. Clin Exp Rheumatol. 2010;28:546–8.
You S, Cho C-S, Lee I, Hood L, Hwang D, Kim W-U. A systems approach to rheumatoid arthritis. PLoS One 2012;7:e51508.
Inman R, El-Gabalawy H. The immunology of ankylosing spondylitis and rheumatoid arthritis: a tale of similarities and dissimilarities. Clin Exp Rheumatol. 2009;27:S26.
Greenjohnson J, Zalcman S, Vriend C, Nance D, Greenberg A. Suppressed T cell and macrophage function in the” reeler”(rl/rl) mutant, a murine strain with elevated cerebellar norepinephrine concentration. Brain Behav Immun. 1995;9:47–60.
Smith JA, Barnes MD, Hong D, DeLay ML, Inman RD, Colbert RA. Gene expression analysis of macrophages derived from ankylosing spondylitis patients reveals interferon-gamma dysregulation. Arthritis Rheum. 2008;58:1640–9.
Sabzevary-Ghahfarokhi M, Shohan M, Shirzad H, Rahimian G, Bagheri N, Soltani A, et al. The expression analysis of Fra-1 gene and IL-11 protein in Iranian patients with ulcerative colitis. BMC Immunol. 2018;19:17.
van den Berg R, Jongbloed EM, de Schepper EIT, Bierma-Zeinstra SMA, Koes BW, Luijsterburg PAJ. The association between pro-inflammatory biomarkers and nonspecific low back pain: a systematic review. Spine J. 2018.
Dougados M, Béhier JM, Jolchine I, Calin A, van der Heijde D, Olivieri I, et al. Efficacy of celecoxib, a cyclooxygenase 2–specific inhibitor, in the treatment of ankylosing spondylitis: A six‐week controlled study with comparison against placebo and against a conventional nonsteroidal antiinflammatory drug. Arthritis Rheum. 2001;44:180–5.
Carvajal AE, Vázquez-Carretero MD, García-Miranda P, Peral MJ, Calonge ML, Ilundain AA. Reelin expression is up-regulated in mice colon in response to acute colitis and provides resistance against colitis. Biochim Biophys Acta. 2017;1863:462–73.
Ding Y, Huang L, Xian X, Yuhanna IS, Wasser CR, Frotscher M, et al. Loss of Reelin protects against atherosclerosis by reducing leukocyte–endothelial cell adhesion and lesion macrophage accumulation. Sci Signal. 2016;9:ra29.
Lories RJ, Luyten FP, De Vlam K. Progress in spondylarthritis. Mechanisms of new bone formation in spondyloarthritis. Arthritis Res Ther. 2009;11:221.
Lee GH, D’Arcangelo G. New insights into reelin-mediated signaling pathways. Front Cell Neurosci. 2016;10:122.
D’Arcangelo G, Homayouni R, Keshvara L, Rice DS, Sheldon M, Curran T. Reelin is a ligand for lipoprotein receptors. Neuron. 1999;24:471–9.
Bock HH, Jossin Y, Liu P, Förster E, May P, Goffinet AM, et al. Phosphatidylinositol 3-kinase interacts with the adaptor protein Dab1 in response to Reelin signaling and is required for normal cortical lamination. J Biol Chem. 2003;278:38772–9.
Day TF, Guo X, Garrett-Beal L, Yang Y. Wnt/β-catenin signaling in mesenchymal progenitors controls osteoblast and chondrocyte differentiation during vertebrate skeletogenesis. Dev Cell. 2005;8:739–50.
Zhang J, Zhang X, Zhang L, Zhou F, van Dinther M, ten Dijke P. LRP8 mediates Wnt/β‐catenin signaling and controls osteoblast differentiation. J Bone Min Res. 2012;27:2065–74.
Acknowledgements
We thank the family for their valuable contributions. We are also grateful for Corinna Jensen, Helmholtz-Zentrum Geesthacht, University of Hamburg, and Christian Sperling for their excellent technical help and Robert Weissmann and Dr Farveh Ehya for their supports in bioinformatics analysis.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: MM, ARJ, and MG. Conducted the experiments: MG, EEGH, ER, EF, ARB, SA, SHP, SMA, and LRJ. Analyzed and interpreted the data: MG, LRJ, EEGH, and ER. Contributed reagents/materials/analysis tools: MM, MG, MV, AWK. Wrote the paper: MG, ARB, EEGH, and ER. All authors read and approved the final paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Garshasbi, M., Mahmoudi, M., Razmara, E. et al. Identification of RELN variant p.(Ser2486Gly) in an Iranian family with ankylosing spondylitis; the first association of RELN and AS. Eur J Hum Genet 28, 754–762 (2020). https://doi.org/10.1038/s41431-020-0573-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-020-0573-4
This article is cited by
-
High genetic heterogeneity of leukodystrophies in Iranian children: the first report of Iranian Leukodystrophy Registry
neurogenetics (2023)
-
Prenatal stress perturbs fetal iron homeostasis in a sex specific manner
Scientific Reports (2022)
-
Novel phenotype and genotype spectrum of WDR62 in two patients with associated primary autosomal recessive microcephaly
Irish Journal of Medical Science (1971 -) (2022)
-
Novel phenotype and genotype spectrum of NARS2 and literature review of previous mutations
Irish Journal of Medical Science (1971 -) (2022)
-
Novel manifestations of Warburg micro syndrome type 1 caused by a new splicing variant of RAB3GAP1: a case report
BMC Neurology (2021)