Key message
Overexpression of K2-NhaD in transgenic cotton resulted in phenotypes with strong salinity and drought tolerance in greenhouse and field experiments, increased expression of stress-related genes, and improved regulation of metabolic pathways, such as the SOS pathway.
Abstract
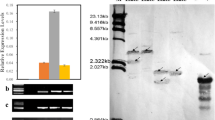
Drought and salinity are major abiotic stressors which negatively impact cotton yield under field conditions. Here, a plasma membrane Na+/H+ antiporter gene, K2-NhaD, was introduced into upland cotton R15 using an Agrobacterium tumefaciens-mediated transformation system. Homozygous transgenic lines K9, K17, and K22 were identified by PCR and glyphosate-resistance. TAIL-PCR confirmed that T-DNA carrying the K2-NhaD gene in transgenic lines K9, K17 and K22 was inserted into chromosome 3, 19 and 12 of the cotton genome, respectively. Overexpression of K2-NhaD in transgenic cotton plants grown in greenhouse conditions and subjected to drought and salinity stress resulted in significantly higher relative water content, chlorophyll, soluble sugar, proline levels, and SOD, CAT, and POD activity, relative to non-transgenic plants. The expression of stress-related genes was significantly upregulated, and this resulted in improved regulation of metabolic pathways, such as the salt overly sensitive pathway. K2-NhaD transgenic plants growing under field conditions displayed strong salinity and drought tolerance, especially at high levels of soil salinity and drought. Seed cotton yields in transgenic line were significantly higher than in wild-type plants. In conclusion, the data indicate that K2-NhaD transgenic lines have great potential for the production of stress-tolerant cotton under field conditions.






Similar content being viewed by others
References
Abraham E, Hourton-Cabassa C, Erdei L, Szabados L (2010) Methods for determination of proline in plants. Methods Mol Biol 639:317–331
Aebi H (1984) Catalase in vitro. Methods Enzymol 105:121–126
Agarwal PK, Agarwal P, Reddy MK, Sopory SK (2006) Role of DREB transcription factors in abiotic and biotic stress tolerance in plants. Plant Cell Rep 25(12):1263–1274
Barajas-Lopez JD, Moreno JR, Gamez-Arjona FM, Pardo JM, Punkkinen M, Zhu JK, Quintero FJ, Fujii H (2018) Upstream kinases of plant SnRKs are involved in salt stress tolerance. Plant J 93(1):107–118
Barragan V, Leidi EO, Andres Z, Rubio L, De Luca A, Fernandez JA, Cubero B, Pardo JM (2012) Ion exchangers NHX1 and NHX2 mediate active potassium uptake into vacuoles to regulate cell turgor and stomatal function in Arabidopsis. Plant Cell 24(3):1127–1142
Bassil E, Ohto M-a, Esumi T, Tajima H, Zhu Z, Cagnac O, Belmonte M, Peleg Z, Yamaguchi T, Blumwald E (2011a) The Arabidopsis intracellular Na+/H+ Antiporters NHX5 and NHX6 are endosome associated and necessary for plant growth and development. Plant Cell 23(1):224–239
Bassil E, Tajima H, Liang YC, Ohto M, Ushijima K, Nakano R, Esumi T, Coku A, Belmonte M, Blumwald E (2011b) The Arabidopsis Na+/H+ antiporters NHX1 and NHX2 control vacuolar pH and K+ homeostasis to regulate growth, flower development, and reproduction. Plant Cell 23(9):3482–3497
Brini F, Hanin M, Mezghani I, Berkowitz GA, Masmoudi K (2007) Overexpression of wheat Na+/H+ antiporter TNHX1 and H+-pyrophosphatase TVP1 improve salt- and drought-stress tolerance in Arabidopsis thaliana plants. J Exp Bot 58(2):301–308
Chen X, Lu X, Shu N, Wang D, Wang S, Wang J, Guo L, Guo X, Fan W, Lin Z et al (2017) GhSOS1, a plasma membrane Na+/H+ antiporter gene from upland cotton, enhances salt tolerance in transgenic Arabidopsis thaliana. PLoS ONE 12(7):e0181450
Cheng C, Zhang Y, Chen XG, Song JL, Guo ZQ, Li KP, Zhang KW (2018) Co-expression of AtNHX1 and TsVP improves the salt tolerance of transgenic cotton and increases seed cotton yield in a saline field. Mol Breed 38:19
Chou KC, Shen HB (2008) Cell-PLoc: a package of Web servers for predicting subcellular localization of proteins in various organisms. Nat Protoc 3(2):153–162
Chu XQ, Wang C, Chen XB, Lu WJ, Li H, Wang XL, Hao LL, Guo XQ (2015) The cotton WRKY gene GhWRKY41 positively regulates salt and drought stress tolerance in transgenic Nicotiana benthamiana. PLoS ONE 10(11):e0143022
Cuming AC, Cho SH, Kamisugi Y, Graham H, Quatrano RS (2007) Microarray analysis of transcriptional responses to abscisic acid and osmotic, salt, and drought stress in the moss, Physcomitrella patens. New Phytol 176:275–287
Gao M, Wang L, Chen SF (2012) Metagenome cloning and functional analysis of Na+/H+ antiporter genes from Keke Salt Lake in China. Curr Microbiol 64(2):179–184
Guo WF, Kevin YW, Wang N, Li J, Li GQ, Liu DH (2018) Rapid and convenient transformation of cotton (Gossypium hirsutum L.) using in planta shoot apex via glyphosate selection. J Integr Agr 17(10):2196–2203
Hasanuzzaman M, Oku H, Nahar K, Bhuyan MHMB, Al Mahmud J, Baluska F, Fujita M (2018) Nitric oxide-induced salt stress tolerance in plants: ROS metabolism, signaling, and molecular interactions. Plant Biotechnol Rep 12:77–92
Hodges DM, DeLong JM, Forney CF, Prange RK (1999) Improving the thiobarbituric acid-reactive-substances assay for estimating lipid peroxidation in plant tissues containing anthocyanin and other interfering compounds. Planta 207(4):604–611
Hooks TN, Picchioni GA, Schutte BJ, Shukla MK, Daniel DL (2018) Sodium chloride effects on seed germination, growth, and water use of Lepidium alyssoides, L. draba, and L. latifolium: traits of resistance and implications for invasiveness on saline soils. Rangel Ecol Manag 71(4): 433–442
Hraskova M, Papouskova K, Sychrova H, Zimmermannova O (2018) Length of the cytoplasmic C-terminal part of yeast Nha1 Na+/H+ antiporter influences its plasma-membrane targeting in cells lacking Erv14 COPII cargo receptor. Febs Open Bio 8:366–366
Huang YJ, Zhao HX, Gao F, Yao PF, Deng RY, Li CL, Chen H, Wu Q (2018) A R2R3-MYB transcription factor gene, FtMYB13, from Tartary buckwheat improves salt/drought tolerance in Arabidopsis. Plant Physiol and Biochem 132(2018):238–248
Ishitani M, Liu JP, Halfter U, Kim CS, Shi WM, Zhu JK (2000) SOS3 function in plant salt tolerance requires N-myristoylation and calcium binding. Plant Cell 12(9):1667–1677
Jia Q, Zheng C, Sun S, Amjad H, Liang K, Lin W (2018) The role of plant cation/proton antiporter gene family in salt tolerance. Biol Plantarum 62(4):617–629
Josephs TM, Morison IM, Day CL, Wilbanks SM, Ledgerwood EC (2014) Enhancing the peroxidase activity of cytochromec by mutation of residue 41: implications for the peroxidase mechanism and cytochrome c release. Biochem J 458:259–265
Kronzucker HJ, Britto DT (2011) Sodium transport in plants: a critical review. New Phytol 189(1):54–81
Leidi EO, Barragan V, Rubio L, El-Hamdaoui A, Ruiz MT, Cubero B, Fernandez JA, Bressan RA, Hasegawa PM, Quintero FJ et al (2010) The AtNHX1 exchanger mediates potassium compartmentation in vacuoles of transgenic tomato. Plant J 61(3):495–506
Li N, Wang X, Ma B, Du C, Zheng L, Wang Y (2017) Expression of a Na(+)/H(+) antiporter RtNHX1 from a recretohalophyte Reaumuria trigyna improved salt tolerance of transgenic Arabidopsis thaliana. J Plant Physiol 218(2017):109–120
Li S, Luo W, Jia ZH, Tang SC, Chen C (2018) The effect of natural rainfall on salt leaching under watertable management. Land Degrad Dev 29:1953–1961
Lu X, Zhang XF, Duan H, Lian CL, Liu C, Yin WL, Xia XL (2018) Three stress-responsive NAC transcription factors from Populus euphratica differentially regulate salt and drought tolerance in transgenic plants. Physiol Plant 162(1):73–97
Maghsoudi K, Emam Y, Niazi A, Pessarakli M, Arvin MJ (2018) P5CS expression level and proline accumulation in the sensitive and tolerant wheat cultivars under control and drought stress conditions in the presence/absence of silicon and salicylic acid. J Plant Interact 13(1):461–471
McCord JM, Fridovich I (1969) Superoxide dismutase. An enzymic function for erythrocuprein (hemocuprein). J Biol Chem 244(22): 6049–6055
McCue KF, Hanson AD (1992) Salt-inducible betaine aldehyde dehydrogenase from sugar beet: cDNA cloning and expression. Plant Mol Biol 18(1):1–11
Mishra S, Alavilli H, Lee BH, Panda SK, Sahoo L (2014) Cloning and functional characterization of a vacuolar Na+/H+ antiporter gene from mungbean (VrNHX1) and its ectopic expression enhanced salt tolerance in Arabidopsis thaliana. PLoS ONE 9(10):e106678
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Ohta M, Hayashi Y, Nakashima A, Hamada A, Tanaka A, Nakamura T, Hayakawa T (2002) Introduction of a Na+/H+ antiporter gene from Atriplex gmelini confers salt tolerance to rice. FEBS Lett 532(3):279–282
Park DY, Shim Y, Gi E, Lee BD, An G, Kang K, Paek NC (2018) The MYB-related transcription factor RADIALIS-LIKE3 (OsRL3) functions in ABA-induced leaf senescence and salt sensitivity in rice. Environ Exp Bot 156:86–95
Paterson AH, Brubaker CL, Wendel JF (1993) A rapid method for extraction of cotton (Gossypium spp.) genomic DNA suitable for RFLP or PCR analysis. Plant Mol Biol Rep 11(2):122–127
Per TS, Khan NA, Reddy PS, Masood A, Hasanuzzaman M, Khan MIR, Anjum NA (2017) Approaches in modulating proline metabolism in plants for salt and drought stress tolerance: phytohormones, mineral nutrients and transgenics. Plant Physiol Bioch 115(2017):126–140
Ritchie RJ (2006) Consistent sets of spectrophotometric chlorophyll equations for acetone, methanol and ethanol solvents. Photosynth Res 89(1):27–41
Shen GX, Wei J, Qiu XY, Hu RB, Kuppu S, Auld D, Blumwald E, Gaxiola R, Payton P, Zhang H (2015) Co-overexpression of AVP1 and AtNHX1 in cotton further improves drought and salt tolerance in transgenic cotton plants. Plant Mol Biol Rep 33(2):167–177
Song JL, Zhang R, Yue D, Chen XG, Guo ZQ, Cheng C, Hu MH, Zhang JR, Zhang KW (2018) Co-expression of ApGSMT2g and ApDMT2g in cotton enhances salt tolerance and increases seed cotton yield in saline fields. Plant Sci 274(2018):369–382
Verma D, Singla-Pareek SL, Rajagopal D, Reddy MK, Sopory SK (2007) Functional validation of a novel isoform of Na+/H+ antiporter from Pennisetum glaucum for enhancing salinity tolerance in rice. J Biosci Bioeng 32(3):621–628
Wang CL, Lu GQ, Hao YQ, Guo HM, Guo Y, Zhao J, Cheng HM (2017a) ABP9, a maize bZIP transcription factor, enhances tolerance to salt and drought in transgenic cotton. Planta 246(3):453–469
Wang N, Qi HK, Qiao WQ, Shi JB, Xu QH, Zhou H, Yan GT, Huang Q (2017b) Cotton (Gossypium hirsutum L.) genotypes with contrasting K+/Na+ ion homeostasis: implications for salinity tolerance. Acta Physiol Plant 39:77
Wang LF, Zhu JF, Li XM, Wang SM, Wu J (2018) Salt and drought stress and ABA responses related to bZIP genes from V. radiata and V. angularis. Gene 651(2018): 152–160
Wani SH, Tripathi P, Zaid A, Challa GS, Kumar A, Kumar V, Upadhyay J, Joshi R, Bhatt M (2018) Transcriptional regulation of osmotic stress tolerance in wheat (Triticum aestivum L.). Plant Mol Biol 97(6):469–487
Wu CA, Yang GD, Meng QW, Zheng CC (2004) The cotton GhNHX1 gene encoding a novel putative tonoplast Na+/H+ antiporter plays an important role in salt stress. Plant Cell Physiol 45(5):600–607
Wu SJ, Wang HH, Li FF, Chen TZ, Zhang J, Jiang YJ, Ding YZ, Guo WZ, Zhang TZ (2008) Enhanced Agrobacterium-mediated transformation of embryogenic calli of upland cotton via efficient selection and timely subculture of somatic embryos. Plant Mol Biol Rep 26(3):174–185
Wu H, Zhu M, Shabala L, Zhou M, Shabala S (2015) K+ retention in leaf mesophyll, an overlooked component of salinity tolerance mechanism: a case study for barley. J Integr Plant Biol 57(2):171–185
Yemm EW, Willis AJ (1954) The estimation of carbohydrates in plant extracts by anthrone. Biochem J 57(3):508–514
Zhao F, Wang Z, Zhang Q, Zhao Y, Zhang H (2006) Analysis of the physiological mechanism of salt-tolerant transgenic rice carrying a vacuolar Na+/H+ antiporter gene from Suaeda salsa. J Plant Res 119(2):95–104
Zhou SF, Chen XY, Zhang XG, Li YX (2008) Improved salt tolerance in tobacco plants by co-transformation of a betaine synthesis gene BADH and a vacuolar Na+/H+ antiporter gene SeNHX1. Biotechnol Lett 30(2):369–376
Zhu JK (2001) Plant salt tolerance. Trends Plant Sci 6:66–71
Funding
This work was supported by the National GMO New Cultivar Science and Technology grant (2016ZX08005-004) entitled: “GM cotton resistance to drought and salinity.
Author information
Authors and Affiliations
Contributions
WG performed the experiments, conceived experiments and research plans, analyzed the data, and participated in the writing of the manuscript; GL and NW provided technical assistance to WG; CY, YZ and HP participated in part of experiments; SC and DL supervised the experiments, conceived research plans, and complemented the writing of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Guo, W., Li, G., Wang, N. et al. A Na+/H+ antiporter, K2-NhaD, improves salt and drought tolerance in cotton (Gossypium hirsutum L.). Plant Mol Biol 102, 553–567 (2020). https://doi.org/10.1007/s11103-020-00969-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-020-00969-1