Abstract
Insect herbivory is pervasive in plant communities, but its impact on microbial plant colonizers is not well-studied in natural systems. By calibrating sequencing-based bacterial detection to absolute bacterial load, we find that the within-host abundance of most leaf microbiome (phyllosphere) taxa colonizing a native forb is amplified within leaves affected by insect herbivory. Herbivore-associated bacterial amplification reflects community-wide compositional shifts towards lower ecological diversity, but the extent and direction of such compositional shifts can be interpreted only by quantifying absolute abundance. Experimentally eliciting anti-herbivore defences reshaped within-host fitness ranks among Pseudomonas spp. field isolates and amplified a subset of putatively phytopathogenic P. syringae in a manner causally consistent with observed field-scale patterns. Herbivore damage was inversely correlated with plant reproductive success and was highly clustered across plants, which predicts tight co-clustering with putative phytopathogens across hosts. Insect herbivory may thus drive the epidemiology of plant-infecting bacteria as well as the structure of a native plant microbiome by generating variation in within-host bacterial fitness at multiple phylogenetic and spatial scales. This study emphasizes that ‘non-focal’ biotic interactions between hosts and other organisms in their ecological settings can be crucial drivers of the population and community dynamics of host-associated microbiomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All sequence data has been deposited to the NIH Sequence Read Archive (SRA) under accession SUB5541275 and can also be accessed via BioProject ID PRJNA587302 (http://www.ncbi.nlm.nih.gov/bioproject/587302). Saved model objects are available for download from Dryad Digital Repository44.
Code availability
Code to reproduce all analyses, figures and tables is available at https://github.com/phumph/coinfection.
References
Lloyd-Smith, J. O., Poss, M. & Grenfell, B. T. HIV-1/parasite co-infection and the emergence of new parasite strains. Parasitology 135, 795–806 (2008).
Laine, A.-L. Context-dependent effects of induced resistance under co-infection in a plant-pathogen interaction. Evol. Appl. 4, 696–707 (2011).
Tollenaere, C., Susi, H. & Laine, A.-L. Evolutionary and epidemiological implications of multiple infection in plants. Trends Plant Sci. 21, 80–90 (2016).
Karvonen, A., Jokela, J. & Laine, A.-L. Importance of sequence and timing in parasite coinfections. Trends Parasitol. 35, 109–118 (2018).
Halliday, F. W., Umbanhowar, J. & Mitchell, C. E. Interactions among symbionts operate across scales to influence parasite epidemics. Ecol. Lett. 20, 1285–1294 (2017).
Susi, H., Barrès, B., Vale, P. F. & Laine, A.-L. Co-infection alters population dynamics of infectious disease. Nat. Commun. 6, 5975 (2015).
Horton, M. W. et al. Genome-wide association study of Arabidopsis thaliana leaf microbial community. Nat. Commun. 5, 5320 (2014).
Bulgarelli, D. et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488, 91–95 (2012).
Bodenhausen, N., Bortfeld-Miller, M., Ackermann, M. & Vorholt, J. A. A synthetic community approach reveals plant genotypes affecting the phyllosphere microbiota. PLoS Genet. 10, e1004283 (2014).
Edwards, J. et al. Structure, variation, and assembly of the root-associated microbiomes of rice. Proc. Natl Acad. Sci. USA 112, E911–E920 (2015).
Wagner, M. R. et al. Host genotype and age shape the leaf and root microbiomes of a wild perennial plant. Nat. Commun. 7, 12151 (2016).
Finkel, O. M., Castrillo, G., Herrera Paredes, S., Salas González, I. & Dangl, J. L. Understanding and exploiting plant beneficial microbes. Curr. Opin. Plant. Biol. 38, 155–163 (2017).
Orozco-Mosqueda, M. D. C., Rocha-Granados, M. D. C., Glick, B. R. & Santoyo, G. Microbiome engineering to improve biocontrol and plant growth-promoting mechanisms. Microbiol. Res. 208, 25–31 (2018).
Maron, J. L. & Crone, E. Herbivory: effects on plant abundance, distribution and population growth. Proc. Biol. Sci. 273, 2575–2584 (2006).
Agrawal, A. A. Induced responses to herbivory and increased plant performance. Science 279, 1201–1202 (1998).
Bressan, M. et al. Exogenous glucosinolate produced by Arabidopsis thaliana has an impact on microbes in the rhizosphere and plant roots. ISME J. 3, 1243–1257 (2009).
Thaler, J. S., Humphrey, P. T. & Whiteman, N. K. Evolution of jasmonate and salicylate signal crosstalk. Trends Plant Sci. 17, 260–270 (2012).
Wagner, M. R. et al. Natural soil microbes alter flowering phenology and the intensity of selection on flowering time in a wild Arabidopsis relative. Ecol. Lett. 17, 717–726 (2014).
Humphrey, P. T. et al. Aversion and attraction to harmful plant secondary compounds jointly shape the foraging ecology of a specialist herbivore. Ecol. Evol. 6, 3256–3268 (2016).
Humphrey, P. T., Nguyen, T. T., Villalobos, M. M. & Whiteman, N. K. Diversity and abundance of phyllosphere bacteria are linked to insect herbivory. Mol. Ecol. 23, 1497–1515 (2014).
Vandeputte, D. et al. Quantitative microbiome profiling links gut community variation to microbial load. Nature 551, 507–511 (2017).
Turcotte, M. M., Davies, T. J., Thomsen, C. J. M. & Johnson, M. T. J. Macroecological and macroevolutionary patterns of leaf herbivory across vascular plants. Proc. Biol. Sci. 281, 20140555 (2014).
Hirano, S. S. & Upper, C. D. Bacteria in the leaf ecosystem with emphasis on Pseudomonas syringae-a pathogen, ice nucleus, and epiphyte. Microbiol. Mol. Biol. Rev. 64, 624–653 (2000).
Lindow, S. E. & Brandl, M. T. Microbiology of the phyllosphere. Appl. Environ. Microbiol. 69, 1875–1883 (2003).
Cui, J. et al. Pseudomonas syringae manipulates systemic plant defenses against pathogens and herbivores. Proc. Natl Acad. Sci. USA 102, 1791–1796 (2005).
Züst, T. et al. Natural enemies drive geographic variation in plant defenses. Science 338, 116–119 (2012).
Underwood, N., Anderson, K. & Inouye, B. D. Induced vs. constitutive resistance and the spatial distribution of insect herbivores among plants. Ecology 86, 594–602 (2005).
Shaw, D. J. & Dobson, A. P. Patterns of macroparasite abundance and aggregation in wildlife populations: a quantitative review. Parasitology 111, S111–S127 (1995).
Karban, R. & Baldwin, I. T. Induced Responses to Herbivory (Univ. Chicago Press, 1997).
Wittstock, U. et al. Successful herbivore attack due to metabolic diversion of a plant chemical defense. Proc. Natl Acad. Sci. USA 101, 4859–4864 (2004).
Alexandre, N. M. et al. Habitat preference of an herbivore shapes the habitat distribution of its host plant. Ecosphere 9, e02372 (2018).
Gloor, G. B., Macklaim, J. M., Pawlowsky-Glahn, V. & Egozcue, J. J. Microbiome datasets are compositional: and this is not optional. Front. Microbiol. 8, 2224 (2017).
Falony, G. et al. Population-level analysis of gut microbiome variation. Science 352, 560–564 (2016).
Raes, J. Editorial overview: it’s the ecology, stupid: microbiome research in the post-stamp collecting age. Curr. Opin. Microbiol. 44, iv–v (2018).
Stämmler, F. et al. Adjusting microbiome profiles for differences in microbial load by spike-in bacteria. Microbiome 4, 28 (2016).
Lovell, D., Pawlowsky-Glahn, V., Egozcue, J. J., Marguerat, S. & Bähler, J. Proportionality: a valid alternative to correlation for relative data. PLoS Comput. Biol. 11, e1004075 (2015).
Foster, K. R. & Bell, T. Competition, not cooperation, dominates interactions among culturable microbial species. Curr. Biol. 22, 1845–1850 (2012).
Thompson, L. R. et al. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature 551, 457–463 (2017).
Lundberg, D. S., Yourstone, S., Mieczkowski, P., Jones, C. D. & Dangl, J. L. Practical innovations for high-throughput amplicon sequencing. Nat. Methods 10, 999–1002 (2013).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Vehtari, A., Gelman, A. & Gabry, J. Practical Bayesian model evaluation using leave-one-out cross-validation and WAIC. Stat. Comput. 27, 1413–1432 (2017).
Humphrey, P. T. et al. Heritable plant phenotypes track light and herbivory levels at fine spatial scales. Oecologia 187, 427–445 (2018).
Lebeis, S. L. et al. Salicylic acid modulates colonization of the root microbiome by specific bacterial taxa. Science 349, 860–864 (2015).
Humphrey, P. T. & Whiteman, N. K. Dryad Data from: Insect herbivory reshapes a native leaf microbiome. (Dryad Digital Repository, 2019); https://doi.org/10.5061/dryad.qz612jm95
Acknowledgements
P.T.H. and N.K.W. gratefully acknowledge funding from the National Science Foundation (Grant Nos. DEB-1309493 to P.T.H. and DEB-1256758 to N.K.W.), the National Institute of General Medical Sciences of the National Institutes of Health (Grant No. R35GM119816 to N.K.W.), as well as the Rocky Mountain Biological Laboratory. We are indebted to field assistance provided by H. Briggs, K. Cromwell, A. Koning, L. Anderson, K. Niezgoda, D. Picklum and N. Alexandre; bioinformatics advice from T. K. O’Connor; and laboratory assistance from H. Pyon and A. Abidov. We thank our contacts at Argonne National Laboratory, S. Owens and J. Koval, for their technical expertise and support.
Author information
Authors and Affiliations
Contributions
P.T.H. and N.K.W. designed the study. P.T.H. carried out the study and analysed the data. P.T.H. and N.K.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
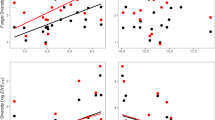
Extended Data Fig. 1 Correspondence between predicted bacterial abundance and herbivory effects between sites EL and NP.
a. Plotted are median (circles) ± 95% posterior distributions of predicted abundance for bacterial bASVs summed at the family level for sites EL (x-axis) and NP (y-axis). b. Comparison between the magnitudes of \({\mathrm{log}}_{2}\)-fold differences between damaged and undamaged leaves at sites EL (x-axis) and NP (y-axis). Middle 50%-ile and 95%-iles of median effects (circles) are depicted by thick and thin bars, respectively. On both plots, we also show data summed across all taxa in the dataset (‘all Bacteria’).
Extended Data Fig. 2 Plant patches with high herbivory harbor a disproportionate fraction of the estimated population of the most abundant P. syringae bASV.
Scaling of percentile rank (high to low) of patch-level herbivore load with total population-level percentage of bacterial propagules present in the sampled patch. At site NP, the top 20% of plant patches with the most herbivory harbor > 50% of bacterial propagules in the plant population.
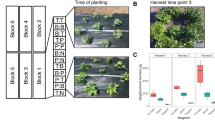
Extended Data Fig. 3 Effects of hormone pretreatment effects on levels of S. nigrita herbivory in bittercress populations at site NP.
a. For each plot separately, plotted is the average (± 1 std error) leaf mines per stem calculated at the patch level (n=16 stems per patch) for mock-treated (x-axis) versus hormonetreated (y-axis) patches. The three panels represent plots assigned to each of the three conditions: mock (that is, control), jasmonic acid (JA) or salicylic acid (SA). b. Histograms of patchlevel leaf miner damage broken down by patch-level treatment. c. Marginal effects for estimates of patch-level treatment on total mined leaves per patch (see table S5 for statistical results). Black bars are posterior means, while thick and thin bars comprises middle 50%- and 95%-iles of posterior distributions of model terms marginalized over all other parameters.
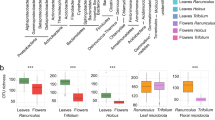
Extended Data Fig. 4 Hormone pre-treatment (Mock, JA, or SA) five weeks prior to sampling does not leave a clear signature in the distribution of γ across samples for undamaged (gray) or herbivore damaged (orange) leaf samples.
For each of the 14 most abundant bacterial families, in addition to all Bacteria, we have plotted the distributions of raw γ for damaged and undamaged samples across mock- (M), JA-, or SA-treated plant patches. Black bars represent medians for the respective distribution, and data points are slightly jittered along the x-axis. Systematic differences in γ can be easily seen between damaged versus undamaged sample classes, whereas no systematic differences can be seen between the different hormone treatment classes within each damage class. If any effects of hormone treatments are indeed present, they do not constitute a discernible feature of these data, supporting the choice to devote minimal attention to this aspect of our experiment.
Supplementary information
Supplementary Information
Supplementary Methods, Figs. 1–7 and Tables 1–5.
Rights and permissions
About this article
Cite this article
Humphrey, P.T., Whiteman, N.K. Insect herbivory reshapes a native leaf microbiome. Nat Ecol Evol 4, 221–229 (2020). https://doi.org/10.1038/s41559-019-1085-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-1085-x
This article is cited by
-
From phyllosphere to insect cuticles: silkworms gather antifungal bacteria from mulberry leaves to battle fungal parasite attacks
Microbiome (2024)
-
Induced responses contribute to rapid adaptation of Spirodela polyrhiza to herbivory by Lymnaea stagnalis
Communications Biology (2024)
-
Small world but large differences: cultivar-specific secondary metabolite-mediated phyllosphere fungal homeostasis in tea plant (Camellia sinensis)
Plant and Soil (2024)
-
Reciprocal influence of soil, phyllosphere, and aphid microbiomes
Environmental Microbiome (2023)
-
Exploration of phyllosphere microbiomes in wheat varieties with differing aphid resistance
Environmental Microbiome (2023)