Abstract
The RAS–RAF–MEK–ERK signaling axis is frequently activated in human cancers. Physiological concentrations of ATP prevent formation of RAF kinase-domain (RAFKD) dimers that are critical for activity. Here we present a 2.9-Å-resolution crystal structure of human BRAFKD in complex with MEK and the ATP analog AMP-PCP, revealing interactions between BRAF and ATP that induce an inactive, monomeric conformation of BRAFKD. We also determine how 14-3-3 relieves the negative regulatory effect of ATP through a 2.5-Å-resolution crystal structure of the BRAFKD–14-3-3 complex, in which dimeric 14-3-3 enforces a dimeric BRAFKD assembly to increase BRAF activity. Our data suggest that most oncogenic BRAF mutations alter interactions with ATP and counteract the negative effects of ATP binding by lowering the threshold for RAF dimerization and pathway activation. Our study establishes a framework for rationalizing oncogenic BRAF mutations and provides new avenues for improved RAF-inhibitor discovery.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lavoie, H. & Therrien, M. Regulation of RAF protein kinases in ERK signalling. Nat. Rev. Mol. Cell Biol. 16, 281–298 (2015).
Haling, J. R. et al. Structure of the BRAF–MEK complex reveals a kinase activity independent role for BRAF in MAPK signaling. Cancer Cell 26, 402–413 (2014).
Thorson, J. A. et al. 14-3-3 proteins are required for maintenance of Raf-1 phosphorylation and kinase activity. Mol. Cell. Biol. 18, 5229–5238 (1998).
Tzivion, G., Luo, Z. J. & Avruch, J. A dimeric 14-3-3 protein is an essential cofactor for Raf kinase activity. Nature 394, 88–92 (1998).
Rajakulendran, T., Sahmi, M., Lefrancois, M., Sicheri, F. & Therrien, M. A dimerization-dependent mechanism drives RAF catalytic activation. Nature 461, 542–545 (2009).
Kubiniok, P., Lavoie, H., Therrien, M. & Thibault, P. Time-resolved phosphoproteome analysis of paradoxical RAF activation reveals novel targets of ERK. Mol. Cell Proteom. 16, 663–679 (2017).
Kondo, Y. et al. Cryo-EM structure of a dimeric B-Raf:14-3-3 complex reveals asymmetry in the active sites of B-Raf kinases. Science 366, 109–115 (2019).
Park, E. et al. Architecture of autoinhibited and active BRAF–MEK1–14-3-3 complexes. Nature 575, 545–550 (2019).
Tate, J. G. et al. COSMIC: the catalogue of somatic mutations in cancer. Nucleic Acids Res. 47, D941–D947 (2019).
Poulikakos, P. I. et al. RAF inhibitor resistance is mediated by dimerization of aberrantly spliced BRAF(V600E). Nature 480, 387–390 (2011).
Vido, M. J., Le, K., Hartsough, E. J. & Aplin, A. E. BRAF splice variant resistance to RAF inhibitor requires enhanced MEK association. Cell Rep. 25, 1501–1510 (2018).
Hatzivassiliou, G. et al. RAF inhibitors prime wild-type RAF to activate the MAPK pathway and enhance growth. Nature 464, 431–435 (2010).
Poulikakos, P. I., Zhang, C., Bollag, G., Shokat, K. M. & Rosen, N. RAF inhibitors transactivate RAF dimers and ERK signalling in cells with wild-type BRAF. Nature 464, 427–430 (2010).
Carnahan, J. et al. Selective and potent Raf inhibitors paradoxically stimulate normal cell proliferation and tumor growth. Mol. Cancer Ther. 9, 2399–2410 (2010).
Heidorn, S. J. et al. Kinase-dead BRAF and oncogenic RAS cooperate to drive tumor progression through CRAF. Cell 140, 209–221 (2010).
Freeman, A. K., Ritt, D. A. & Morrison, D. K. The importance of Raf dimerization in cell signaling. Small GTPases 4, 180–185 (2013).
Durrant, D. E. & Morrison, D. K. Targeting the Raf kinases in human cancer: the Raf dimer dilemma. Br. J. Cancer 118, 3–8 (2018).
Lavoie, H. et al. Inhibitors that stabilize a closed RAF kinase domain conformation induce dimerization. Nat. Chem. Biol. 9, 428–436 (2013).
Bridges, D. & Moorhead, G. B. 14-3-3 proteins: a number of functions for a numbered protein. Sci. STKE 2005, re10 (2005).
Muslin, A. J., Tanner, J. W., Allen, P. M. & Shaw, A. S. Interaction of 14-3-3 with signaling proteins is mediated by the recognition of phosphoserine. Cell 84, 889–897 (1996).
Knighton, D. R. et al. Structure of a peptide inhibitor bound to the catalytic subunit of cyclic adenosine monophosphate-dependent protein kinase. Science 253, 414–420 (1991).
Taylor, S. S. & Radzio-Andzelm, E. Three protein kinase structures define a common motif. Structure 2, 345–355 (1994).
Xie, T. et al. Structural insights into RIP3-mediated necroptotic signaling. Cell Rep. 5, 70–78 (2013).
Huang, C. S. et al. Crystal structure of Ripk4 reveals dimerization-dependent kinase activity. Structure 26, 767–777 (2018).
Wan, P. T. et al. Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell 116, 855–867 (2004).
Molzan, M. et al. Stabilization of physical RAF/14-3-3 interaction by cotylenin A as treatment strategy for RAS mutant cancers. ACS Chem. Biol. 8, 1869–1875 (2013).
Morrison, D. K., Heidecker, G., Rapp, U. R. & Copeland, T. D. Identification of the major phosphorylation sites of the Raf-1 kinase. J. Biol. Chem. 268, 17309–17316 (1993).
Shaw, A. S., Kornev, A. P., Hu, J., Ahuja, L. G. & Taylor, S. S. Kinases and pseudokinases: lessons from RAF. Mol. Cell. Biol. 34, 1538–1546 (2014).
Thevakumaran, N. et al. Crystal structure of a BRAF kinase domain monomer explains basis for allosteric regulation. Nat. Struct. Mol. Biol. 22, 37–43 (2015).
Schreiber, G., Haran, G. & Zhou, H. X. Fundamental aspects of protein–protein association kinetics. Chem. Rev. 109, 839–860 (2009).
Morrison, J. F. Kinetics of the reversible inhibition of enzyme-catalysed reactions by tight-binding inhibitors. Biochim. Biophys. Acta 185, 269–286 (1969).
Yao, Z. et al. Tumours with class 3 BRAF mutants are sensitive to the inhibition of activated RAS. Nature 548, 234–238 (2017).
Yao, Z. et al. RAF inhibitor PLX8394 selectively disrupts BRAF dimers and RAS-independent BRAF-mutant-driven signaling. Nat. Med. 25, 284–291 (2019).
Michaud, N. R. et al. KSR stimulates Raf-1 activity in a kinase-independent manner. Proc. Natl Acad. Sci. USA 94, 12792–12796 (1997).
Lavoie, H. et al. MEK drives BRAF activation through allosteric control of KSR proteins. Nature 554, 549–553 (2018).
Verlande, A. et al. Metabolic stress regulates ERK activity by controlling KSR-RAF heterodimerization. EMBO Rep. 19, 320–336 (2018).
Tsai, J. et al. Discovery of a selective inhibitor of oncogenic B-Raf kinase with potent antimelanoma activity. Proc. Natl Acad. Sci. USA 105, 3041–3046 (2008).
Mccoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Buss, O., Rudat, J. & Ochsenreither, K. FoldX as protein engineering tool: better than random based approaches? Comput. Struct. Biotechnol. J. 16, 25–33 (2018).
Schrodinger, L. L. C. The PyMOL Molecular Graphics System, Version 1.3r1. (2010).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Bern, M., Cai, Y. H. & Goldberg, D. Lookup peaks: a hybrid of de novo sequencing and database search for protein identification by tandem mass spectrometry. Anal. Chem. 79, 1393–1400 (2007).
Bern, M., Kil, Y. J. & Becker, C. Byonic: advanced peptide and protein identification software. Curr. Protoc. Bioinformatics 40, 13.20.1–13.20.14 (2012).
Acknowledgements
We thank the BioMolecular Resources (BMR) group at Genentech for their support in generating constructs and biomass for protein production; Stanford Synchrotron Radiation Lightsource (SSRL) and Advanced Light Source (ALS) for access to the synchrotrons; and W. Wang, C. Noland for help with data collection and C. Sneeringer for assistance with enzyme assays. Use of the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, is supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences under Contract No. DE-AC02-76SF00515. The SSRL Structural Molecular Biology Program is supported by the DOE Office of Biological and Environmental Research, and by the National Institutes of Health, National Institute of General Medical Sciences (P41GM103393). The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views of NIGMS or NIH. Beamline 5.0.2 of the Advanced Light Source, a DOE Office of Science User Facility under Contract No. DE-AC02-05CH11231, is supported in part by the ALS-ENABLE program funded by the National Institutes of Health, National Institute of General Medical Sciences, grant P30 GM124169-01.
Author information
Authors and Affiliations
Contributions
N.P.D.L, T.J.W and J.S. designed, generated and biochemically characterized proteins and their complexes. N.P.D.L, S.G.H. and J.S. solved crystal structures. M.S., A.S.S. and J.G.Q. designed experiments and generated SPR data. F.J.I. and S.I. ran MD simulations. W.P., J.T. and P.L. generated mass spectrometry data. T.J.W., N.P.D.L., S.G.H., S.M., A.S.S. and J.S. analyzed all the data and wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
All authors were employees and/or shareholders of Genentech/Roche at the time the work was done.
Additional information
Peer review information Inês Chen was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 ATP analog interacts intimately with and stabilizes monomers of BRAF and CRAF kinase domains.
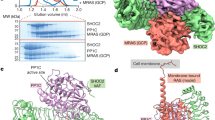
a, SDS-PAGE gels from SEC traces in Fig. 1a. b, Size exclusion chromatography traces (left) showing CRAFKD-MEK1 complex alone (blue), in the presence of ACP (red) and in the presence of GDC-0879, a RAF dimer promoter (magenta). SDS-PAGE analysis of size exclusion experiments (right). The positions of CRAF and MEK1 are labeled to the right of each gel. Elution volumes at top correspond to all gels. c, BRAF-ACP key interactions diagram. d, BRAF-ACP interaction with key residues highlighted. Blue mesh represents electron density of ACP bound to BRAF, 2Fo-Fc map contoured at 2 σ.
Extended Data Fig. 2 14-3-3 induced RAF dimer (+/- MEK) is stable in presence of ATP analog when C-terminus of RAF is phosphorylated.
a, CRAFKD(FCT) (left) and BRAFKD(CFT) co-purify as a 2:2 complex with 14-3-3 as judged by preparative SEC (top) and SDS-PAGE (bottom). b, Identification of phosphorylation on BRAF-S729 and CRAF-S621 quantified at >90% by LC-MS/MS analysis. c, Proteins co-purifying with RAFKD(FCT) identified as insect 14-3-3ζ and 14-3-3ε by LC-MS/MS analysis. d, SDS-PAGE analysis of size exclusion experiments in Fig. 2a. The corresponding SEC trace for each gel is labeled to the left, while the positions of CRAF, BRAF, MEK and 14-3-3 are labeled to the right of each gel. Elution volumes at top correspond to all gels.
Extended Data Fig. 3 BRAFKD:14-3-3 complex assembled in vitro results in active conformation of BRAFKD, with BRAFKD dimer and 14-3-3 dimer flexibly tethered by phosphopeptide:14-3-3 interactions.
a, Identification of phosphorylation on pBRAFKD(BRAFKD-14m phosphorylated at S729) - S729 phosphorylation was quantified at >99% by LC-MS/MS analysis. b, SDS-PAGE analysis of size exclusion experiments in Fig. 2b. c, Alignment of the BRAF dimer (green) in an active conformation from the previously reported BRAFKD:MEK1 hetero-tetramer structure (PDB ID = 4MNE) aligned with the BRAF dimer (blue) from the BRAFKD:14-3-3 hetero-tetramer in Fig. 2c. d, Equilibrium molecular dynamics simulation showing crystal lattice induced BRAF-14-3-3 interface is transient, with increased solvent accessibility between BRAF and 14-3-3 in 3/5 simulations over time (top left, green, blue and black vs red and purple) and a decreased number of intermolecular contacts in 3/5 simulations over time (bottom left). Right panels show initial (cyan) and dissociated (red) conformations and beginning and ending non-pS729 region contacts (bottom). e, BRAF:BRAF, 14-3-3:14-3-3 and pS729 contacts do not change over time, indicating a real interaction.
Extended Data Fig. 4 Model of BRAF-KD:MEK:14-3-3 hexamer and unambiguous electron density of pS729 in BRAF bound to 14-3-3.
a, Model of hetero-hexamer formed by MEK1 (red, salmon), BRAF (blue, marine) and 14-3-3 (orange, yellow). Compared to the BRAF/14-3-3 crystal structure (Fig. 2c), the BRAF dimer must undock from the α9 helix of one 14-3-3 protomer to bind MEK1 (as observed by SEC in Fig. 2a). Undocking is easily feasible (inset) while maintaining pS729 binding (peptide shown in pink) to 14-3-3. b, Detailed view of one BRAF-pSer729 (blue) – 14-3-3 (orange) interaction. Electron density map is 2Fo – Fc contoured to 2.0σ.
Extended Data Fig. 5 Active site titration of various BRAF variants to estimate active enzyme concentration using Morrison kinetics.
(Left) Active site titration of GDC-0879 against indicated enzymes. Activities for each enzyme were normalized to a no inhibitor control and fitted to Morrison Ki. For BRAFKD, low inhibitor concentration points below the observed paradoxical activation maximum activity were removed to allow for fitting of Morrison Ki. Each point represents three time points assayed in duplicate (n=6), normalized to reaction time. Error bars represent SD of six replicates. (Right) Zoomed version of active site titration of BRAFKD including all inhibitor points, showing paradoxical activation.
Extended Data Fig. 6 Comparison of BRAF:14-3-3 complex from this study (x-ray) with recently published BRAF:14-3-3 complexes (CryoEM).
a, Crystal structure of BRAF dimer (blue) bound to 14-3-3 (yellow, orange) reported in this work, aligned by 14-3-3 dimer onto the BRAF:14-3-3 structure (left) reported in PDB: 6UAN, and (right) the BRAF:MEK1:14-3-3 structure reported by in PDB:6Q0J (BRAF-14-3-3 in grey, MEK1 in pink). b, Overlay of structures aligned by BRAF dimer, with structure pairs and coloring as described in a.
Extended Data Fig. 7 Conserved residues in all RAF isoforms stabilize ACP induced inactive state, which is disrupted by activating mutations like V600E in BRAF.
a, Alignment of ARAF, BRAF and CRAF across species shows conservation of key residues involved in the hydrophobic packing interactions displayed in Fig. 3c. b, BRAFKD-V600E:MEK1 complex alone (top), in the presence of ACP (middle) and in the presence of GDC-0879 (bottom) from SEC experiments in main Fig. 3c were analyzed by SDS-PAGE. The positions of MEK1 and BRAFV600E are labeled to the right of each gel.
Extended Data Fig. 8 BRAF inhibitors that are “Paradox busters” disrupt BRAF dimers by causing steric clash at the dimer interface.
BRAFKD dimer (yellow) overlaid with two PLX-4720 bound BRAFKD monomers (PDB code 4WO5) (blue), aligned by their C-lobes. While most of the N-lobe and C-lobe regions overlay well, PLX4720 induces a large shift of the aC helix, causing a steric clash at the dimer interface, destabilizing the dimer interface. This is likely the mechanism of the current series of “paradox buster” molecules. With PLX4720, this shift can be accommodated in a BRAF dimer, however this requires slight conformational changes at the dimer interface and in the binding orientation of one of two bound PLX4720 molecules (PDB code 3C4C).
Supplementary information
Supplementary Video 1
Conformational changes at the BRAF dimer interface upon ACP binding
Supplementary Video 2
Conformational changes in BRAF upon ACP binding (only one molecule shown)
Supplementary Video 3
Molecular dynamics simulation showing that the BRAF N-lobe–14-3-3 interaction is unstable and that the BRAF dimer and 14-3-3 dimer interaction via phospho-peptide–14-3-3 interaction is stable
Supplementary Video 4
Alignment of all known BRAFKD crystal structures suggesting that the BRAF-ATP analogue-bound structure is unique amongst all known BRAF kinase domain structures
Source data
Source Data Fig. 3
Data used to generate curves in Fig. 3 for SPR data (BRAF–14-3-3 binding) and kinase activity data for various forms of BRAF tested
Rights and permissions
About this article
Cite this article
Liau, N.P.D., Wendorff, T.J., Quinn, J.G. et al. Negative regulation of RAF kinase activity by ATP is overcome by 14-3-3-induced dimerization. Nat Struct Mol Biol 27, 134–141 (2020). https://doi.org/10.1038/s41594-019-0365-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0365-0
This article is cited by
-
Live-cell target engagement of allosteric MEKi on MEK–RAF/KSR–14-3-3 complexes
Nature Chemical Biology (2024)
-
Structural insights into regulation of the PEAK3 pseudokinase scaffold by 14-3-3
Nature Communications (2023)
-
SHOCing RAF into action
Nature Structural & Molecular Biology (2022)
-
Structural basis for SHOC2 modulation of RAS signalling
Nature (2022)
-
Structural insights into the BRAF monomer-to-dimer transition mediated by RAS binding
Nature Communications (2022)