Abstract
Virulence mechanisms typically evolve through the continual interaction of a pathogen with its host. In contrast, it is poorly understood how environmentally acquired pathogens are able to cause disease without prior interaction with humans. Here, we provide experimental evidence for the model that Legionella pathogenesis in humans results from the cumulative selective pressures of multiple amoebal hosts in the environment. Using transposon sequencing, we identify Legionella pneumophila genes required for growth in four diverse amoebae, defining universal virulence factors commonly required in all host cell types and amoeba-specific auxiliary genes that determine host range. By comparing genes that promote growth in amoebae and macrophages, we show that adaptation of L. pneumophila to each amoeba causes the accumulation of distinct virulence genes that collectively allow replication in macrophages and, in some cases, leads to redundancy in this host cell type. In contrast, some bacterial proteins that promote replication in amoebae restrict growth in macrophages. Thus, amoebae-imposed selection is a double-edged sword, having both positive and negative impacts on disease. Comparing the genome composition and host range of multiple Legionella species, we demonstrate that their distinct evolutionary trajectories in the environment have led to the convergent evolution of compensatory virulence mechanisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Tn-seq raw sequence data supporting the findings of this study are available through the NCBI Sequence Read Archive, accession number PRJNA593054. Fitness ratios, Z scores and statistical test values for all genes examined by Tn-seq are presented in Supplementary Table 1. All other data supporting the findings of this study are available in the indicated Source Data. All plasmids and strains developed during the course of this work will be provided to all investigators upon request.
Code availability
The custom script generated in this study for Tn-seq data analysis is available through GitHub at OConnorLab/TnseqSA.
References
Young, B. C. et al. Severe infections emerge from commensal bacteria by adaptive evolution. eLife 6, e30637 (2017).
Horwitz, M. A. Formation of a novel phagosome by the Legionnaires’ disease bacterium (Legionella pneumophila) in human monocytes. J. Exp. Med. 158, 1319–1331 (1983).
Fliermans, C. B. et al. Ecological distribution of Legionella pneumophila. Appl. Environ. Microbiol. 41, 9–16 (1981).
Dilger, T., Melzl, H. & Gessner, A. Legionella contamination in warm water systems: a species-level survey. Int. J. Hyg. Environ. Health 221, 199–210 (2018).
Horwitz, M. A. & Silverstein, S. C. Legionnaires’ disease bacterium (Legionella pneumophila) multiplies intracellularly in human monocytes. J. Clin. Invest. 66, 441–450 (1980).
Yu, V. L., Zuravleff, J. J., Gavlik, L. & Magnussen, M. H. Lack of evidence for person-to-person transmission of Legionnaires’ disease. J. Infect. Dis. 147, 362 (1983).
Edelstein, P. H., Meyer, R. D. & Finegold, S. M. Laboratory diagnosis of Legionnaires’ disease. Am. Rev. Resp. Dis. 121, 317–327 (1980).
Schlossberg, D. & Bonoan, J. Legionella and immunosuppression. Semin. Resp. Infect. 13, 128–131 (1998).
Lanternier, F. et al. Incidence and risk factors of Legionella pneumophila pneumonia during anti-tumor necrosis factor therapy: a prospective French study. Chest 144, 990–998 (2013).
Segal, G. & Shuman, H. A. Intracellular multiplication and human macrophage killing by Legionella pneumophila are inhibited by conjugal components of IncQ plasmid RSF1010. Mol. Microbiol. 30, 197–208 (1998).
Vogel, J. P., Andrews, H. L., Wong, S. K. & Isberg, R. R. Conjugative transfer by the virulence system of Legionella pneumophila. Science 279, 873–876 (1998).
de Felipe, K. S. et al. Evidence for acquisition of Legionella type IV secretion substrates via interdomain horizontal gene transfer. J. Bacteriol. 187, 7716–7726 (2005).
Huang, L. et al. The E Block motif is associated with Legionella pneumophila translocated substrates. Cell Microbiol. 13, 227–245 (2011).
Zhu, W. et al. Comprehensive identification of protein substrates of the Dot/Icm type IV transporter of Legionella pneumophila. PLoS ONE 6, e17638 (2011).
Berger, K. H. & Isberg, R. R. Two distinct defects in intracellular growth complemented by a single genetic locus in Legionella pneumophila. Mol. Microbiol. 7, 7–19 (1993).
Brand, B. C., Sadosky, A. B. & Shuman, H. A. The Legionella pneumophila icm locus: a set of genes required for intracellular multiplication in human macrophages. Mol. Microbiol. 14, 797–808 (1994).
M Luo, Z. Q. & Isberg, R. R. Multiple substrates of the Legionella pneumophila Dot/Icm system identified by interbacterial protein transfer. Proc. Natl Acad. Sci. USA 101, 841–846 (2004).
O’Connor, T. J., Boyd, D., Dorer, M. S. & Isberg, R. R. Aggravating genetic interactions allow a solution to redundancy in a bacterial pathogen. Science 338, 1440–1444 (2012).
Ghosh, S. & O’Connor, T. J. Beyond paralogs: the multiple layers of redundancy in bacterial pathogenesis. Front. Cell Infect. Microbiol. 7, 467 (2017).
O’Connor, T. J., Adepoju, Y., Boyd, D. & Isberg, R. R. Minimization of the Legionella pneumophila genome reveals chromosomal regions involved in host range expansion. Proc. Natl Acad. Sci. USA 108, 14733–14740 (2011).
Hahn, M. W. & Höfle, M. G. Grazing of protozoa and its effect on populations of aquatic bacteria. FEMS Microbiol. Ecol. 35, 113–121 (2001).
Boamah, D. K., Zhou, G., Ensminger, A. W. & O’Connor, T. J. From many hosts, one accidental pathogen: the diverse protozoan hosts of Legionella. Front. Cell Infect. Microbiol. 7, 477 (2017).
Amaro, F., Wang, W., Gilbert, J. A., Anderson, O. R. & Shuman, H. A. Diverse protist grazers select for virulence-related traits in Legionella. ISME J. 9, 1607–1618 (2015).
Segal, G. & Shuman, H. A. Legionella pneumophila utilizes the same genes to multiply within Acanthamoeba castellanii and human macrophages. Infect. Immun. 67, 2117–2124 (1999).
Harb, O. S., Gao, L. Y. & Abu Kwaik, Y. From protozoa to mammalian cells: a new paradigm in the life cycle of intracellular bacterial pathogens. Envrion. Microbiol. 2, 251–265 (2000).
Molmeret, M., Horn, M., Wagner, M., Santic, M. & Abu Kwaik, Y. Amoebae as training grounds for intracellular bacterial pathogens. Appl. Environ. Microbiol. 71, 20–28 (2005).
Ensminger, A. W. Legionella pneumophila, armed to the hilt: justifying the largest arsenal of effectors in the bacterial world. Curr. Opin. Microbiol. 29, 74–80 (2016).
Gomez-Valero, L. & Buchrieser, C. Intracellular parasitism, the driving force of evolution of Legionella pneumophila and the genus Legionella. Genes Immun. 20, 394–402 (2019).
Scheikl, U. et al. Free-living amoebae (FLA) co-occurring with legionellae in industrial waters. Eur. J. Protistol. 50, 422–429 (2014).
Marciano-Cabral, F., Jamerson, M. & Kaneshiro, E. S. Free-living amoebae, Legionella and Mycobacterium in tap water supplied by a municipal drinking water utility in the USA. J. Water Health 8, 71–82 (2010).
Ruggiero, M. A. et al. A higher level classification of all living organisms. PLoS ONE 10, e0119248 (2015).
van Opijnen, T., Bodi, K. L. & Camilli, A. Tn-seq: high-throughput parallel sequencing for fitness and genetic interaction studies in microorganisms. Nat. Methods 6, 767–772 (2009).
Isaac, D. T., Laguna, R. K., Valtz, N. & Isberg, R. R. MavN is a Legionella pneumophila vacuole-associated protein required for efficient iron acquisition during intracellular growth. Proc. Natl Acad. Sci. USA 112, e5208–e5217 (2015).
Portier, E. et al. IroT/mavN, a new iron-regulated gene involved in Legionella pneumophila virulence against amoebae and macrophages. Environ. Microbiol. 17, 1338–1350 (2015).
Laguna, R. K., Creasey, E. A., Li, Z., Valtz, N. & Isberg, R. R. A. Legionella pneumophila-translocated substrate that is required for growth within macrophages and protection from host cell death. Proc. Natl Acad. Sci. USA 103, 18745–18750 (2006).
Shames, S. R. et al. Multiple Legionella pneumophila effector virulence phenotypes revealed through high-throughput analysis of targeted mutant libraries. Proc. Natl Acad. Sci. USA 114, e10446–e10454 (2017).
Burstein, D. et al. Genomic analysis of 38 Legionella species identifies large and diverse effector repertoires. Nat. Genet. 48, 167–175 (2016).
Gomez-Valero, L. et al. More than 18,000 effectors in the Legionella genus genome provide multiple, independent combinations for replication in human cells. Proc. Natl Acad. Sci. USA 116, 2265–2273 (2019).
Abu Khweek, A., Kanneganti, A., Guttridge, D. D. C. & Amer, A. O. The sphingosine-1-phosphate lyase (LegS2) contributes to the restriction of Legionella pneumophila in murine macrophages. PLoS ONE 11, e0146410 (2016).
Ensminger, A. W., Yassin, Y., Miron, A. & Isberg, R. R. Experimental evolution of Legionella pneumophila in mouse macrophages leads to strains with altered determinants of environmental survival. PLoS Pathog. 8, e1002731 (2012).
Feeley, J. C. et al. Charcoal-yeast extract agar: primary isolation medium for Legionella pneumophila. J. Clin. Microbiol. 10, 437–441 (1979).
Ensminger, A. W. & Isberg, R. R. E3 ubiquitin ligase activity and targeting of BAT3 by multiple Legionella pneumophila translocated substrates. Infect. Immun. 78, 3905–3919 (2010).
Sundstrom, C. & Nilsson, K. Establishment and characterization of a human histiocytic lymphoma cell line (U-937). Int. J. Cancer 17, 565–577 (1976).
Chien, M. et al. The genomic sequence of the accidental pathogen Legionella pneumophila. Science 305, 1966–1968 (2004).
Rao, C., Benhabib, H. & Ensminger, A. W. Phylogenetic reconstruction of the Legionella pneumophila Philadelphia-1 laboratory strains through comparative genomics. PLoS ONE 8, e64129 (2013).
Moffat, J. F. & Tompkins, L. S. A quantitative model of intracellular growth of Legionella pneumophila in Acanthamoeba castellanii. Infect. Immun. 60, 296–301 (1992).
Afgan, E., Baker, D., Batut, B., van den Beek, M. & Bouvier, D. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 46, W537–W544 (2018).
McCoy, K. M., Antonio, M. L. & van Opijnen, T. MAGenTA: a Galaxy implemented tool for complete Tn-seq analysis and data visualization. Bioinformatics 33, 2781–2783 (2017).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Sassetti, C. M., Boyd, D. H. & Rubin, E. J. Comprehensive identification of conditionally essential genes in mycobacteria. Proc. Natl Acad. Sci. USA 98, 12712–12717 (2001).
Iglewicz B. & Hoaglin D. C. How to Detect and Handle Outliers (ASQC, 1993).
Kropp, K. P. The Rorschah Z score. J. Proj. Tech. 19, 443–452 (1955).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. 57, 289–300 (1995).
Merriam, J. J., Mathur, R., Maxfield-Boumil, R. & Isberg, R. R. Analysis of the Legionella pneumophila fliI gene: intracellular growth of a defined mutant defective for flagellum biosynthesis. Infect. Immun. 65, 2497–2501 (1997).
Stewart, C. R., Rossier, O. & Cianciotto, N. P. Surface translocation by Legionella pneumophila: a form of sliding motility that is dependent upon type II protein secretion. J. Bacteriol. 191, 1537–1546 (2009).
Kolter, R., Inuzuka, M. & Helinski, D. R. Trans-complementation-dependent replication of a low molecular weight origin fragment from plasmid R6K. Cell 15, 1199–1208 (1978).
Sexton, J. A. et al. The Legionella pneumophila PilT homologue DotB exhibits ATPase activity that is critical for intracellular growth. J. Bacteriol. 186, 1658–1666 (2004).
Bardill, J. P., Miller, J. L. & Vogel, J. P. IcmS-dependent translocation of SdeA into macrophages by the Legionella pneumophila type IV secretion system. Mol. Microbiol. 56, 90–103 (2005).
O’Connor, T. J. et al. Iron limitation triggers early egress by the intracellular bacterial pathogen Legionella pneumophila. Infect. Immun. 84, 2185–2197 (2016).
Zera, K. & Zastre, J. Thiamine deficiency activates hypoxia inducible factor-1α to facilitate pro-apoptotic responses in mouse primary astrocytes. PLoS ONE 12, e0186707 (2017).
Stamatakis, A. et al. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690 (2006).
Cui, X. & Churchill, G. A. Statistical tests for differential expression in cDNA microarray experiments. Genome Biol. 4, 210 (2003).
Acknowledgements
We thank J. Vogel, M. Hossain and S. Rego for thoughtful review of the manuscript. We are grateful to R. Isberg (Tufts University School of Medicine) for sharing data, R. Isberg, J. Vogel (Washington University School of Medicine) and N. Cianciotto (Northwestern University School of Medicine) for plasmids and/or strains, and H. Shuman (University of Chicago School of Medicine) for Legionella species. This work was supported by the United States National Institutes of Health (grant no. AI119580-01) to T.J.O.
Author information
Authors and Affiliations
Contributions
T.J.O. generated the Legionella transposon mutant libraries. J.M.P. performed Tn-seq screens. T.J.O. performed TraSH experiments in U937 cells. J.M.P., S.G. and T.J.O. performed Tn-seq and TraSH data analysis. T.J.O. performed comparative analysis between amoebae and U937 macrophages data sets. T.J.O. performed experiments to assess growth kinetic parameters in amoebae and macrophages. J.M.P., S.G. and T.J.O. contributed to reagent preparations and Legionella plasmid and strain construction. J.M.P. and S.G. performed in vitro growth experiments of L. pneumophila null mutants. J.M.P. and T.J.O. performed L. pneumophila null mutant phenotypic analyses in amoebae and macrophages. S.G. and T.J.O. performed genetic interaction analysis experiments in macrophages. S.G. performed null mutant phenotypic analysis of L. pneumophila strains Paris and 130b. J.M.P., S.G. and T.J.O. performed in trans complementation experiments. T.J.O. and J.M.P. performed nutritional requirement experiments in amoebae and macrophages. J.M.P. and T.J.O. performed phenotypic analyses of Legionella species in vitro and in amoebae. J.M.P., S.G. and T.J.O. performed intracellular growth assays of Legionella species in macrophages. S.G. and T.J.O. performed Legionella species phenotype correlation analyses and add-back experiments in amoebae. S.G. wrote and executed custom scripts and phylogenetic cluster analyses. T.J.O., J.M.P. and S.G. wrote the manuscript. T.J.O. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
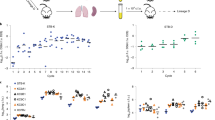
Extended Data Fig. 1 L. pneumophila growth in amoebal hosts and macrophages.
a, Intracellular growth cycle of L. pneumophila in A. castellanii, A. polyphaga, H. vermiformis and N. gruberi. Differences in the sets of bacterial genes required for growth in each amoeba were not due to variations in the duration of the intracellular growth cycle, or the cumulative replication, as no correlation between the number of genes identified and the extent of growth was observed and host-specific genes were identified for all hosts examined (Fig. 1). b, The intracellular growth cycle of L. pneumophila in U937 cells is similar to amoebae. In a-b, Host cells were challenged with wild type (WT) or Dot/Icm translocation deficient (dot-) bacteria constitutively expressing the enhanced green fluorescence protein and bacterial growth was monitored by measuring fluorescence over time. Plotted is the relative fluorescence units (RFU) normalized to the WT strain by the RFU at the 1 hour time point (for a, the WT strain in A. castellanii was used). Data are the mean ± standard deviation of 3 biological replicates, each generated from 3 technical replicates (Source Data for Extended Data Fig. 1).
Extended Data Fig. 2 Validation of IDTS mutant phenotypes predicated by Tn-seq.
a, Null mutations in IDTS genes do not impair L. pneumophila replication in nutrient rich bacteriological medium. The wild type (WT) and indicated null mutant strains were grown in CYE medium and the absorbance at 600 nm was monitored over time. Data are the mean ± standard deviation of 2 biological replicates, each generated from 2 technical replicates (Source Data 6). b, IDTS genes for which mutations were predicted to be neutral based on Tn-seq do not impair L. pneumophila growth in amoebae when compared to the WT strain. c, The phenotype predicted by Tn-seq for mavB (lgp1752) is due to polar effects of transposon insertion mutations on lpg1751. Top panel: The L. pneumophila genetic locus of lpg1751 and lpg1752 (mavB). Bottom panel: Deletion of lpg1751 but not mavB severely impairs L. pneumophila replication within A. castellanii and a ∆mavB∆lpg1751 double mutant phenocopies the ∆lpg1751 single deletion strain. d, The intracellular growth defects of IDTS null mutants can be rescued by reintroducing the corresponding gene on a self-replicating plasmid. Bacterial strains harboring the empty vector pJB908 or the respective complementation plasmid were used to challenge A. castellanii or H. vermiformis. In b,c,d, The WT and indicated IDTS null mutant strains were used to challenge amoebae and bacterial growth, based on recovered colony forming units (cfus) on solid media from lysed host cells, was assessed after 24 hours. Plotted is the total bacterial yield 24 hours post infection (hpi) normalized to the wild type strain by the number of intracellular bacteria 1–2 hpi. e, Uptake and/or survival phenotypes of IDTS mutants can be rescued by reintroducing the corresponding gene on a self-replicating plasmid. Bacterial strains harboring the empty vector pJB908 or the respective complementation plasmid were used to challenge A. castellanii or H. vermiformis for 1.5 hours. Cells were then treated with gentamycin to kill extracellular bacteria and internal bacteria were enumerated based on recovered cfus from host cell lysates plated on bacteriological medium. In b-e, Data are the mean ± standard deviation of 2–5 biological replicates, each generated from 3 technical replicates; *P < 0.02, 1-way ANOVA, relative to the WT strain (Source Data for Extended Data Fig. 2).
Extended Data Fig. 3 Universal IDTS are required for replication of L. pneumophila strains 130b and Paris in amoebae.
The wild type (WT) and indicated IDTS null mutant strains were used to challenge amoebae and bacterial growth, based on recovered colony forming units (cfus) on solid media from lysed host cells, was assessed after 24 hours. Plotted is the total bacterial yield 24 hours post infection (hpi) normalized to the WT strain by the number of intracellular bacteria 1–2 hpi. Data are the mean ± standard deviation of 2–3 biological replicates, each generated from 3 technical replicates, *P<0.02, 1-way ANOVA, relative to the WT strain (Source Data for Extended Data Fig. 3).
Extended Data Fig. 4 IDTS mutants exhibit similar phenotypes in primary bone marrow-derived murine macrophages.
a, Intracellular growth of the wild type (WT) and universal IDTS mutant strains in primary bone-marrow-derived macrophages from A/J mice. b, Intracellular replication of auxiliary IDTS mutants. c, Amoebae-specific IDTS genes are a source of redundancy in primary macrophages. In a-c, Macrophages were challenge with the indicated strains and bacterial replication, based on recovered colony forming units (cfus) on solid media from lysed host cells, was monitored 24 hpi and normalized to the WT strain by the number of intracellular bacteria 1 hpi. Data are the mean ± standard deviation of 2–5 biological replicates, each generated from 3 technical replicates; *P < 0.03, 1-way ANOVA, relative to the WT strain (Source Data for Extended Data Fig. 4).
Extended Data Fig. 5 The amoebae-specific growth defect of the Δbio mutant strain is not medium-dependent.
To rule out prior culturing of A. castellanii in PYG medium and H. vermiformis and N. gruberi in PYNFH medium as a source of the amoebae-specific requirement for the biotin biosynthetic genes, growth of the Δbio mutant in A. castellanii pre-cultured in PYNFH was compared to A. castellanii pre-cultured in PYG. Under both conditions, the Δbio mutant show a growth defect. While this does not rule out trace amounts of biotin in PYNFH being sufficient for L. pneumophila growth in H. vermiformis and N. gruberi, this result demonstrates that in the same medium, the intracellular levels of biotin in A. castellanii are sufficiently lower than in N. gruberi or H. vermiformis, defining distinct differences in the availability of biotin between these amoebal hosts. Data are the mean ± standard deviation of 2 biological replicates, each generated from 3 technical replicates; *P<0.01, 1-way ANOVA, relative to the WT strain (Source Data for Extended Data Fig. 5).
Extended Data Fig. 6 Growth of 12 Legionella species compared to L. pneumophila in nutrient rich bacteriological medium.
The intracellular growth phenotypes of the 12 Legionella species were not due to differences in growth kinetics as all species grew as well as L. pneumophila in bacteriological medium. L. pneumophila and the indicated species of Legionella were grown in CYE medium and the absorbance at 600 nm was monitored over time. Data are the mean ± standard deviation of 3 biological replicates, each generated from 3 technical replicates. The doubling time (dt) of each species during the exponential growth phase was determined as dt = t/n, where t = time and n = number of generations as describe in Methods. Data are the mean ± standard deviation of 3 biological replicates, each generated from 3 technical replicates; *P < 0.05, two-tailed Student t test relative to L. pneumophila (Source Data for Extended Data Fig. 6).
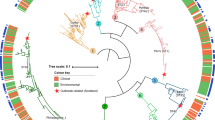
Extended Data Fig. 7 Correlates between the fitness profiles in amoebae and phylogenetic distances of individual Legionella species.
a, Pairwise comparisons define no correlation between the fitness profiles of 12 Legionella species (Fig. 5) and their phylogenetic relatedness. b, Pairwise comparisons between members of the same phenotypic group (Group I, II, III and IV as defined in Fig. 5) show strong positive correlations (blue) with the exception of L. dumoffii (Ld) and other Group I members (red). c, The fitness profile of L. dumoffii negatively correlates with that of all members of Group I, II and IV species (left panel) and is due to the decreased fitness of L. dumoffii in H. vermiformis relative to A. castellanii and N. gruberi (right panel). Plots are based on the phylogenetic distances and percent L. pneumophila intracellular growth reported in Fig. 5.
Extended Data Fig. 8 Different evolutionary trajectories of Legionella in the environment result in the convergent evolution of distinct but compensatory virulence mechanisms.
In the environment, a common ancestor of Legionella encounters distinct, possibly overlapping, sets of diverse amoebal hosts. Amoebal heterogeneity in different environmental niches provides the selective pressure driving the diversification of individual Legionella species through the assembly of both common and distinct sets of virulence genes. The resulting virulence gene repertoires collectively determined fitness in the environment and human macrophages. In some cases, the evolutionary trajectory of the bacterium results in a set of virulence factors that is not sufficient to support replication in macrophages (a). In other cases, different evolutionary trajectories lead to distinct sets of virulence factors that each promote replication in macrophages (b, c) and thus distinct but compensatory virulence mechanisms. Conversely, some bacteria accumulate virulence factors that while important for growth in amoebae restrict replication in macrophages (d).
Supplementary information
Supplementary Information
References for supplementary tables
Supplementary Information
Supplementary Tables 1–6.
Source data
Source Data Fig. 2
Source Data for Fig. 2.
Source Data Fig. 3
Source Data for Fig. 3.
Source Data Fig. 4
Source Data for Fig. 4.
Source Data Fig. 5
Source Data for Fig. 5.
Source Data Extended Data Fig. 1
Source Data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Source Data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Source Data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Source Data for Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Source Data for Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Source Data for Extended Data Fig. 6.
Rights and permissions
About this article
Cite this article
Park, J.M., Ghosh, S. & O’Connor, T.J. Combinatorial selection in amoebal hosts drives the evolution of the human pathogen Legionella pneumophila. Nat Microbiol 5, 599–609 (2020). https://doi.org/10.1038/s41564-019-0663-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0663-7
This article is cited by
-
Exploring the microbial savanna: predator-prey interactions in the soil
Molecular Systems Biology (2024)
-
Sodium levels and grazing pressure shape natural communities of the intracellular pathogen Legionella
Microbiome (2023)
-
Microbial warfare in the wild—the impact of protists on the evolution and virulence of bacterial pathogens
International Microbiology (2021)
-
Protozoa hosts lead to virulence
Nature Microbiology (2020)