Abstract
Supergenes are clusters of linked genetic loci that jointly affect the expression of complex phenotypes, such as social organization. Little is known about the origin and evolution of these intriguing genomic elements. Here we analyse whole-genome sequences of males from native populations of six fire ant species and show that variation in social organization is under the control of a novel supergene haplotype (termed Sb), which evolved by sequential incorporation of three inversions spanning half of a ‘social chromosome’. Two of the inversions interrupt protein-coding genes, resulting in the increased expression of one gene and modest truncation in the primary protein structure of another. All six socially polymorphic species studied harbour the same three inversions, with the single origin of the supergene in their common ancestor inferred by phylogenomic analyses to have occurred half a million years ago. The persistence of Sb along with the ancestral SB haplotype through multiple speciation events provides a striking example of a functionally important trans-species social polymorphism presumably maintained by balancing selection. We found that while recombination between the Sb and SB haplotypes is severely restricted in all species, a low level of gene flux between the haplotypes has occurred following the appearance of the inversions, potentially mitigating the evolutionary degeneration expected at genomic regions that cannot freely recombine. These results provide a detailed picture of the structural genomic innovations involved in the formation of a supergene controlling a complex social phenotype.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The genome assembly, gene models and sequence reads are available at the NCBI under the BioProject PRJNA421367.
References
Ross, K. G. Multilocus evolution in fire ants: effects of selection, gene flow and recombination. Genetics 145, 961–974 (1997).
Ross, K. G. & Keller, L. Genetic control of social organization in an ant. Proc. Natl Acad. Sci. USA 95, 14232–14237 (1998).
Krieger, M. J. B. & Ross, K. G. Identification of a major gene regulating complex social behavior. Science 295, 328–332 (2002).
Gotzek, D. & Ross, K. G. Genetic regulation of colony social organization in fire ants: an integrative overview. Q. Rev. Biol. 82, 201–226 (2007).
Wang, J. et al. A Y-like social chromosome causes alternative colony organization in fire ants. Nature 493, 664–668 (2013).
Ross, K. & Keller, L. Experimental conversion of colony social organization by manipulation of worker genotype composition in fire ants (Solenopsis invicta). Behav. Ecol. Sociobiol. 51, 287–295 (2002).
Charlesworth, D. & Charlesworth, B. Theoretical genetics of Batesian mimicry II. Evolution of supergenes. J. Theor. Biol. 55, 305–324 (1975).
Schwander, T., Libbrecht, R. & Keller, L. Supergenes and complex phenotypes. Curr. Biol. 24, R288–R294 (2014).
Thompson, M. J. & Jiggins, C. D. Supergenes and their role in evolution. Heredity 113, 1–8 (2014).
Zhang, W., Westerman, E., Nitzany, E., Palmer, S. & Kronforst, M. R. Tracing the origin and evolution of supergene mimicry in butterflies. Nat. Commun. 8, 1269 (2017).
Huang, Y.-C., Dang, V. D., Chang, N.-C. & Wang, J. Multiple large inversions and breakpoint rewiring of gene expression in the evolution of the fire ant social supergene. Proc. R. Soc. B 285, 20180221 (2018).
Ho, M. S., Tsai, P.-I. & Chien, C.-T. F-box proteins: the key to protein degradation. J. Biomed. Sci. 13, 181–191 (2006).
Long, M., Betrán, E., Thornton, K. & Wang, W. Chromosome rearrangement by ectopic recombination in Drosophila melanogaster: genome structure and evolution. Genetics 129, 1085–1098 (1991).
Huang, Y.-C. et al. Evolution of long centromeres in fire ants. BMC Evol. Biol. 16, 189 (2016).
Ross, K. G. & Shoemaker, D. Unexpected patterns of segregation distortion at a selfish supergene in the fire ant Solenopsis invicta. BMC Genet. 19, 101 (2018).
Jaenike, J. Sex chromosome meiotic drive. Annu. Rev. Ecol. Syst. 32, 25–49 (2001).
Fritz, G. N., Vander Meer, R. K. & Preston, C. A. Selective male mortality in the red imported fire ant, Solenopsis invicta. Genetics 173, 207–213 (2006).
Lawson, L. P., Vander Meer, R. K. & Shoemaker, D. Male reproductive fitness and queen polyandry are linked to variation in the supergene Gp-9 in the fire ant Solenopsis invicta. Proc. R. Soc. B 279, 3217–3222 (2012).
Gotzek, D., Shoemaker, D. & Ross, K. G. Molecular variation at a candidate gene implicated in the regulation of fire ant social behavior. PLoS ONE 2, e1088 (2007).
Krieger, M. J. B. & Ross, K. G. Molecular evolutionary analyses of the odorant-binding protein gene Gp-9 in fire ants and other Solenopsis species. Mol. Biol. Evol. 22, 2090–2103 (2005).
Charlesworth, D. The status of supergenes in the 21st century: recombination suppression in Batesian mimicry and sex chromosomes and other complex adaptations. Evol. Appl. 9, 74–90 (2016).
DeHeer, C. J., Goodisman, M. A. D. & Ross, K. G. Queen dispersal strategies in the multiple-queen form of the fire ant Solenopsis invicta. Am. Nat. 153, 660–675 (1999).
Hallar, B. L., Krieger, M. J. B. & Ross, K. G. Potential cause of lethality of an allele implicated in social evolution in fire ants. Genetica 131, 69–79 (2007).
Remis, M. I. Chromosome polymorphisms in natural populations of the South American grasshopper Sinipta dalmani. Genet. Mol. Biol. 31, 42–48 (2008).
Campos, J. L., Charlesworth, B. & Haddrill, P. R. Molecular evolution in nonrecombining regions of the Drosophila melanogaster genome. Genome Biol. Evol. 4, 278–288 (2012).
Stolle, E. et al. Degenerative expansion of a young supergene. Mol. Biol. Evol. 36, 553–561 (2018).
Gotzek, D., Clarke, J. & Shoemaker, D. Mitochondrial genome evolution in fire ants (Hymenoptera: Formicidae). BMC Evol. Biol. 10, 300 (2010).
Pracana, R., Priyam, A., Levantis, I., Nichols, R. A. & Wurm, Y. The fire ant social chromosome supergene variant Sb shows low diversity but high divergence from SB. Mol. Ecol. 26, 2864–2879 (2017).
Ross, K. G. & Shoemaker, D. Estimation of the number of founders of an invasive pest insect population: the fire ant Solenopsis invicta in the USA. Proc. R. Soc. B 275, 2231–2240 (2008).
Ross, K. G., Vargo, E. L. & Keller, L. Social evolution in a new environment: the case of introduced fire ants. Proc. Natl Acad. Sci. USA 93, 3021–3025 (1996).
Bachtrog, D. Y-chromosome evolution: emerging insights into processes of Y-chromosome degeneration. Nat. Rev. Genet. 14, 113–124 (2013).
Charlesworth, D., Charlesworth, B. & Marais, G. Steps in the evolution of heteromorphic sex chromosomes. Heredity 95, 118–128 (2005).
Huang, Y.-C. & Wang, J. Did the fire ant supergene evolve selfishly or socially? Bioessays 36, 200–208 (2014).
Comeron, J. M., Williford, A. & Kliman, R. M. The Hill–Robertson effect: evolutionary consequences of weak selection and linkage in finite populations. Heredity 100, 19–31 (2008).
Kamdem, C., Fouet, C. & White, B. J. Chromosome arm-specific patterns of polymorphism associated with chromosomal inversions in the major African malaria vector, Anopheles funestus. Mol. Ecol. 26, 5552–5566 (2017).
Jay, P. et al. Supergene evolution triggered by the introgression of a chromosomal inversion. Curr. Biol. 28, 1839–1845 (2018).
Llaurens, V., Whibley, A. & Joron, M. Genetic architecture and balancing selection: the life and death of differentiated variants. Mol. Ecol. 26, 2430–2448 (2017).
Tschinkel, W. R. The Fire Ants (Harvard Univ. Press, 2006).
Stevison, L. S., Hoehn, K. B. & Noor, M. A. F. Effects of inversions on within- and between-species recombination and divergence. Genome Biol. Evol. 3, 830–841 (2011).
Crown, K. N., Miller, D. E., Sekelsky, J. & Hawley, R. S. Local inversion heterozygosity alters recombination throughout the genome. Curr. Biol. 28, 2984–2990.e3 (2018).
Kelemen, R. K. & Vicoso, B. Complex history and differentiation patterns of the t-haplotype, a mouse meiotic driver. Genetics 208, 365–375 (2018).
Grossen, C., Neuenschwander, S. & Perrin, N. The evolution of XY recombination: sexually antagonistic selection versus deleterious mutation load. Evolution 66, 3155–3166 (2012).
Muller, H. J. The relation of recombination to mutational advance. Mutat. Res. Mol. Mech. Mutagen. 1, 2–9 (1964).
Manfredini, F. et al. Molecular and social regulation of worker division of labour in fire ants. Mol. Ecol. 23, 660–672 (2014).
Shoemaker, D. & Ascunce, M. S. A new method for distinguishing colony social forms of the fire ant, Solenopsis invicta. J. Insect Sci. 10, 73 (2010).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive κ-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Tang, H. et al. ALLMAPS: robust scaffold ordering based on multiple maps. Genome Biol. 16, 3 (2015).
Kiełbasa, S. M., Wan, R., Sato, K., Horton, P. & Frith, M. C. Adaptive seeds tame genomic sequence comparison. Genome Res. 21, 487–493 (2011).
Crooks, G. E., Hon, G., Chandonia, J.-M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Ometto, L., Shoemaker, D., Ross, K. G. & Keller, L. Evolution of gene expression in fire ants: the effects of developmental stage, caste, and species. Mol. Biol. Evol. 28, 1381–1392 (2011).
Valles, S. M. & Porter, S. D. Identification of polygyne and monogyne fire ant colonies (Solenopsis invicta) by multiplex PCR of Gp-9 alleles. Insectes Soc. 50, 199–200 (2003).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2009).
Van der Auwera, G. A. et al. From fastQ data to high-confidence variant calls: the genome analysis toolkit best practices pipeline. Curr. Protoc. Bioinf. 43, 11.10.1–11.10.33 (2013).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Chen, K. et al. TIGRA: a targeted iterative graph routing assembler for breakpoint assembly. Genome Res. 24, 310–317 (2014).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. Preprint at http://arXiv.org/abs/1207.3907 (2012).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Pitts, J. P., Camacho, G. P., Gotzek, D., Mchugh, J. V. & Ross, K. G. Revision of the fire ants of the Solenopsis saevissima species-group (Hymenoptera: Formicidae). Proc. Entomol. Soc. Wash. 120, 308–411 (2018).
Lee, T.-H., Guo, H., Wang, X., Kim, C. & Paterson, A. H. SNPhylo: a pipeline to construct a phylogenetic tree from huge SNP data. BMC Genom. 15, 162 (2014).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Reed, E. et al. A guide to genome-wide association analysis and post-analytic interrogation. Stat. Med. 34, 3769–3792 (2015).
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012).
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010).
Moreau, C. S. & Bell, C. D. Testing the museum versus cradle tropical biological diversity hypothesis: phylogeny, diversification, and ancestral biogeographic range evolution of the ants. Evolution 67, 2240–2257 (2013).
Drummond, A. J. & Rambaut, A. BEAST: Bayesian evolutionary analysis by somping trees. BMC Evol. Biol. 7, 214 (2007).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff. Fly 6, 80–92 (2012).
Yang, Z. PAML: a program package for phylogenetic analysis by maximum likelihood. Bioinformatics 13, 555–556 (1997).
Xu, B. & Yang, Z. PAMLX: a graphical user interface for PAML. Mol. Biol. Evol. 30, 2723–2724 (2013).
Acknowledgements
We thank K. Harshman for Illumina and Pacbio sequencing support and R. Arguello, H. Darras, T. Flatt, J. Goudet, M. Qiaowei Pan, T. Schwander and J. Wang for comments on earlier versions of the manuscript. All computations were performed at the Vital-IT (http://www.vital-it.ch) Center for High-Performance Computing of the Swiss Institute of Bioinformatics. This work was supported by grants from the Swiss NSF to L.K., an ERC advanced grant to L.K., US NSF grants to K.G.R. and D.S. (no. 1354479) and K.G.R. and B.G.H. (no. 1755130) and US Federal Hatch funds to K.G.R. and B.G.H.
Author information
Authors and Affiliations
Contributions
D.G., K.G.R., D.S. and L.K. designed the experiments. K.G.R., D.S. and D.G. performed sample collection, DNA extraction and genotyping. Z.Y. performed PacBio sequence data collection and genome assembly and analysed the population genomic data. Z.Y., P.D., N.S. and L.K. conducted phylogenomic analyses. S.V.A., Z.Y., O.R.-G. and B.G.H. performed RNA-seq analyses. Q.H. conducted analyses of the structure of the genes interrupted by the inversions. S.H.M. performed population genetic simulations. Z.Y., K.G.R. and L.K. wrote the manuscript with the help of all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended Data
Extended Data Fig. 1 Dot plot of the alignment between chromosomes 1-15 of Sb and SB males of S. invicta from the invasive (United States) range.
The red line indicates the forward strand alignment.
Extended Data Fig. 2 Sequence features for the breakpoints of the three Sb supergene inversions, In(16)1 [chr16:14,549,064-24,031,576], In(16)2 [chr16: 13,705,210-24,030,990], and In(16)3 [chr16: 12,612,565-13,683,100].
a, The red and blue blocks represent the 200nt segments adjacent to the In(16)1 proximal breakpoint (red arrow), and the magenta and gold blocks represent the 200nt segments adjacent to the distal breakpoint (black arrow) of this inversion. The blue and magenta blocks, and the segment between them, are inverted between SB and Sb, as indicated by the grey arrow. The Sb haplotype has a 9nt insertion at the proximal breakpoint and a 27,953nt insertion at the distal breakpoint (black blocks). Percentages are the sequence similarity (disregarding deletions) between SB and Sb for the 200nt segments immediately upstream and downstream of the breakpoints. b, The In(16)1 proximal breakpoint (red arrow) on the SB haplotype is located in exon 1 of the “F-box/WD repeat-containing protein 4-like” gene (NCBI Gene symbol: LOC105199310; green blocks depict exons; pale green represents the UTRs and dark green the coding sequence [CDS] regions). The red and blue lines under exon 1 indicate the segments that are upstream or downstream of the proximal breakpoint. c, The In(16)1 distal breakpoint (black arrow) on the SB haplotype is located in the 5′ UTR of the “Phosphoglycerate mutase 2” gene (NCBI Gene symbol: LOC105193833; green blocks depict exons; pale green represents the UTRs and dark green the CDS regions). The red and blue lines under exon 1 indicate the segments that are upstream or downstream of the distal breakpoint. d, The red and blue blocks represent the 200nt segments adjacent to the In(16)2 proximal breakpoint (red arrow), and the magenta and gold blocks the 200nt segments adjacent to the distal breakpoint (black arrow) of this inversion. The red block contains the 3′ end of a Jockey-like mobile element. The Sb haplotype has a 14nt insertion (black block) at the proximal breakpoint as well as a second Jockey-like mobile element gene (pink block) and a 10,310nt insertion (black block) just upstream of the distal breakpoint. e, The In(16)2 proximal breakpoint (red arrow) in the SB haplotype is located in the single exon (dark green) of a Jockey-like mobile element. f, The In(16)2 distal breakpoint (black arrow) in the SB haplotype is located in the 5′ UTR of the uncharacterized gene “LOC105193832” (containing 3 exons depicted as green blocks; pale green represents the UTR and dark green the CDS region). g, The red and blue blocks represent the 200nt segments adjacent to the In(16)3 proximal breakpoint (red arrow), and the magenta and gold blocks the 200nt segments adjacent to the distal breakpoint (black arrow) of this inversion. The SB haplotype has a 334nt insertion at the proximal breakpoint and a 3,100nt insertion at the distal breakpoint (black blocks); both are absent in the Sb haplotype, which instead has a 62,682nt insertion at the proximal breakpoint and a 142nt insertion at the distal breakpoint (black blocks). h, The In(16)3 proximal breakpoint (red arrow) in the SB haplotype is located just upstream of a region containing 24 dinucleotide (AT) repeats. i, The In(16)3 distal breakpoint (black arrow) in the SB haplotype is located within a region containing 21 dinucleotide (AT) repeats.
Extended Data Fig. 3 Functional features of the two protein-coding genes interrupted by the inversion In(16)1 breakpoints.
a, The S. invicta Phosphoglycerate mutase 2 (PGAM2) protein sequence shows a very high level of similarity with putative orthologs in Drosophila melanogaster (fruit fly), Danio rerio (zebrafish), Mus musculus (mouse), and Homo sapiens. Virtually all described functional sites are conserved across the five species: the catalytic core is strictly conserved and all but one amino acid of the substrate binding sites are also conserved. This high level of amino acid sequence conservation suggests conservation of the function of PGAM2 among putative orthologs. b, The S. invicta FBXW4 protein contains the typical domains of F-box protein ubiquitin ligase complexes (F-box and WD40 repeats from InterPro and NCBI predictions) and is therefore also likely to be involved in ubiquitination and proteasome degradation. In the alignment, identical sites are shown with black bars, 75% similar with dark grey, 50% with light grey, and 25% with white. c: Disruption of PGAM in D. melanogaster, the ortholog of PGAM2 in S. invicta, is not lethal. Paralogs of S. invicta FBXW4 in D. melanogaster have a wide range of substrates and their disruption can be lethal.
Extended Data Fig. 4 The box plots of the numbers of anomalous read pairs (ARPs) connecting the downstream and upstream regions (400nt) adjacent to the breakpoints of the three supergene inversions [In(16)1, In(16)2, In(16)3] for Sb and SB males of six socially polymorphic fire ant species.
Each box ranges from the first (Q1) to the third quartile (Q3) of the distribution and represents the interquartile range (IQR). A line across the box indicates the median. The whiskers are lines extending from Q1 and Q3 to end points that are defined as the most extreme data points within Q1 − 1.5 × IQR and Q3 + 1.5 × IQR, respectively. Each outlier outside the whiskers is represented by a solid dot. a-c, ARPs connecting proximal and distal inversion breakpoints when samples are mapped to the Sb reference genome. d-f, ARPs connecting proximal and distal inversion breakpoints when samples are mapped to the SB reference genome. There are four Gp-9b individuals with zero values when samples are mapped on the SB reference genome (one S. AdRX for inversion In(16)1); one S. richteri, one S. nr. interrupta, and one S. macdonaghi for inversion In(16)3)). However, the targeted local assembly of the breakpoint sequences yielded contigs that bridge these breakpoints in Sb males of all these four individuals.
Extended Data Fig. 5 Linkage disequilibrium (r2; = gametic disequilibrium for haploid males) in native S. invicta estimated using SNPs across the social chromosome (chr16).
a, LD dot plot for pooled SB (N = 60) and Sb (N = 19) males. b, LD plot for Sb males. c: LD plot for SB males. The coloured bar under each plot represents the physical map of the chromosome, with the red segment indicating the region where recombination is suppressed between the Sb and SB haplotypes in invasive (United States) S. invicta. SNPs are ordered according to physical position on the chromosome. The blue and red dashed lines link SNPs on the LD plot to their position on the physical map. The centromere occupies the approximate region 7.5-11 Mb.
Extended Data Fig. 6 Nucleotide divergence (dXY) values between sequences from conspecific Sb and SB males of the six socially polymorphic fire ant species estimated using 50 kb non-overlapping sliding windows across the social chromosome.
The boundaries of the differently shaded intervals correspond to the breakpoints of the three inversions, with the inversions depicted by thick horizontal coloured lines. The x axes represent physical position along chromosome 16, with the location of gene Gp-9 indicated. For each of the six species, dXY values between Sb and SB males were significantly higher for the region corresponding to the inversions than for the rest of the social chromosome (Mann–Whitney U-tests, all P < 0.01). The region of elevated dXY proximal to the inversions (at ca. 8-10 Mb) appears to experience reduced recombination in S. invicta based on progeny studies;15 this area may correspond at least partly to the centomeric region or some other, unknown feature may be responsible for the reduced recombination and elevated dXY there.
Extended Data Fig. 7 Linkage disequilibrium (r2; = gametic disequilibrium) for pooled conspecific Sb and SB males of five native populations of socially polymorphic fire ants estimated using SNPs across the social chromosome.
a, LD dot plot for pooled Sb (N = 30) and SB (N = 26) S. richteri males. b:, LD plot for Sb (N = 5) and SB (N = 11) S. AdRX males. c, LD plot for Sb (N = 2) and SB (N = 2) S. nr. interrupta males. d, LD plot for Sb (N = 1) and SB (N = 2) S. megergates males. e, LD plot for Sb (N = 1) and SB (N = 2) S. macdonaghi males. Mean LD estimates for exclusively Sb haplotypes are r2 = 0.41, 0.36, 0.85, 0.87, and 0.80 for S. richteri, S. AdRX, S. megergates, S. nr. interrupta, and S. macdonaghi, respectively. See Extended Data Fig. 5 caption for additional information.
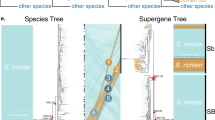
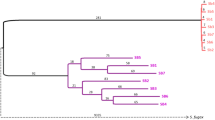
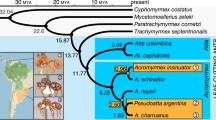
Extended Data Fig. 8 Phylogenetic trees for the supergene region of the social chromosome and for the fire ant species included in this study.
a, Alternative maximum-likelihood (ML) tree for the supergene region that disregards the LD (non-independence) of the 30,921 included SNPs. b, Alternative ML tree for the supergene region that accounts for LD, using a pruning threshold of 0.5. Trees in (a) and (b) were rooted with the outgroup species S. saevissima, which lacks the three chr16 inversions. The red scale bars are substitutions per site. c, Bayesian-inference tree with divergence time estimates for the study species based on sequences of five nuclear genes. The analysis incorporated the uncorrelated molecular clock method (BEAST), with the age of the basal divergence calibrated using data from a previous study (see Methods). The tree was rooted with the outgroup species S. xyloni. The number at each node represents the mean estimate of divergence time (in Myr), with the green bars representing the 95% confidence intervals (CIs) about the estimates; divergence time for the two major lineages of the socially polymorphic clade is highlighted with yellow background. Note that the topology of this tree is fully congruent with that of the ML species tree based on 12,237,341 non-supergene SNPs (Fig. 4).
Extended Data Fig. 9 Evidence from dXY and dS estimates that the three inversions comprising the Sb haplotype emerged in the order In(16)1 → In(16)2 → In(16)3.
a, Values of dXY between conspecific Sb and SB haplotypes for the three inversions in each of the six socially polymorphic fire ant species studied (left panel) and for groups of species for which data were pooled (right panel) (means and 95% CIs from 1000 bootstrap replicates). Values for subsets of species were pooled if they did not differ significantly (see Methods). Note that dXY values for S. macdonaghi, S. megergates, and S. nr. interrupta may not be highly accurate because of the small sample sizes for Sb in these species. b, Values of dS between conspecific Sb and SB haplotypes for the three inversions in each of the six socially polymorphic fire ant species studied (left panel) and for groups of species for which data were pooled (right panel) (means and 95% CIs from 1000 bootstrap replicates). Values for subsets of species were pooled if they did not differ significantly (see Methods). Note that dS values for S. macdonaghi, S. megergates, and S. nr. interrupta may be inflated because the small sample sizes of Sb for these species mean that some intrahaplotype polymorphic synonymous changes are interpreted to be fixed synonymous differences.
Supplementary information
Supplementary Table
Supplementary Tables 1–5.
Rights and permissions
About this article
Cite this article
Yan, Z., Martin, S.H., Gotzek, D. et al. Evolution of a supergene that regulates a trans-species social polymorphism. Nat Ecol Evol 4, 240–249 (2020). https://doi.org/10.1038/s41559-019-1081-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-1081-1
This article is cited by
-
Genomic structural variation is associated with hypoxia adaptation in high-altitude zokors
Nature Ecology & Evolution (2024)
-
Advances in genome sequencing reveal changes in gene content that contribute to arthropod macroevolution
Development Genes and Evolution (2023)
-
Functional properties of ant queen pheromones as revealed by behavioral experiments
Behavioral Ecology and Sociobiology (2023)
-
A novel distribution of supergene genotypes is present in the socially polymorphic ant Formica neoclara
BMC Ecology and Evolution (2022)
-
Recurring adaptive introgression of a supergene variant that determines social organization
Nature Communications (2022)