Abstract
Main conclusion
Circular RNA (circRNA) identification and expression profiles, and construction of circRNAs-miRNAs-mRNAs networks indicates that circRNAs are involved in wood formation of poplars in acclimation to low nitrogen availability.
Abstract
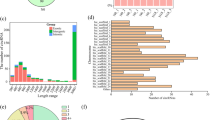
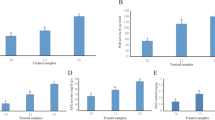
Circular RNAs (circRNAs) are covalently closed non-coding RNAs that play pivotal roles in various biological processes. However, circRNAs’ roles in wood formation of poplars in acclimation to low nitrogen (N) availability are currently unknown. Here, we undertook a systematic identification and characterization of circRNAs in the wood of Populus × canescens exposed to either 50 (low N) or 500 (normal N) µM NH4NO3 using rRNA-depleted RNA-sequencing. A total of 2,509 unique circRNAs were identified, and 163 (ca. 6.5%) circRNAs were significantly differentially expressed (DE) under low N condition. We observed a positive correlation between the expression patterns of DE circRNAs and their hosting protein-coding genes. Moreover, circRNAs–miRNAs–mRNAs’ networks were identified in the wood of poplars under low N availability. For instance, upregulated several circRNAs, such as circRNA1226, circRNA 1732, and circRNA392 induced increases in nuclear factor Y, subunit A1-A (NFYA1-A), NFYA1-B, and NFYA10 transcript levels via the mediation of miR169b members, which is in line with reduced xylem width and cell layers of the xylem in the wood of low N-supplied poplars. Upregulation of circRNA1006, circRNA1344, circRNA1941, circRNA901, and circRNA146 caused increased transcript level of MYB61 via the mediation of a miR5021 member, corresponding well to the higher lignin concentration in the wood of low N-treated poplars. Overall, these results indicated that DE circRNAs play an essential role in regulating gene expression via circRNAs–miRNAs–mRNAs’ networks to modulate wood anatomical and chemical properties of poplars in acclimation to low N availability.






Similar content being viewed by others
Abbreviations
- CSLG3 :
-
Cellulose synthase-like G3
- P5CS1 :
-
Delta1-pyrroline-5-carboxylate synthase 1
- DE:
-
Differentially expressed
- GO:
-
Gene Ontology
- MYB61 :
-
MYB domain protein 61
- NFYA1-A/1-B/10 :
-
Nuclear factor Y, subunit A1-A/1-B/10
References
Bao M, Bian H, Zha Y, Li F, Sun Y, Bai B, Chen Z, Wang J, Zhu M, Han N (2014) miR396a-mediated basic helix-loop-helix transcription factor bHLH74 repression acts as a regulator for root growth in Arabidopsis seedlings. Plant Cell Physiol 55(7):1343–1353
Camargo ELO, Nascimento LC, Soler M, Salazar MM, Lepikson-Neto J, Marques WL, Alves A, Teixeira PJPL, Mieczkowski P, Carazzolle MF (2014) Contrasting nitrogen fertilization treatments impact xylem gene expression and secondary cell wall lignification in Eucalyptus. BMC Plant Biol 14(1):256
Chen X, Mao XZ, Huang JJ, Ding Y, Wu JM, Dong S, Kong L, Gao G, Li CY, Wei LP (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39:316–322 (Web Server issue)
Chen B, Chen J, Du Q, Zhou D, Wang L, Xie J, Li Y, Zhang D (2018a) Genetic variants in microRNA biogenesis genes as novel indicators for secondary growth in Populus. New Phytol 219(4):1263–1282. https://doi.org/10.1111/nph.15262
Chen L, Zhang P, Fan Y, Lu Q, Li Q, Yan J, Muehlbauer GJ, Schnable PS, Dai M, Li L (2018b) Circular RNAs mediated by transposons are associated with transcriptomic and phenotypic variation in maize. New Phytol 217(3):1292–1306. https://doi.org/10.1111/nph.14901
Combier JP, Frugier F, de Billy F, Boualem A, El-Yahyaoui F, Moreau S, Vernie T, Ott T, Gamas P, Crespi M, Niebel A (2006) MtHAP2-1 is a key transcriptional regulator of symbiotic nodule development regulated by microRNA169 in Medicago truncatula. Genes Dev 20(22):3084–3088. https://doi.org/10.1101/gad.402806
Conesa A, Götz S, Garcíagómez JM, Terol J, Talón M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676
Debernardi JM, Mecchia MA, Vercruyssen L, Smaczniak C, Kaufmann K, Inze D, Rodriguez RE, Palatnik JF (2014) Post-transcriptional control of GRF transcription factors by microRNA miR396 and GIF co-activator affects leaf size and longevity. Plant J 79(3):413–426
Desrochers A (2006) NPK fertilization at planting of three hybrid poplar clones in the boreal region of Alberta. For Ecol Manage 232(1):216–225
Feng XJ, Li JR, Qi SL, Lin QF, Jin JB, Hua XJ (2016) Light affects salt stress-induced transcriptional memory of P5CS1 in Arabidopsis. Proc Natl Acad Sci USA 113(51):E8335
Frazee AC, Pertea G, Jaffe AE, Langmead B, Salzberg SL, Leek JT (2015) Ballgown bridges the gap between transcriptome assembly and expression analysis. Nat Biotechnol 33(3):243
Gao Y, Zhang J, Zhao F (2017) Circular RNA identification based on multiple seed matching. Brief Bioinf 5:803–810
Gao Z, Li J, Luo M, Li H, Chen Q, Wang L, Song S, Zhao L, Xu W, Zhang C, Wang S, Ma C (2019) Characterization and cloning of grape circular RNAs identified the cold resistance-related vv-circATS1. Plant Physiol 180:966–985
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, Kjems J (2013) Natural RNA circles function as efficient microRNA sponges. Nature 495(7441):384–388
Jeck WR, Sharpless NE (2014) Detecting and characterizing circular RNAs. Nat Biotechnol 32(5):453–461
Joshi CP, Thammannagowda S, Fujino T, Gou J-Q, Avci U, Haigler CH, McDonnell LM, Mansfield SD, Mengesha B, Carpita NC (2011) Perturbation of wood cellulose synthesis causes pleiotropic effects in transgenic aspen. Mol Plant 4(2):331–345
Kato S, Hayashi M, Kitagawa M, Kajiura H, Maeda M, Kimura Y, Igarashi K, Kasahara M, Ishimizu T (2018) Degradation pathway of plant complex-type N-glycans: identification and characterization of a key α1,3-fucosidase from glycoside hydrolase family 29. Biochem J 475(1):305–317
Kiełbasa SM, Blüthgen N, Fähling M, Mrowka R (2010) Targetfinder.org: a resource for systematic discovery of transcription factor target genes. Nucleic Acids Res 38:W233 (Web Server issue)
Kim D, Salzberg SL (2011) TopHat-Fusion: an algorithm for discovery of novel fusion transcripts. Genome Biol 12(8):R72
Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, Zhong G, Yu B, Hu W, Dai L (2015) Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol 22(3):256
Li X, Yang L, Chen LL (2018) The biogenesis, functions, and challenges of circular RNAs. Mol Cell 71(3):428–442. https://doi.org/10.1016/j.molcel.2018.06.034
Liu S, Wu L, Qi H, Xu M (2019) LncRNA/circRNA–miRNA–mRNA networks regulate the development of root and shoot meristems of Populus. Ind Crops Prod 133:333–347. https://doi.org/10.1016/j.indcrop.2019.03.048
Lu Y, Deng S, Li Z, Wu J, Liu Q, Liu W, Yu W-J, Zhang Y, Shi W, Zhou J, Li H, Polle A, Luo Z-B (2019) Competing endogenous RNA networks underlying anatomical and physiological characteristics of poplar wood in acclimation to low nitrogen availability. Plant Cell Physiol 60(11):2478–2495. https://doi.org/10.1093/pcp/pcz146
Luo J, Zhou J-J (2019) Growth performance, photosynthesis, and root characteristics are associated with nitrogen use efficiency in six poplar species. Environ Exp Bot 164:40–51. https://doi.org/10.1016/j.envexpbot.2019.04.013
Luo J, Li H, Liu T, Polle A, Peng C, Luo Z-B (2013a) Nitrogen metabolism of two contrasting poplar species during acclimation to limiting nitrogen availability. J Exp Bot 64(14):4207–4424
Luo J, Qin J, He F, Li H, Liu T, Polle A, Peng C, Luo Z-B (2013b) Net fluxes of ammonium and nitrate in association with H+ fluxes in fine roots of Populus popularis. Planta 237(4):919–931
Luo J, Zhou J, Li H, Shi W, Polle A, Lu M, Sun X, Luo Z-B (2015) Global poplar root and leaf transcriptomes reveal links between growth and stress responses under nitrogen starvation and excess. Tree Physiol 35(12):1283–1302
Luo J, Liang Z, Wu M, Mei L (2019a) Genome-wide identification of BOR genes in poplar and their roles in response to various environmental stimuli. Environ Exp Bot 164:101–113. https://doi.org/10.1016/j.envexpbot.2019.04.006
Luo J, Zhou J-J, Masclaux-Daubresse C, Wang N, Wang H, Zheng B (2019b) Morphological and physiological responses to contrasting nitrogen regimes in Populus cathayana is linked to resources allocation and carbon/nitrogen partition. Environ Exp Bot 162:247–255. https://doi.org/10.1016/j.envexpbot.2019.03.003
Memczak S, Jens M, Elefsinioti A et al (2013) Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495(7441):333–338. https://doi.org/10.1038/nature11928
Meng Y, Shao C, Wang H, Jin Y (2012) Target mimics: an embedded layer of microRNA-involved gene regulatory networks in plants. BMC Genomics 13(1):197
Metge F, Czaja-Hasse LF, Reinhardt R, Dieterich C (2017) FUCHS-towards full circular RNA characterization using RNAseq. PeerJ 5:e2934. https://doi.org/10.7717/peerj.2934
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8(19):4321–4325
Pan T, Sun X, Liu Y, Li H, Deng G, Lin H, Wang S (2018) Heat stress alters genome-wide profiles of circular RNAs in Arabidopsis. Plant Mol Biol 96(3):217–229. https://doi.org/10.1007/s11103-017-0684-7
Paris S, Wessel PM, Dumas R (2002) Overproduction, purification, and characterization of recombinant aspartate semialdehyde dehydrogenase from Arabidopsis thaliana. Protein Expres Purif 24(1):99–104
Piwecka M, Glažar P, Hernandezmiranda LR, Memczak S, Wolf SA, Rybakwolf A, Filipchyk A, Klironomos F, Cerda Jara CA, Fenske P (2017) Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Science 357(6357):eaam8526
Quan M, Du Q, Xiao L, Lu W, Wang L, Xie J, Song Y, Xu B, Zhang D (2018) Genetic architecture underlying the lignin biosynthesis pathway involves noncoding RNAs and transcription factors for growth and wood properties in Populus. Plant Biotechnol J 17(1):302–315. https://doi.org/10.1111/pbi.12978
Ratke C, Terebieniec BK, Winestrand S, Derba-Maceluch M, Grahn T, Schiffthaler B, Ulvcrona T, Ozparpucu M, Ruggeberg M, Lundqvist SO, Street NR, Jonsson LJ, Mellerowicz EJ (2018) Downregulating aspen xylan biosynthetic GT43 genes in developing wood stimulates growth via reprograming of the transcriptome. New Phytol 219(1):230–245. https://doi.org/10.1111/nph.15160
Ren Y, Yue H, Li L, Xu Y, Wang Z, Xin Z, Lin T (2018) Identification and characterization of circRNAs involved in the regulation of low nitrogen-promoted root growth in hexaploid wheat. Biol Res 51(1):43. https://doi.org/10.1186/s40659-018-0194-3
Roberts AW, Bushoven JT (2007) The cellulose synthase (CESA) gene superfamily of the moss Physcomitrella patens. Plant Mol Biol 63(2):207–219
Rubinelli PM, Chuck G, Li X, Meilan R, Mugnozza GS, Massacci A, Tognetti R (2013) Constitutive expression of the Corngrass1 microRNA in poplar affects plant architecture and stem lignin content and composition. Biomass Bioenergy 54(4):312–321
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP (2011) A ceRNA hypothesis: the rosetta stone of a hidden RNA language? Cell 146(3):353–358. https://doi.org/10.1016/j.cell.2011.07.014
Sorin C, Declerck M, Christ A, Blein T, Ma L, Lelandais-Briere C, Njo MF, Beeckman T, Crespi M, Hartmann C (2014) A miR169 isoform regulates specific NF-YA targets and root architecture in Arabidopsis. New Phytol 202(4):1197–1211. https://doi.org/10.1111/nph.12735
Sun X, Wang C, Xiang N, Li X, Yang S, Du J, Yang Y, Yang Y (2017) Activation of secondary cell wall biosynthesis by miR319-targeted TCP4 transcription factor. Plant Biotechnol J 15(10):1284–1294. https://doi.org/10.1111/pbi.12715
Tong W, Yu J, Hou Y, Li F, Zhou Q, Wei C, Bennetzen JL (2018) Circular RNA architecture and differentiation during leaf bud to young leaf development in tea (Camellia sinensis). Planta 248(6):1417–1429. https://doi.org/10.1007/s00425-018-2983-x
Trapnell C, Pachter L, Salzberg SL (2009) TopHat: discovering splice junctions with RNA-seq. Bioinformatics 25(9):1105–1111. https://doi.org/10.1093/bioinformatics/btp120
Wang T, Zhao M, Zhang X, Liu M, Yang C, Chen Y, Chen R, Wen J, Mysore KS, Zhang WH (2017) Novel phosphate deficiency-responsive long non-coding RNAs in the legume model plant Medicago truncatula. J Exp Bot 68:5937–5948. https://doi.org/10.1093/jxb/erx384
Wilkins O, Nahal H, Foong J, Provart NJ, Campbell MM (2009) Expansion and diversification of the Populus R2R3-MYB family of transcription factors. Plant Physiol 149(2):981–993
Ye ZH, Zhong R (2015) Molecular control of wood formation in trees. J Exp Bot 66(14):4119–4131. https://doi.org/10.1093/jxb/erv081
Ye CY, Chen L, Liu C, Zhu QH, Fan L (2015) Widespread noncoding circular RNAs in plants. New Phytol 208(1):88–95. https://doi.org/10.1111/nph.13585
Yuan G, Wang J, Yi Z, Zhang J, Shuai C, Zhao F (2016) Comprehensive identification of internal structure and alternative splicing events in circular RNAs. Nat Commun 7:12060
Zhang J, Nieminen K, Serra JA, Helariutta Y (2014) The formation of wood and its control. Curr Opin Plant Biol 17:56–63. https://doi.org/10.1016/j.pbi.2013.11.003
Zhang G, Diao S, Zhang T, Chen D, He C, Zhang J (2019) Identification and characterization of circular RNAs during the sea buckthorn fruit development. RNA Biol 16(3):354–361. https://doi.org/10.1080/15476286.2019.1574162
Zheng Q, Bao C, Guo W, Li S, Chen J, Chen B, Luo Y, Lyu D, Li Y, Shi G (2016) Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat Commun 7:11215
Acknowledgements
This study was jointly supported by the Key Forestry Public Welfare Project (201504105), the Fundamental Research Funds for the Central Non-profit Research Institution of CAF (No. CAFYBB2016QB005), the Jiangsu Agriculture Science and Technology Innovation Fund (JASTIF) (No. CX(16)1005), and the National Natural Science Foundation of China (No. 31500507).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, H., Yu, W., Wu, J. et al. Identification and characterization of circular RNAs during wood formation of poplars in acclimation to low nitrogen availability. Planta 251, 47 (2020). https://doi.org/10.1007/s00425-020-03338-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-020-03338-w