Abstract
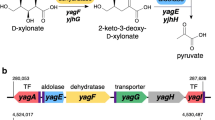
The xylose oxidative pathway (XOP) is continuously gaining prominence as an alternative for the traditional pentose assimilative pathways in prokaryotes. It begins with the oxidation of D-xylose to D-xylonic acid, which is further converted to α-ketoglutarate or pyruvate + glycolaldehyde through a series of enzyme reactions. The persistent drawback of XOP is the accumulation of D-xylonic acid intermediate that causes culture media acidification. This study addresses this issue through the development of a novel pH-responsive synthetic genetic controller that uses a modified transmembrane transcription factor called CadCΔ. This genetic circuit was tested for its ability to detect extracellular pH and to control the buildup of D-xylonic acid in the culture media. Results showed that the pH-responsive genetic sensor confers dynamic regulation of D-xylonic acid accumulation, which adjusts with the perturbation of culture media pH. This is the first report demonstrating the use of a pH-responsive transmembrane transcription factor as a transducer in a synthetic genetic circuit that was designed for XOP. This may serve as a benchmark for the development of other genetic controllers for similar pathways that involve acidic intermediates.







Similar content being viewed by others
Change history
16 January 2020
In the published version, the y-axis data of Fig. 3c was incorrectly inserted (OD600 instead of D-xylonate (g L<Superscript>-1</Superscript>) and the x-axes of Figs. 3b, 3d, 3e and 3f ended at 48 h instead of 72 h. See the correct Fig. 3 below.
References
Baldoma L, Aguilar J (1987) Involvement of lactaldehyde dehydrogenase in several metabolic pathways of Escherichia coli K12. J Biol Chem 262:13991–13996
Bañares AB, Valdehuesa KNG, Ramos KRM, Nisola GM, Lee W-K, Chung W-J (2019) Discovering a novel D-xylonate-responsive promoter: the PyjhI-driven genetic switch towards better 1,2,4-butanetriol production. Appl Microbiol Biotechnol 103:8063–8074. https://doi.org/10.1007/s00253-019-10073-0
Berghäll S, Hilditch S, Penttilä M, Richard P (2007) Identification in the mould Hypocrea jecorina of a gene encoding an NADP+: D-xylose dehydrogenase. FEMS Microbiol Lett 277:249–253. https://doi.org/10.1111/j.1574-6968.2007.00969.x
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Cabulong RB, Valdehuesa KNG, Ramos KRM, Nisola GM, Lee W-K, Lee CR, Chung W-J (2017) Enhanced yield of ethylene glycol production from D-xylose by pathway optimization in Escherichia coli. Enzym Microb Technol 97:11–20. https://doi.org/10.1016/j.enzmictec.2016.10.020
Cabulong RB, Lee W-K, Bañares AB, Ramos KRM, Nisola GM, Valdehuesa KNG, Chung W-J (2018a) Engineering Escherichia coli for glycolic acid production from D-xylose through the Dahms pathway and glyoxylate bypass. Appl Microbiol Biotechnol 102:2179–2189. https://doi.org/10.1007/s00253-018-8744-8
Cabulong RB, Valdehuesa KNG, Bañares AB, Ramos KRM, Nisola GM, Lee W-K, Chung W-J (2018b) Improved cell growth and biosynthesis of glycolic acid by overexpression of membrane-bound pyridine nucleotide transhydrogenase. J Ind Microbiol Biotechnol 46:159–169. https://doi.org/10.1007/s10295-018-2117-2
Choi SY, Park SJ, Kim WJ, Yang JE, Lee H, Shin J, Lee SY (2016) One-step fermentative production of poly(lactate-co-glycolate) from carbohydrates in Escherichia coli. Nat Biotechnol 34:435–440. https://doi.org/10.1038/nbt.3485
Chung CT, Niemela SL, Miller RH (1989) One-step preparation of competent Escherichia coli: transformation and storage of bacterial cells in the same solution. Proc Natl Acad Sci U S A 86:2172–2175
Dahms AS (1974) 3-deoxy-D-pentulosonic acid aldolase and its role in a new pathway of D-xylose degradation. Biochem Biophys Res Commun 60:1433–1439. https://doi.org/10.1016/0006-291X(74)90358-1
Deacon J, Cooper RA (2001) D-galactonate utilisation by enteric bacteria. FEBS Lett 77:201–205. https://doi.org/10.1016/0014-5793(77)80234-2
Dell CL, Neely MN, Olson ER (1994) Altered pH lysine signalling mutants of cadC, a gene encoding a membrane-bound transcriptional activator of the Escherichia coli cadBA operon. Mol Microbiol 14:7–16. https://doi.org/10.1111/j.1365-2958.1994.tb01262.x
Eichinger A, Haneburger I, Koller C, Jung K, Skerra A (2011) Crystal structure of the sensory domain of Escherichia coli CadC, a member of the ToxR-like protein family: crystal structure of CadC. Protein Sci 20:656–669. https://doi.org/10.1002/pro.594
Fraenkel DG, Levisohn SR (1967) Glucose and gluconate metabolism in an Escherichia coli mutant lacking phosphoglucose isomerase. J Bacteriol 93:1571–1578
Fritz G, Koller C, Burdack K, Tetsch L, Haneburger I, Jung K, Gerland U (2009) Induction kinetics of a conditional pH stress response system in Escherichia coli. J Mol Biol 393:272–286. https://doi.org/10.1016/j.jmb.2009.08.037
Gibson DG, Young L, Chuang R-Y, Venter JC, Hutchison CA, Smith HO (2009) Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6:343–345. https://doi.org/10.1038/nmeth.1318
Gírio FM, Fonseca C, Carvalheiro F, Duarte LC, Marques S, Bogel-Łukasik R (2010) Hemicelluloses for fuel ethanol: a review. Bioresour Technol 101:4775–4800. https://doi.org/10.1016/j.biortech.2010.01.088
Green MR, Sambrook J (2012) Molecular cloning: a laboratory manual, 4 edn. Cold Spring Harbor Laboratory Press, New York
Haas BL, Matson JS, Dirita VJ, Biteen JS (2015) Single-molecule tracking in live Vibrio cholerae reveals that ToxR recruits the membrane-bound virulence regulator TcpP to the toxT promoter. Mol Microbiol 96:4–13. https://doi.org/10.1111/mmi.12834
Haneburger I, Eichinger A, Skerra A, Jung K (2011) New insights into the signaling mechanism of the pH-responsive, membrane-integrated transcriptional activator CadC of Escherichia coli. J Biol Chem 286:10 681–10 689. https://doi.org/10.1074/jbc.M110.196923
Haneburger I, Fritz G, Jurkschat N, Tetsch L, Eichinger A, Skerra A, Gerland U, Jung K (2012) Deactivation of the E. coli pH stress sensor CadC by cadaverine. J Mol Biol 424:15–27. https://doi.org/10.1016/j.jmb.2012.08.023
Hoynes-OConnor A, Shopera T, Hinman K, Creamer JP, Moon TS (2017) Enabling complex genetic circuits to respond to extrinsic environmental signals. Biotechnol Bioeng 114:1626–1631. https://doi.org/10.1002/bit.26279
Jung K, Fabiani F, Hoyer E, Lassak J (2018) Bacterial transmembrane signalling systems and their engineering for biosensing. Open Biol 8:180023. https://doi.org/10.1098/rsob.180023
Lee C, Park C (2017) Bacterial responses to glyoxal and methylglyoxal: reactive electrophilic species. Int J Mol Sci 18:169. https://doi.org/10.3390/ijms18010169
Lindner E, White SH (2014) Topology, dimerization and stability of the single-span membrane protein CadC. J Mol Biol 426:2942–2957. https://doi.org/10.1016/j.jmb.2014.06.006
Liu H, Lu T (2015) Autonomous production of 1,4-butanediol via a de novo biosynthesis pathway in engineered Escherichia coli. Metab Eng 29:135–141. https://doi.org/10.1016/j.ymben.2015.03.009
Liu H, Valdehuesa KNG, Nisola GM, Ramos KRM, Chung W-J (2012) High yield production of D-xylonic acid from D-xylose using engineered Escherichia coli. Bioresour Technol 115:244–248. https://doi.org/10.1016/j.biortech.2011.08.065
Liu H, Ramos KRM, Valdehuesa KNG, Nisola GM, Lee W-K, Chung W-J (2013) Biosynthesis of ethylene glycol in Escherichia coli. Appl Microbiol Biotechnol 97:3409–3417. https://doi.org/10.1007/s00253-012-4618-7
Meijnen J-P, de Winde JH, Ruijssenaars HJ (2009) Establishment of oxidative D-xylose metabolism in Pseudomonas putida S12. Appl Environ Microbiol 75:2784–2791. https://doi.org/10.1128/AEM.02713-08
Nechooshtan G, Elgrably-Weiss M, Sheaffer A, Westhof E, Altuvia S (2009) A pH-responsive riboregulator. Genes Dev 23:2650–2662. https://doi.org/10.1101/gad.552209
Neely MN, Olson ER (1996) Kinetics of expression of the Escherichia coli cad operon as a function of pH and lysine. J Bacteriol 178:5522–5528
Neely MN, Dell CL, Olson ER (1994) Roles of LysP and CadC in mediating the lysine requirement for acid induction of the Escherichia coli cad operon. J Bacteriol 176:3278–3285. https://doi.org/10.1128/jb.176.11.3278-3285.1994
Pédelacq J-D, Cabantous S, Tran T, Terwilliger TC, Waldo GS (2006) Engineering and characterization of a superfolder green fluorescent protein. Nat Biotechnol 24:79–88. https://doi.org/10.1038/nbt1172
Pham HL, Wong A, Chua N, Teo WS, Yew WS, Chang MW (2017) Engineering a riboswitch-based genetic platform for the self-directed evolution of acid-tolerant phenotypes. Nat Commun 8:411. https://doi.org/10.1038/s41467-017-00511-w
Radek A, Krumbach K, Gätgens J, Wendisch VF, Wiechert W, Bott M, Noack S, Marienhagen J (2014) Engineering of Corynebacterium glutamicum for minimized carbon loss during utilization of D-xylose containing substrates. J Biotechnol 192(Pt a):156–160. https://doi.org/10.1016/j.jbiotec.2014.09.026
Rajkumar AS, Liu G, Bergenholm D, Arsovska D, Kristensen M, Nielsen J, Jensen MK, Keasling JD (2016) Engineering of synthetic, stress-responsive yeast promoters. Nucleic Acids Res 44:e136. https://doi.org/10.1093/nar/gkw553
Roberts TM, Rudolf F, Meyer A, Pellaux R, Whitehead E, Panke S, Held M (2016) Identification and characterisation of a pH-stable GFP. Sci Rep 6:28166. https://doi.org/10.1038/srep28166
Salusjärvi L, Toivari M, Vehkomäki M-L, Koivistoinen O, Mojzita D, Niemelä K, Penttilä M, Ruohonen L (2017) Production of ethylene glycol or glycolic acid from D-xylose in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 14:127–113. https://doi.org/10.1007/s00253-017-8547-3
Šeputienė V, Sužiedėlis K, Normark S, Melefors Ö, Sužiedėlienė E (2004) Transcriptional analysis of the acid-inducible asr gene in enterobacteria. Res Microbiol 155:535–542. https://doi.org/10.1016/j.resmic.2004.03.010
Slonczewski JL, Gonzalez TN, Bartholomew FM, Holt NJ (1987) Mu d-directed lacZ fusions regulated by low pH in Escherichia coli. J Bacteriol 169:3001–3006. https://doi.org/10.1128/jb.169.7.3001-3006.1987
Stirling F, Naydich A, Bramante J, Barocio R, Certo M, Wellington H, Redfield E, O’Keefe S, Gao S, Cusolito A, Way J, Silver P (2019) Synthetic cassettes for pH-mediated sensing, counting and containment. bioRxiv 740902. doi: https://doi.org/10.1101/740902
Sun L, Yang F, Sun H, Zhu T, Li X, Li Y, Xu Z, Zhang Y (2016) Synthetic pathway optimization for improved 1,2,4-butanetriol production. J Ind Microbiol Biotechnol 43:67–78. https://doi.org/10.1007/s10295-015-1693-7
Takayama M, Ohyama T, Igarashi K, Kobayashi H (1994) Escherichia coli cad operon functions as a supplier of carbon dioxide. Mol Microbiol 11:913–918. https://doi.org/10.1111/j.1365-2958.1994.tb00370.x
Tetsch L, Koller C, Haneburger I, Jung K (2008) The membrane-integrated transcriptional activator CadC of Escherichia coli senses lysine indirectly via the interaction with the lysine permease LysP. Mol Microbiol 67:570–583. https://doi.org/10.1111/j.1365-2958.2007.06070.x
Valdehuesa KNG, Liu H, Ramos KRM, Park SJ, Nisola GM, Lee W-K, Chung W-J (2014) Direct bioconversion of D-xylose to 1,2,4-butanetriol in an engineered Escherichia coli. Process Biochem 49:25–32. https://doi.org/10.1016/j.procbio.2013.10.002
Valdehuesa KNG, Lee W-K, Ramos KRM, Cabulong RB, Choi J, Liu H, Nisola GM, Chung W-J (2015) Identification of aldehyde reductase catalyzing the terminal step for conversion of xylose to butanetriol in engineered Escherichia coli. Bioprocess Biosyst Eng 38:1761–1772. https://doi.org/10.1007/s00449-015-1417-4
Valdehuesa KNG, Ramos KRM, Nisola GM, Bañares AB, Cabulong RB, Lee W-K, Liu H, Chung W-J (2018) Everyone loves an underdog: metabolic engineering of the xylose oxidative pathway in recombinant microorganisms. Appl Microbiol Biotechnol 102:7703–7716. https://doi.org/10.1007/s00253-018-9186-z
Wang C, Wei D, Zhang Z, Wang D, Shi J, Kim CH, Jiang B, Han Z, Hao J (2016) Production of xylonic acid by Klebsiella pneumoniae. Appl Microbiol Biotechnol 100:10055–10063. https://doi.org/10.1007/s00253-016-7825-9
Wang X, Xu N, Hu S, Yang J, Gao Q, Xu S, Chen K, Ouyang P (2018) D-1,2,4-Butanetriol production from renewable biomass with optimization of synthetic pathway in engineered Escherichia coli. Bioresour Technol 250:406–412. https://doi.org/10.1016/j.biortech.2017.11.062
Watson N, Dunyak DS, Rosey EL, Slonczewski JL, Olson ER (1992) Identification of elements involved in transcriptional regulation of the Escherichia coli cad operon by external pH. J Bacteriol 174:530–540
Weimberg R (1961) Pentose oxidation by Pseudomonas fragi. J Biol Chem 236:629–635
Wong TY, Yao XT (1994) The DeLey-Doudoroff pathway of galactose metabolism in Azotobacter vinelandii. Appl Environ Microbiol 60:2065–2068
Xu P (2018) Production of chemicals using dynamic control of metabolic fluxes. Curr Opin Biotechnol 53:12–19. https://doi.org/10.1016/j.copbio.2017.10.009
Xu P, Vansiri A, Bhan N, Koffas MAG (2012) ePathBrick: a synthetic biology platform for engineering metabolic pathways in E. coli. ACS Synth Biol 1:256–266. https://doi.org/10.1021/sb300016b
Funding
This work was supported by Korea Research Fellowship Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science and ICT (2015H1D3A1062172), by the Ministry of Education (2018R1D1A1B07043993), and by the Korea Institute of Energy Technology Evaluation and Planning (KETEP) funded by the Ministry of Trade, Industry & Energy (MOTIE, No. 20194010201750).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict(s) of interest.
Statement of informed consent
This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 617 kb)
Rights and permissions
About this article
Cite this article
Bañares, A.B., Valdehuesa, K.N.G., Ramos, K.R.M. et al. A pH-responsive genetic sensor for the dynamic regulation of D-xylonic acid accumulation in Escherichia coli. Appl Microbiol Biotechnol 104, 2097–2108 (2020). https://doi.org/10.1007/s00253-019-10297-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-10297-0