Abstract
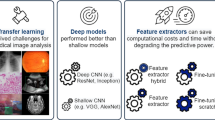
The tumor–stroma ratio (TSR) reflected on hematoxylin and eosin (H&E)-stained histological images is a potential prognostic factor for survival. Automatic image processing techniques that allow for high-throughput and precise discrimination of tumor epithelium and stroma are required to elevate the prognostic significance of the TSR. As a variant of deep learning techniques, transfer learning leverages nature-images features learned by deep convolutional neural networks (CNNs) to relieve the requirement of deep CNNs for immense sample size when handling biomedical classification problems. Herein we studied different transfer learning strategies for accurately distinguishing epithelial and stromal regions of H&E-stained histological images acquired from either breast or ovarian cancer tissue. We compared the performance of important deep CNNs as either a feature extractor or as an architecture for fine-tuning with target images. Moreover, we addressed the current contradictory issue about whether the higher-level features would generalize worse than lower-level ones because they are more specific to the source-image domain. Under our experimental setting, the transfer learning approach achieved an accuracy of 90.2 (vs. 91.1 for fine tuning) with GoogLeNet, suggesting the feasibility of using it in assisting pathology-based binary classification problems. Our results also show that the superiority of the lower-level or the higher-level features over the other ones was determined by the architecture of deep CNNs.





Similar content being viewed by others
References
Adeli, E., G. Wu, B. Saghafi, L. An, and D. Shen. Kernel-based joint feature selection and max-margin classification for early diagnosis of Parkinson’s disease. Sci. Rep. 7:41069, 2017.
Antony, J., K. McGuinness, N. E. O. Connor, and K. Moran. Quantifying radiographic knee osteoarthritis severity using deep convolutional neural networks. In: ICPR 2016 Proceedings, 2016, pp. 1195–1200. http://arxiv.org/abs/1609.02469.
Beck, A. H., A. R. Sangoi, S. Leung, R. J. Marinelli, T. O. Nielsen, M. J. van de Vijver, R. B. West, M. van de Rijn, and D. Koller. Systematic analysis of breast cancer morphology uncovers stromal features associated with survival. Sci. Transl. Med. 3:108ra113, 2011.
Bengio, Y., A. Courville, and P. Vincent. Representation learning: a review and new perspectives. IEEE Trans. Pattern Anal. Mach. Intell. 35:1798–1828, 2013.
Carneiro, G., J. Nascimento, and A. P. Bradley. Unregistered multiview mammogram analysis with pre-trained deep learning models. In: Lect. Notes Comput. Sci. (including Subser. Lect. Notes Artif. Intell. Lect. Notes Bioinformatics), vol. 9351, pp. 652–660, 2015.
Cireşan, D. C., A. Giusti, L. M. Gambardella, and J. Schmidhuber. Mitosis detection in breast cancer histology images with deep neural networks. Med. Image Comput. Comput. Interv. 16(Pt2):411–418, 2013.
de Kruijf, E. M., J. G. H. van Nes, C. J. H. van de Velde, H. Putter, V. T. H. B. M. Smit, G. J. Liefers, P. J. K. Kuppen, R. A. E. M. Tollenaar, and W. E. Mesker. Tumor–stroma ratio in the primary tumor is a prognostic factor in early breast cancer patients, especially in triple-negative carcinoma patients. Breast Cancer Res. Treat. 125:687–696, 2011.
Esteva, A., B. Kuprel, R. A. Novoa, J. Ko, S. M. Swetter, H. M. Blau, and S. Thrun. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542:115–118, 2017.
Girshick, R., J. Donahue, T. Darrell, and J. Malik. Region-based convolutional networks for accurate object detection and segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 38:142–158, 2016.
Gurcan, M. N., L. E. Boucheron, A. Can, A. Madabhushi, N. M. Rajpoot, and B. Yener. Histopathological image analysis: a review. IEEE Rev. Biomed. Eng. 2:147–171, 2009.
Huynh, B. Q., H. Li, and M. L. Giger. Digital mammographic tumor classification using transfer learning from deep convolutional neural networks. J. Med. Imaging 3:34501, 2016.
Ioffe, S., and C. Szegedy. Batch normalization: accelerating deep network training by reducing internal covariate shift. Arxiv 1–11, 2015. https://doi.org/10.1007/s13398-014-0173-7.2.
Krizhevsky, A., I. Sutskever, and G. E. Hinton. ImageNet classification with deep convolutional neural networks. Adv. Neural Inf. Process. Syst. 2012. https://doi.org/10.1016/j.protcy.2014.09.007.
LeCun, Y., Y. Bengio, and G. Hinton. Deep learning. Nature 521:436–444, 2015.
Liu, J., J. Liu, J. Li, Y. Chen, X. Guan, X. Wu, C. Hao, Y. Sun, Y. Wang, and X. Wang. Tumor–stroma ratio is an independent predictor for survival in early cervical carcinoma. Gynecol. Oncol. 132:81–86, 2014.
Long, X., L. Chen, C. Jiang, and L. Zhang. Prediction and classification of Alzheimer disease based on quantification of MRI deformation. PLoS ONE 12:1–19, 2017.
Lu, P., L. Barazzetti, V. Chandran, K. Gavaghan, S. Weber, N. Gerber, and M. Reyes. Highly accurate facial nerve segmentation refinement from CBCT/CT Imaging using a super resolution classification approach. IEEE Trans. Biomed. Eng. 65:178–188, 2017.
Mehrtash, A., A. Sedghi, M. Ghafoorian, M. Taghipour, C. M. Tempany, W. M. Wells, T. Kapur, P. Mousavi, P. Abolmaesumi, and A. Fedorov. Classification of clinical significance of MRI prostate findings using 3D convolutional neural networks. In: Proc SPIE Int Soc Opt Eng, 2017. https://doi.org/10.1117/12.2277123.
Menegola, A., M. Fornaciali, R. Pires, S. Avila, and E. Valle. Towards automated melanoma screening: exploring transfer learning schemes, 2016. http://arxiv.org/abs/1609.01228.
Moorman, A. M., R. Vink, H. J. Heijmans, J. van der Palen, and E. A. Kouwenhoven. The prognostic value of tumour-stroma ratio in triple-negative breast cancer. Eur. J. Surg. Oncol. 38:307–313, 2012.
Mossotto, E., J. J. Ashton, T. Coelho, R. M. Beattie, B. D. MacArthur, and S. Ennis. Classification of paediatric inflammatory bowel disease using machine learning. Sci. Rep. 7:2427, 2017.
Owjimehr, M., H. Danyali, M. S. Helfroush, and A. Shakibafard. Staging of fatty liver diseases based on hierarchical classification and feature fusion for back-scan–converted ultrasound images. Ultrason. Imaging 39:79–95, 2017.
Pan, Y., W. Huang, Z. Lin, W. Zhu, J. Zhou, J. Wong, and Z. Ding. Brain tumor grading based on neural network s and convolutional neural networks. In: Eng. Med. Biol. Soc. (EMBC), 2015 37th Annu. Int. Conf. IEEE, pp. 699–702, 2015. https://doi.org/10.1109/embc.2015.7318458.
Pota, M., E. Scalco, G. Sanguineti, A. Farneti, G. M. Cattaneo, G. Rizzo, and M. Esposito. Early prediction of radiotherapy-induced parotid shrinkage and toxicity based on CT radiomics and fuzzy classification. Artif. Intell. Med. 2017. https://doi.org/10.1016/j.artmed.2017.03.004.
Reis, S., P. Gazinska, J. Hipwell, T. Mertzanidou, K. Naidoo, N. Williams, S. Pinder, and D. J. Hawkes. Automated classification of breast cancer stroma maturity from histological images. IEEE Trans. Biomed. Eng. 2017. https://doi.org/10.1109/tbme.2017.2665602.
Roy, D. AUTO CONTRAST, 2009. http://www.mathworks.com/matlabcentral/fileexchange/10566-auto-contrast.
Sethi, A., L. Sha, A. R. Vahadane, R. J. Deaton, N. Kumar, V. Macias, and P. H. Gann. Empirical comparison of color normalization methods for epithelial-stromal classification in H and E images. J. Pathol. Inform. 7:17, 2016.
Shin, H. C., H. R. Roth, M. Gao, L. Lu, Z. Xu, I. Nogues, J. Yao, D. Mollura, and R. M. Summers. Deep convolutional neural networks for computer-aided detection: CNN architectures, dataset characteristics and transfer learning. IEEE Trans. Med. Imaging 35:1285–1298, 2016.
Stanford Tissue Microarray Database. https://tma.im/cgi-bin/home.pl.
Tran, H., H. Phan, A. Kumar, J. Kim, and D. Feng. Transfer learning of a convolutional neural network for Hep-2 cell image classification. In: ISBI, pp. 1208–1211, 2016.
Wang, J., X. Yang, H. Cai, W. Tan, C. Jin, and L. Li. Discrimination of breast cancer with microcalcifications on mammography by deep learning. Sci. Rep. 6:27327, 2016.
Xi, K.-X., Y.-S. Wen, C.-M. Zhu, X.-Y. Yu, R.-Q. Qin, X.-W. Zhang, Y.-B. Lin, T.-H. Rong, W.-D. Wang, Y.-Q. Chen, and L.-J. Zhang. Tumor–stroma ratio (TSR) in non-small cell lung cancer (NSCLC) patients after lung resection is a prognostic factor for survival. J. Thorac. Dis. 9:4017–4026, 2017.
Xu, J., X. Luo, G. Wang, H. Gilmore, and A. Madabhushi. A deep convolutional neural network for segmenting and classifying epithelial and stromal regions in histopathological images. Neurocomputing 191:214–223, 2016.
Yang, M., X. Li, Z. Li, Z. Ou, M. Liu, S. Liu, X. Li, and S. Yang. Gene features selection for three-class disease classification via multiple orthogonal partial least square discriminant analysis and S-plot using microarray data. PLoS ONE 8:1–12, 2013.
Yoshida, H., T. Shimazu, T. Kiyuna, A. Marugame, Y. Yamashita, E. Cosatto, H. Taniguchi, S. Sekine, and A. Ochiai. Automated histological classification of whole-slide images of gastric biopsy specimens. Gastric Cancer 2017. https://doi.org/10.1007/s10120-017-0731-8.
Zhang, R., Y. Zheng, T. W. C. Mak, R. Yu, S. H. Wong, J. Y. W. Lau, and C. C. Y. Poon. Automatic detection and classification of colorectal polyps by transferring low-level CNN features from nonmedical domain. IEEE J. Biomed. Heal. Inform 21:41–47, 2017.
Zhou, B., A. Lapedriza, J. Xiao, A. Torralba, and A. Oliva. Learning deep features for scene recognition using places database. Adv. Neural Inf. Process. Syst. 27:487–495, 2014. http://papers.nips.cc/paper/5349-learning-deep-features-for-scene-recognition-using-places-database.pdf.
Acknowledgments
The authors gratefully acknowledge the support from Oklahoma Center for the Advancement of Science & Technology (OCAST) grant HR15-016 and National Institutes of Health (NIH) grant R01 CA197150. This work was also partially sponsored by SCC research award from Stephenson Cancer Center at the University of Oklahoma Health Sciences Center (OUHSC).
Conflict of interest
The authors declare no competing financial interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
Associate Editor Mona Kamal Marei oversaw the review of this article.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Du, Y., Zhang, R., Zargari, A. et al. Classification of Tumor Epithelium and Stroma by Exploiting Image Features Learned by Deep Convolutional Neural Networks. Ann Biomed Eng 46, 1988–1999 (2018). https://doi.org/10.1007/s10439-018-2095-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10439-018-2095-6