Abstract
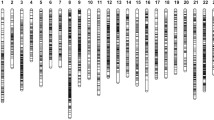
Bighead carp (Hypophthalmichthys nobilis) and silver carp (Hypophthalmichthys molitrix) are genetically close aquaculture fish in the Cyprinidae, which have been confirmed to hold XX/XY sex determination. However, genomic locations of potential sex-related loci in these two fishes are still unknown. In this study, a high-resolution genetic linkage map was constructed by using 2976 SNP and 924 microsatellite markers in a F1 full-sib family of bighead carp, the length of which spanned 2022.34 cM with an average inter-marker distance of 0.52 cM. Comparative genomics revealed a high level of genomic synteny between bighead carp and zebrafish as well as grass carp. QTL fine mapping for sex trait was performed based on this linkage map of bighead carp and an unpublished linkage map of silver carp. A map distance of 3.863 cM (69.787–73.650 cM) on LG19 of bighead carp and 4.705 cM (79.096–83.801 cM) on LG21 of silver carp was significantly associated with sex phenotypes, and these two LGs are homologous between two fish species. Fourteen markers harboring in these regions were in strong linkage disequilibrium with the sex phenotype variance explained (PVE) varying from 89 to 100%. Two common markers were mapped on the QTL regions of bighead carp and silver carp, suggesting that these two carp species may have similar genetic bases for sex determination. Eleven potentially sex-related genes were identified within or near the sex QTL markers in two species. This study provided insights into elucidating mechanisms and evolution of sex determination in cyprinid fishes.






Similar content being viewed by others
References
Anderson JL, Mari AR, Braasch I, Amores A, Hohenlohe P, Batzel P et al (2012) Multiple sex-associated regions and a putative sex chromosome in zebrafish revealed by RAD mapping and population genomics. PLoS One 7:e40701
Ankra-Badu GA, Shriner D, Le Bihan-Duval E, Mignon-Grasteau S, Pitel F, Beaumont C et al (2010) Mapping main, epistatic and sex-specific QTL for body composition in a chicken population divergently selected for low or high growth rate. BMC Genomics 11:107
Bradley KM, Breyer JP, Melville DB, Broman KW, Knapik EW, Smith JR (2011) An SNP-based linkage map for zebrafish reveals sex determination loci. G3 (Bethesda) 1:3–9
Brown JK et al (2016) Mapping the sex determination locus in the hapuku (Polyprion oxygeneios) using ddRAD sequencing. BMC Genomics 17:448
Chakravarti A, Lasher LK, Reefer JE (1991) A maximum likelihood method for estimating genome length using genetic linkage data. Genetics 128:175–182
Chen J, Xie P, Zhang DW, Lei HH (2007) In situ studies on the distribution patterns and dynamics of microcystins in a biomanipulation fish - bighead carp (Aristichthys nobilis). Environ Pollut 147:150–157
Cui Z et al (2015) High-density linkage mapping aided by transcriptomics documents ZW sex determination system in the Chinese mitten crab Eriocheir sinensis. Heredity 115:206–215
Davey JW, Hohenlohe PA, Etter PD, Boone JQ, Catchen JM, Blaxter ML (2011) Genome-wide genetic marker discovery and genotyping using next generation sequencing. Nat Rev Genet 12:499–510
Devlin RH, Nagahama Y (2002) Sex determination and sex differentiation in fish: an overview of genetic, physiological, and environmental influences. Aquaculture 208:191–364
Ezaz T, Stiglec R, Veyrunes F, Jennifer A, Graves M (2006) Relationships between vertebrate ZW and XY sex chromosome systems. Curr Biol 16:5
Feng X et al (2018) A high-resolution genetic linkage map and QTL fine mapping for growth-related and sex in the Yangtze River common carp (Cyprinus carpio haematopterus). BMC Genomics 19:230
Fishman EL, Kelly HA, Morgan LE, Willis JH (2001) A genetic map in the Mimulus guttatus species complex reveals transmission ratio distortion due to heterospecific interactions. Genetics 159:1701–1716
Froschauer A, Körting C, Katagiri T (2002) Construction and initial analysis of bacterial artificial chromosome (BAC) contigs from the sex-determining region of the platyfish Xiphophorus maculatus. Gene 295:247–254
Fu XT et al (2013) RADtyping: an integrated package for accurate de novo codominant and dominant RAD genotyping in mapping populations. PLoS One 8:e79960
Fu BD, Liu HY, Yu XM, Tong JG (2016) A high-density genetic map and growth related QTL mapping in bighead carp (Hypophthalmichthys nobilis). Sci Rep 6:28679
Gagnaire PA, Normandeau E, Pavey SA, Bernatchez L (2013) Mapping phenotypic, expression and transmission ratio distortion QTL using RAD markers in the Lake Whitefish (Coregonus clupeaformis). Mol Ecol 22:3036–3048
Gamble T, Zarkower D (2014) Identification of sex-specific molecular markers using restriction site-associated DNA sequencing. Mol Ecol Resour 14:902–913
Gilbert JR (2005) A field guide to north atlantic wildlife: marine mammals, seabirds, fish and other sea life, Noble S. Proctor and Patrick J. Lynch. Yale University, New Haven, Connecticut, and London, United Kingdom. Wildl Soc Bull 34:898–899
Guo W et al (2013) A second generation genetic linkage map for silver carp (Hypophthalmichthys molitrix) using microsatellite markers. Aquaculture 412-413:97–106
Hattori RS, Gould RJ, Fujioka T, Saito T, Kurita J, Strüssmann CA, Yokota M, Watanabe S (2007) Temperature-dependent sex-determination in Hd-rR medaka Oryzias latipes: gender sensitivity, thermal threshold, critical period, and DMRT1 expression profile. Sex Dev 1:138–146
Kamiya T et al (2012) A trans-species missense SNP in Amhr2 is associated with sex-determination in the tiger pufferfish, Takifugu rubripes (fugu). PLoS Genet 8:e1002798
Kawajiri M, Yoshida K, Fujimoto S, Mokodongan DF, Ravinet M, Kirkpatrick M, Yamahira K, Kitano J (2014) Ontogenetic stage-specific quantitative trait loci contribute to divergence in developmental trajectories of sexually dimorphic fins between medaka populations. Mol Ecol 23:5258–5275
Koji F, Conte MA, Kocher TD et al (2011) Circular DNA intermediate in the duplication of Nile tilapia vasa genes. PLoS One 6:e29477
Kolar CS, Chapman DC, Courtenay WR Jr, Housel CM, Williams JD, Jennings DP (2007) Bigheaded carps: A biological synopsis and environmental risk assessment. American Fisheries Society, Special Publica on 33, Bethesda
Korstanje R, Li R, Howard T, Kelmenson P, Marshall J, Paigen B, Churchill G (2004) Influence of sex and diet on quantitative trait loci for HDL cholesterol levels inan SM/J by NZB/BlNJ intercross population. J Lipid Res 45:881–888
Krzywinski M, Schein J, Birol I (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645
Liao M, Zhang L, Yang G, Zhu M, Wang D, Wei Q, Zou G, Chen D (2007) Development of silver carp (Hypophthalmichthys molitrix) and bighead carp (Aristichthys nobilis) genetic maps using microsatellite and AFLP markers and a pseudo-testcross strategy. Anim Genet 38:364–370
Liu HY et al (2016) QTL fine mapping and identification of candidate genes for growth-related traits in bighead carp (Hypophthalmichthys nobilis). Aquaculture 465:134–143
Liu HY et al (2018) Sex-specific markers developed by next generation sequencing confirmed an XX/XY sex determination system in bighead carp (Hypophthalmichthys nobilis) and silver carp (Hypophthalmichthys molitrix). DNA Res 25:257–254
Llamas B, Jiang Z, Rainville ML, Picard S, Deschepper CF (2005) Distinct QTLs are linked to cardiac left ventricular mass in a sex-specific manner in a normotensive inbred rat intercross. Mamm Genome 16:700–711
Luhmann LM, Knorr C, Horstgen-Schwark G, Wessels S (2012) First evidence for family-specific QTL for temperature-dependent sex reversal in Nile Tilapia (Oreochromis niloticus). Sex Dev 6:247–256
Martínez P et al (2009) Identification of the major sex-determining region of turbot (Scophthalmus maximus). Genetics 183:1443–1452
Martínez P, Viñas AM, Sánchez L, Díaz N, Ribas L, Piferrer F (2014) Genetic architecture of sex determination in fish: applications to sex ratio control in aquaculture. Front Genet 5:340
Massault C, Bovenhuis H, Haley C, Koning D (2008) QTL mapping designs for aquaculture. Aquaculture 285:23–29
Mei J, Gui JF (2015) Genetic basis and biotechnological manipulation of sexual dimorphism and sex determination in fish. Sci China 58:124
Myosho T et al (2012) Tracing the emergence of a novel sex-determining gene in medaka, Oryzias luzonensis. Genetics 191:163–170
Palaiokostas C et al (2013a) Mapping the sex determination locus in the Atlantic halibut (Hippoglossus hippoglossus) using RAD sequencing. BMC Genomics 14:566
Palaiokostas C, Bekaert M, Khan MG, Taggart JB, Gharbi K, McAndrew B, Penman DJ (2013b) Mapping and validation of the major sex-determining region in Nile Tilapia (Oreochromis niloticus L.) using RAD sequencing. PLoS One 8:e68389
Palaiokostas C et al (2015a) A new SNP-based vision of the genetics of sex determination in European sea bass (Dicentrarchus labrax). Genet Sel Evol 47:68
Palaiokostas C et al (2015b) A novel sex-determining QTL in Nile tilapia (Oreochromis niloticus). BMC Genomics 16:171
Penman DJ, Piferrer F (2008) Fish gonadogenesis. Part I: genetic and environmental mechanisms of sex determination. Rev Fish Sci 16:16–34
Piferrer F, Guiguen Y (2008) Fish gonadogenesis. Part II: molecular biology and genomics of sex differentiation. Rev Fish Sci 16:35–55
Pompini M, Buser AM, Thali MR, von Siebenthal B, Nusslé S, Guduff S, Wedekind C (2013) Temperature-induced sex reversal is not responsible for sex-ratio distortions in grayling Thymallus thymallus or brown trout Salmo trutta. J Fish Biol 83:404–411
Sambrook J, Russell D (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Santos D, Luzio A, Coimbra AM (2017) Zebrafish sex differentiation and gonad development: a review on the impact of environmental factors. Aquat Toxicol 191:141–163
Slate J, Gratten J, Beraldi D, Stapley J, Hale M, Pemberton JM (2009) Gene mapping in the wild with SNPs: guidelines and future directions. Genetica 136:97–107
Song W et al (2012) Construction of high density microsatellite genetic linkage map and mapping of sexual and growth-related traits in half-smooth tongue sole (Cynoglossus semilaevis). PLoS One 7:e52097
Sun YL et al (2014) Screening and characterization of sex-linked DNA markers and marker-assisted selection in the Nile tilapia (Oreochromis niloticus). Aquaculture 433:19–27
Sun C et al (2017) Construction of a high-density linkage map and mapping of sex determination and growth-related loci in the mandarin fish (Siniperca chuatsi). BMC Genomics 18:446
Tong JG, Sun XW (2015) Genetic and genomic analyses for economically important traits and their applications in molecular breeding of cultured fish. Sci China Life Sci 58:178–186
Tríbulo P, Siqueira LGB, Oliveira LJ, Scheffler T, Hansen PJ (2018) Identification of potential embryokines in the bovine reproductive tract. J Dairy Sci 101:690–704
Vale L, Dieguez R, Sánchez L, Martínez P, Viñas A (2014) A sex-associated sequence identified by RAPD screening in gynogenetic individuals of turbot (Scophthalmus maximus). Mol Biol Rep 41:1501–1509
Van O (2006) JoinMap 4. Software for the calculation of genetic linkage maps in experimental populations. Kyazma BV, Wageningen, Netherlands
Vinas A et al (2012) Mapping of DNA sex-specific markers and genes related to sex differentiation in Turbot (Scophthalmus maximus). Mar Biotechnol 14:655–663
Volff JN, Nanda I, Schmid M, Schartl M (2007) Governing sex determination in fish: regulatory putsches and ephemeral dictators. Sex Dev 1:85–99
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Wang S, Meyer E, McKay JK, Matz MV (2012) 2b-RAD: a simple and flexible method for genome-wide genotyping. Nat Methods 9:808–910
Wang YP et al (2015) The draft genome of the grass carp (Ctenopharyngodon idellus) provides insights into its evolution and vegetarian adaptation. Nat Genet 47:625–631
Wang L, Xie N, Shen Y, Ye B, Yue GH, Feng X (2019) Constructing high-density genetic maps and developing sexing markers in northern snakehead (Channa argus). Mar Biotechnol 21:348–358
Wilson CA, High SK, McCluskey BM, Amores A, Yan YL, Titus TA et al (2014) Wild sex in zebrafish: loss of the natural sex determinant indomesticated strains. Genetics 198:1291
Zhu C, Tong J, Yu X, Guo W (2015) Comparative mapping for bighead carp (Aristichthys nobilis) against model and non-model fishes provides insights into the genomic evolution of cyprinids. Mol Gen Genomics 290:1313–1326
Acknowledgments
We would like to thank all colleagues of our lab (Meixia Pang, Xueli Liu, Baojiang Gan) for sample preparation and laboratory technical assistance.
Funding
This research was supported by the National Key Research and Development Program (No. 2018YFD009102) and State Key Laboratory of Freshwater Ecology and Biotechnology (2019FBZ04).
Author information
Authors and Affiliations
Contributions
JT conceived this study and guided the experiments. YZ and HL carried out the experiments and wrote the manuscript. BF performed bioinformatic analysis. XW and XY performed experiments and prepared mapping animals. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Table S1
The information of primers used for qPCR in this study (XLS 20 kb)
Table S2
The information of 3,900 markers located in the high-density linkage map of bighead carp (XLSX 358 kb)
Figure S1
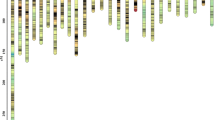
The lengths and marker distributions of 24 linkage groups in the female map of bighead carp (PNG 200 kb)
Figure S2
The lengths and marker distributions of 24 linkage groups in the male map of bighead carp (PNG 140 kb)
Figure S3
The syntenic relationships between female (left) and male (right) linkage groups of bighead carp. The common markers are indicated with lines (PNG 527 kb)
Rights and permissions
About this article
Cite this article
Zhou, Y., Liu, H., Wang, X. et al. QTL Fine Mapping for Sex Determination Region in Bighead Carp (Hypophthalmichthys nobilis) and Comparison with Silver Carp (Hypophthalmichthys molitrix). Mar Biotechnol 22, 41–53 (2020). https://doi.org/10.1007/s10126-019-09929-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-019-09929-3