Abstract
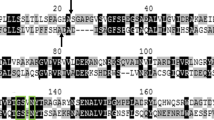
Non-specific nuclease (NSN) can be applied in industrial downstream processing to remove nucleic acids from crude protein extracts or in cell-sorting systems to degrade nucleic acids derived from lysed cells. PsNuc from the ice-nucleating bacterium Pseudomonas syringae has the ability to decompose double- and single-stranded DNA in linear or circular form and RNA. It is not affected by the presence of metal-ion chelators such as EDTA and tolerates several protease inhibitors and reducing agents. A multiple sequence alignment of PsNuc with closely related enzymes (97–99% identity on the protein level) within the family Pseudomonaceae revealed the presence of only six amino acid residues that are variable in putative NSN from different members of the genus Pseudomonas. Single amino acid variants were produced in recombinant form in Escherichia coli, purified, and characterized. They showed similar activity compared to PsNuc, but a single variant even displayed an improved performance with an activity of > 20,000 U/mg at 35 °C, while amino acid residues S148 and V161 were found to be essential for enzymatic functionality. These results suggest that homologous nucleases from Pseudomonaceae display high activity levels in a metal-ion-independent manner and are therefore of interest for applications in biotechnology.





Similar content being viewed by others
Abbreviations
- AEBSF:
-
4-(2-Aminoethyl)benzenesulfonyl fluoride hydrochloride
- dsDNA:
-
Double-stranded deoxyribonucleic acid
- DTT:
-
Dithiothreitol
- EDTA:
-
Ethylenediaminetetraacetic acid
- IPTG:
-
Isopropyl β-d-1-thiogalactopyranoside
- NSN:
-
Non-specific nuclease
- PMSF:
-
Phenylmethylsulfonyl fluoride
- RNA:
-
Ribonucleic acid
- ssDNA:
-
Single-stranded deoxyribonucleic acid
- SP:
-
Signal peptide
References
Bao, Y., Higgins, L., Zhang, P., Chan, S. H., Laget, S., Sweeney, S., et al. (2008). Expression and purification of BmrI restriction endonuclease and its N-terminal cleavage domain variants. Protein Expression and Purification,58, 42–52.
Belkebir, A., & Azeddoug, H. (2012). Characterization of LlaKI, a new metal ion-independent restriction endonuclease from Lactococcus lactis KLDS4. ISRN Biochemistry,2012, 287230.
Biasini, M., Bienert, S., Waterhouse, A., Arnold, K., Studer, G., Schmidt, T., et al. (2014). SWISS-MODEL: Modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Research,42, W252–W258.
Binnenkade, L., Kreienbaum, M., & Thormann, K. M. (2018). Characterization of ExeM, an extracellular nuclease of Shewanella oneidensis MR-1. Frontiers in Microbiology,9, 1761.
Conant, G. C., & Wolfe, K. H. (2008). Turning a hobby into a job: How duplicated genes find new functions. Nature Reviews Genetics,9, 938–950.
Damnjanovic, J., & Iwasaki, Y. (2013). Phospholipase D as a catalyst: Application in phospholipid synthesis, molecular structure and protein engineering. Journal of Bioscience and Bioengineering,116, 271–280.
Elleuche, S., Schröder, C., Sahm, K., & Antranikian, G. (2014). Extremozymes–biocatalysts with unique properties from extremophilic microorganisms. Current Opinion in Biotechnology,29, 116–123.
Gottlin, E. B., Rudolph, A. E., Zhao, Y., Matthews, H. R., & Dixon, J. E. (1998). Catalytic mechanism of the phospholipase D superfamily proceeds via a covalent phosphohistidine intermediate. Proc Natl Acad Sci USA,95, 9202–9207.
Grazulis, S., Manakova, E., Roessle, M., Bochtler, M., Tamulaitiene, G., Huber, R., et al. (2005). Structure of the metal-independent restriction enzyme BfiI reveals fusion of a specific DNA-binding domain with a nonspecific nuclease. Proc Natl Acad Sci USA,102, 15797–15802.
Horikoshi, K., Antranikian, G., Bull, A. T., Robb, F. T., & Stetter, K. O. (2011). Extremophiles handbook (1st ed.). New York: Springer.
Jones, D. T. (1999). Protein secondary structure prediction based on position-specific scoring matrices. Journal of Molecular Biology,292, 195–202.
Larkin, M. A., Blackshields, G., Brown, N. P., Chenna, R., McGettigan, P. A., McWilliam, H., et al. (2007). Clustal W and Clustal X version 2.0. Bioinformatics,23, 2947–2948.
Li, L., Lin, S., & Yanga, F. (2005). Functional identification of the non-specific nuclease from white spot syndrome virus. Virology,337, 399–406.
Li, L., & Rohrmann, G. F. (2000). Characterization of a baculovirus alkaline nuclease. Journal of Virology,74, 6401–6407.
Maciejewska, N., Walkusz, R., Olszewski, M., & Szymanska, A. (2019). New nuclease from extremely psychrophilic microorganism Psychromonas ingrahamii 37: Identification and Characterization. Molecular Biotechnology,61, 122–133.
MacLellan, S. R., & Forsberg, C. W. (2001). Properties of the major non-specific endonuclease from the strict anaerobe Fibrobacter succinogenes and evidence for disulfide bond formation in vivo. Microbiology,147, 315–323.
Meiss, G., Franke, I., Gimadutdinow, O., Urbanke, C., & Pingoud, A. (1998). Biochemical characterization of Anabaena sp. strain PCC 7120 non-specific nuclease NucA and its inhibitor NuiA. European Journal of Biochemistry,251, 924–934.
Miltenyi, S., Hübel, T., & Nölle, V. (2018). Process for sorting cells by microfabricated components using a nuclease. USA Patent US 10018541 B2, July 10, 2018.
Nielsen, H. (2017). Predicting secretory proteins with signalP. Methods in Molecular Biology,1611, 59–73.
Nilsen, I. W., Overbo, K., Jensen Havdalen, L., Elde, M., Gjellesvik, D. R., & Lanes, O. (2010). The enzyme and the cDNA sequence of a thermolabile and double-strand specific DNase from Northern shrimps (Pandalus borealis). PLoS ONE,5, e10295.
Pommer, A. J., Wallis, R., Moore, G. R., James, R., & Kleanthous, C. (1998). Enzymological characterization of the nuclease domain from the bacterial toxin colicin E9 from Escherichia coli. Biochemical Journal,334(Pt 2), 387–392.
Rangarajan, E. S., & Shankar, V. (2001). Sugar non-specific endonucleases. FEMS Microbiology Reviews,25, 583–613.
Rangarajan, S., & Shankar, V. (1999). Extracellular nuclease from Rhizopus stolonifer: purification and characteristics of single strand preferential—deoxyribonuclease activity. Biochimica et Biophysica Acta,1473, 293–304.
Rudolph, A. E., Stuckey, J. A., Zhao, Y., Matthews, H. R., Patton, W. A., Moss, J., et al. (1999). Expression, characterization, and mutagenesis of the Yersinia pestis murine toxin, a phospholipase D superfamily member. Journal of Biological Chemistry,274, 11824–11831.
Schaumburg, F., Pauly, M., Schubert, G., Shittu, A., Tong, S., Leendertz, F., et al. (2014). Characterization of a novel thermostable nuclease homolog (NucM) in a highly divergent Staphylococcus aureus clade. Journal of Clinical Microbiology,52, 4036–4038.
Schmitz, S., Börner, P., Nölle, V., & Elleuche, S. (2019). Comparative analysis of two non-specific nucleases of the phospholipase D family from the plant pathogen competitor bacterium Pantoea agglomerans. Applied Microbiology and Biotechnology,103, 2635–2648.
Schmitz, S., Nölle, V., & Elleuche, S. (2019). A non-specific nucleolytic enzyme and its application potential in EDTA-containing buffer solutions. Biotechnology Letters,41, 129–136.
Song, Q., & Zhang, X. (2008). Characterization of a novel non-specific nuclease from thermophilic bacteriophage GBSV1. BMC Biotechnology,8, 43.
Stuckey, J. A., & Dixon, J. E. (1999). Crystal structure of a phospholipase D family member. Natural Structural Biology,6, 278–284.
Sung, T. C., Roper, R. L., Zhang, Y., Rudge, S. A., Temel, R., Hammond, S. M., et al. (1997). Mutagenesis of phospholipase D defines a superfamily including a trans-Golgi viral protein required for poxvirus pathogenicity. EMBO Journal,16, 4519–4530.
Wang, D., Miyazono, K. I., & Tanokura, M. (2016). Tetrameric structure of the restriction DNA glycosylase R.PabI in complex with nonspecific double-stranded DNA. Scientific Reports,6, 35197.
Wilkinson, S. G. (1967). The sensitivity of pseudomonads to ethylenediaminetetra-acetic acid. Journal of General Microbiology,47, 67–76.
Xin, X. F., Kvitko, B., & He, S. Y. (2018). Pseudomonas syringae: What it takes to be a pathogen. Nature Reviews Microbiology,16, 316–328.
Yang, W. (2011). Nucleases: Diversity of structure, function and mechanism. Quarterly Reviews of Biophysics,44, 1–93.
Zhao, Y., Stuckey, J. A., Lohse, D. L., & Dixon, J. E. (1997). Expression, characterization, and crystallization of a member of the novel phospholipase D family of phosphodiesterases. Protein Science,6, 2655–2658.
Acknowledgements
This work was funded by Miltenyi Biotec B.V. & Co. KG. The authors thank Jens Helmer (Miltenyi Biotec B.V. & Co. KG) for peptide mass fingerprinting, mass spectrometry analysis, and for the preparation of Supplementary Fig. S1.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
SS, MW, VN, and SE are employees of Miltenyi Biotec B.V. & Co. KG.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Schmitz, S., Wieczorek, M., Nölle, V. et al. Characterization of Single Amino Acid Variations in an EDTA-Tolerating Non-specific Nuclease from the Ice-Nucleating Bacterium Pseudomonas syringae. Mol Biotechnol 62, 67–78 (2020). https://doi.org/10.1007/s12033-019-00229-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-019-00229-8