Abstract
The prokaryotic communities of water bodies contaminated by acid mine drainage from the São Domingos mining area in southern Portugal were analyzed using a meta-taxonomics approach with 16S rRNA gene sequences. Samples were collected in two seasonal sampling campaigns (summer and winter of 2017) from the most contaminated sites from where the water flows downstream to the freshwater reservoir of the river Chança. The physicochemical data indicate a trend of decreasing acid mine drainage contamination downstream of the mining area to the Chança’s reservoir. The most contaminated sites (pH = 2.3–3.1) are distinguished by prokaryotic diversity with high abundances of operational taxonomics units related to acidophiles (genera Metallibacterium, Acidibacter, Leptospirillum, Acidobacterium, Thiomonas, Acidicapsa, Acidocella, Acidiphilium; family Acidobacteriaceae, order CPla-3 termite group). Likewise, in the transition zone in the mouth of the contaminated water flow into the Chança´s reservoir (pH = 6.4), a specific prokaryotic flora exists with some acidophiles, but notably with a cyanobacteria bloom and a high abundance of the genus Sediminibacterium (family I; order Subsection III). Moreover, the strong correlation between the abundance of acidophiles and characteristic physiochemical parameters (metals, acidity, and sulfate) confirm their potential as biomarkers of acid mine drainage pollution.



Similar content being viewed by others
References
Abdi H, Williams LJ (2010) Principal component analysis. WIREs Comp. Stat 2:433–459. https://doi.org/10.1002/wics.101
Abreu MM, Tavares MT, Batista MJ (2008) Potential use of Erica andevalensis and Erica australis in phytoremediation of sulphide mine environments: São Domingos, Portugal. J Geochem Explor 96:210–222. https://doi.org/10.1016/j.gexplo.2007.04.007
Albertsen M, Karst SM, Ziegler AS, Kirkegaard RH, Nielsen PH (2015) Back to basics—the Influence of DNA extraction and primer choice on phylogenetic analysis of activated sludge communities. PLoS ONE 10(7):e0132783. https://doi.org/10.1371/journal.pone.0132783
Alvarenga P, Palma P, de Varennes A, Cunha-queda AC (2012) A contribution towards the risk assessment of soils from the São Domingos Mine (Portugal): chemical, microbial and ecotoxicological indicators. Environ Pollut 161:50–56. https://doi.org/10.1016/j.envpol.2011.09.044
Álvarez-Valero AM, Pérez-López R, Matos J, Capitán MA, Nieto JM, Sáez R, Delgado J, Caraballo M (2008) Potential environmental impact at São Domingos mining district (Iberian Pyrite Belt, SW Iberian Peninsula): evidence from a chemical and mineralogical characterization. Environ Geol 55:1797–1809. https://doi.org/10.1007/s00254-007-1131-x
Barns SM, Takala SL, Kuske CR (1999) Wide distribution and diversity of members of the bacterial kingdom acidobacterium in the environment. Appl Environ Microbiol 65:1731–1737
Bond PL, Smriga SP, Banfield JF (2000) Phylogeny of microorganisms populating a thick, subaerial, predominantly lithotrophic biofilm at an extreme acid mine drainage site. Appl Environ Microbiol 66:3842–3849. https://doi.org/10.1128/AEM.66.9.3842-3849.2000
Bruneel O, Duran R, Casiot C, Elbaz-Poulichet F, Personné J-C (2006) Diversity of microorganisms in Fe–As-rich acid mine drainage waters of Carnoulès, France. Appl Environ Microbiol 72:551–556. https://doi.org/10.1128/AEM.72.1.551-556.2006
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA 108:4516–4522. https://doi.org/10.1073/pnas.1000080107
Chen W, Simpson J, Levesque CA (2018) RAM: R for amplicon-sequencing-based microbial-ecology. R package version 1.2.1.7. https://cran.r-project.org/package=RAM
Clarke A, Raymond J, Leakey RJG (1996) The seasonal cycle of phytoplankton, macronutrients, and the microbial community in a nearshore antarctic marine ecosystem. Limnol Oceanogr 41:1281–1294. https://doi.org/10.4319/lo.1996.41.6.1281
Cowan DA, Ramond J-B, Makhalanyane TP, De Maayer P (2015) Metagenomics of extreme environments. Curr Opin Microbiol 25:97–102. https://doi.org/10.1016/j.mib.2015.05.005
Dias-Sardinha I, Craveiro D, Milheiras S (2013) A sustainability framework for redevelopment of rural brownfields: stakeholder participation at SÃO DOMINGOS mine, Portugal. J Cleaner Prod 57:200–208. https://doi.org/10.1016/j.jclepro.2013.05.042
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998. https://doi.org/10.1038/nmeth.2604
Edwards KJ, Gihring TM, Banfield JF (1999) Seasonal variations in microbial populations and environmental conditions in an extreme acid mine drainage environment. Appl Environ Microbiol 65:3627–3632
García-Moyano A, Austnes AE, Lanzén A, González-Toril E, Aguilera A, Øvreås L (2015) Novel and unexpected microbial diversity in acid mine drainage in Svalbard (78°N), revealed by culture-independent approaches. Microorganisms 3:667–694. https://doi.org/10.3390/microorganisms3040667
Gilbert JA, Field D, Swift P, Newbold L, Oliver A, Smyth T, Somerfield PJ, Huse S, Joint I (2009) The seasonal structure of microbial communities in the Western English Channel. Environ Microbiol 11:3132–3139. https://doi.org/10.1111/j.1462-2920.2009.02017.x
González-Toril E, Aguilera A, Rodriguez N, Fernández-Remolar D, Gómez F, Diaz E, García-Moyano A, Sanz JL, Amils R (2010) Microbial ecology of Río Tinto, a natural extreme acidic environment of biohydrometallurgical interest. Hydrometallurgy 104:329–333. https://doi.org/10.1016/j.hydromet.2010.01.011
González-Toril E, Aguilera A, Souza-Egipsy V, Pamo EL, España JS, Amils R (2011) Geomicrobiology of La Zarza-Perrunal acid mine effluent (Iberian Pyritic Belt, Spain). Appl Environ Microbiol 77:2685–2694. https://doi.org/10.1128/AEM.02459-10
Hao C-B, Zhang H-X, Bai Z-H, Hu Q, Zhang B-G (2007) A novel acidophile community populating waste ore deposits at an acid mine drainage site. J Environ Sci 19:444–450. https://doi.org/10.1016/S1001-0742(07)60074-6
Hubbard CG, Black S, Coleman ML (2009) Aqueous geochemistry and oxygen isotope compositions of acid mine drainage from the Río Tinto, SW Spain, highlight inconsistencies in current models. Chem Geol 265:321–334. https://doi.org/10.1016/J.CHEMGEO.2009.04.009
Illumina I (2015) 16S metagenomic sequencing library preparation. Part # 15044223 Rev. B. https://support.illumina.com/downloads/16s_metagenomic_sequencing_library_preparation.html
Janda JM, Abbott SL (2007) 16S rRNA gene sequencing for bacterial identification in the diagnostic laboratory: pluses, perils, and pitfalls. J Clin Microbiol 45:2761–2764. https://doi.org/10.1128/JCM.01228-07
Jovel J, Patterson J, Wang W, Hotte N, O'Keefe S, Mitchel T, Perry T, Kao D, Mason AL, Madsen KL, Wong GK-S (2016) Characterization of the Gut microbiome using 16S or shotgun metagenomics. Front Microbiol 7:459. https://doi.org/10.3389/fmicb.2016.00459
Kennedy J, O’Leary ND, Kiran GS, Morrissey JP, O’Gara F, Selvin J, Dobson ADW (2011) Functional metagenomic strategies for the discovery of novel enzymes and biosurfactants with biotechnological applications from marine ecosystems. J Appl Microbiol 111:787–799. https://doi.org/10.1111/j.1365-2672.2011.05106.x
Keshri J, Pradeep Ram AS, Sime-Ngando T (2018) Distinctive patterns in the taxonomical resolution of bacterioplankton in the sediment and pore waters of contrasted freshwater lakes. Microb Ecol 75:662–673. https://doi.org/10.1007/s00248-017-1074-z
Kim J, Koo S-Y, Kim J-Y, Lee EH, Lee S-D, Ko K-S, Dong-Chan K, Cho K-S (2009) Influence of acid mine drainage on microbial communities in stream and groundwater samples at Guryong Mine. S Korea Environ Geol 58:1567. https://doi.org/10.1007/s00254-008-1663-8
Kishimoto N, Kosako Y, Tano T (1991) Acidobacterium capsulatum gen. nov., sp. nov.: an acidophilic chemoorganotrophic bacterium containing menaquinone from acidic mineral environment. Curr Microbiol 22:1–7. https://doi.org/10.1007/BF02106205
Kuang J-L, Huang L-N, Chen L-X, Hua Z-S, Li S-J, Hu M, Li JT, Shu W-S (2013) Contemporary environmental variation determines microbial diversity patterns in acid mine drainage. ISME J 7:1038–1050. https://doi.org/10.1038/ismej.2012.139
Laplante K, Sébastien B, Derome N (2013) Parallel changes of taxonomic interaction networks in lacustrine bacterial communities induced by a polymetallic perturbation. Evol Appl 6:643–659. https://doi.org/10.1111/eva.12050
Lee D-G, Trujillo ME, Kang S, Nam J, Kim Y (2018) Epidermidibacterium keratini gen. nov., sp. nov., a member of the family Sporichthyaceae, isolated from keratin epidermis. Int J Syst Evol Microbiol 68:745–750. https://doi.org/10.1099/ijsem.0.002579
Llirós M, Inceoğlu Ö, García-Armisen T, Anzil A, Leporcq B, Pigneur L-M, Viroux L, Darchambeau F, Descy J-P, Servais P (2014) Bacterial community composition in three freshwater reservoirs of different alkalinity and trophic status. PLoS ONE 9:e116145. https://doi.org/10.1371/journal.pone.0116145
Marchesi JR, Ravel J (2015) The vocabulary of microbiome research: a proposal. Microbiome 3:31. https://doi.org/10.1186/s40168-015-0094-5
Martins M, Santos ES, Leonor M, Chaves S, Tenreiro R, Barros RJ, Barreiros A, Costa MC (2011) Performance and bacterial community shifts during bioremediation of acid mine drainage from two Portuguese mines. Int Biodeterior Biodegrad 65:972–981. https://doi.org/10.1016/j.ibiod.2011.07.006
MATLAB and Statistics Toolbox Release (2016) The MathWorks, Inc., Natick, Massachusetts, United States
Mcilroy SJ, Kirkegaard RH, McIlroy B, Nierychlo M, Kristensen JM, Karst SM, Albertsen M, Nielsen PH (2017) MiDAS 2.0: an ecosystem-specific taxonomy and online database for the organisms of wastewater treatment systems expanded for anaerobic digester groups. Database 2017:1–9. https://doi.org/10.1093/database/bax016
Mekki A, Sayadi S (2017) Study of heavy metal accumulation and residual toxicity in soil saturated with phosphate processing wastewater. Water Air Soil Pollut 228:215. https://doi.org/10.1007/s11270-017-3399-0
Normand P (2006) The Families Frankiaceae, Geodermatophilaceae, Acidothermaceae and Sporichthyaceae. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (eds) The prokaryotes. Springer, New York, pp 669–681. https://doi.org/10.1007/0-387-30743-5-26
Okabayashi A, Wakai S, Kanao T, Sugio T, Kamimura K (2005) Diversity of 16S ribosomal DNA-defined bacterial population in acid rock drainage from Japanese pyrite mine. J Biosci Bioeng 100:644–652. https://doi.org/10.1263/jbb.100.644
Pereira R, Ribeiro R, Gonçalves F (2015) Plan for an Integrated human and environmental risk assessment in the S. Domingos Mine Area (Portugal). Hum Ecol Risk Assess 10:543–578. https://doi.org/10.1080/10807030490452197
Pérez-López R, Delgado J, Nieto JM, Márquez-García B (2010) Rare earth element geochemistry of sulphide weathering in the São Domingos mine area (Iberian Pyrite Belt): a proxy for fluid–rock interaction and ancient mining pollution. Chem Geol 276:29–40. https://doi.org/10.1016/j.chemgeo.2010.05.018
Podda F, Zuddas P, Minacci A, Pepi M, Baldi F (2000) Heavy metal coprecipitation with hydrozincite [Zn5(CO3)2(OH)6] from mine waters caused by photosynthetic microorganisms. Appl Environ Microbiol 66:5092–5098. https://doi.org/10.1128/AEM.66.11.5092-5098.2000
R Core Team (2017) R version 1.0.153: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Reis MP, Dias MF, Costa PS, Ávila MP, Leite LR, de Araújo FMG, Salim ACM, Bucciarelli-Rodriguez M et al (2016) Metagenomic signatures of a tropical mining-impacted stream reveal complex microbial and metabolic networks. Chemosphere 161:266–273. https://doi.org/10.1016/j.chemosphere.2016.06.097
Salomons W (1995) Environmental impact of metals derived from mining activities: processes, predictions, prevention. J Geochem Explor 52:5–23. https://doi.org/10.1016/0375-6742(94)00039-E
Schrenk MO, Edwards KJ, Goodman RM, Hamers RJ, Banfield JF (1998) Distribution of Thiobacillus ferrooxidans and Leptospirillum ferrooxidans: implications for generation of acid mine drainage. Science 279:1519–1522. https://doi.org/10.1126/science.279.5356.1519
Shannon CE (1948) A mathematical theory of communication. Bell Syst Tech J 27:379–423. https://doi.org/10.1002/j.1538-7305.1948.tb01338.x
Simpson EH (1949) Measurement of diversity. Nature 163:688. https://doi.org/10.1038/163688a0
Spellerberg IF, Fedor PJ (2003) A tribute to Claude Shannon (1916–2001) and a plea for more rigorous use of species richness, species diversity and the ‘Shannon–Wiener’ Index. Global Ecol Biogeogr 12:177–179. https://doi.org/10.1046/j.1466-822X.2003.00015.x
Stewart EJ (2012) Growing unculturable bacteria. J Bacteriol 194:4151–4160. https://doi.org/10.1128/JB.00345-12
Tavares MT, Sousa AJ, Abreu MM (2008) Ordinary kriging and indicator kriging in the cartography of trace elements contamination in São Domingos mining site (Alentejo, Portugal). J Geochem Explor 98:43–56. https://doi.org/10.1016/j.gexplo.2007.10.002
Thomas T, Gilbert J, Meyer F (2012) Metagenomics—a guide from sampling to data analysis. Microb Inform Exp 2:3. https://doi.org/10.1186/2042-5783-2-3
Vicente-Villardón JL (2015) MULTBIPLOT version 16.0430: a package for multivariate analysis using biplots Departamento de Estadística. Universidad de Salamanca. Spain. https://biplot.usal.es/ClassicalBiplot/index.html
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/AEM.00062-07
Zhang X, Tang S, Wang M, Sun W, Xie Y, Peng H, Zhong A, Liu H et al (2019) Acid mine drainage affects the diversity and metal resistance gene profile of sediment bacterial community along a river. Chemosphere 217:790–799. https://doi.org/10.1016/j.chemosphere.2018.10.210
Ziegler S, Waidner B, Itoh T, Schumann P, Spring S, Gescher J (2013) Metallibacterium scheffleri gen. nov., sp. nov., an alkalinizing gammaproteobacterium isolated from an acidic biofilm. Int J Syst Evol Microbiol 63:1499–1504. https://doi.org/10.1099/ijs.0.042986-0
Acknowledgements
Sampling and samples preparation received Portuguese national funds from the Foundation for Science and Technology (FCT) through project UID/Multi/04326/2019. The bioinformatics and interpretation of the results were framed in the project METALCHEMBIO (no. 29251) financed by national funds through the FCT and co-financed by the Algarve´s Regional Operational Program (CRESC Algarve 2020), through Portugal 2020 and European Regional Development Fund (FEDER).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Atomi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
792_2019_1136_MOESM1_ESM.pdf
Supplementary Figure 1: ESM 1. Numbers of retrieved sequence reads and operational taxonomic units (OTUs) from each sample as well as respective Shannon and Simpson diversity indices (PDF 203 kb)
792_2019_1136_MOESM2_ESM.pdf
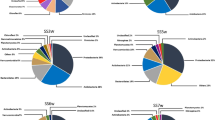
Supplementary Figure 2: ESM 2. Figure. Relative abundances (%) of prokaryotic phyla at different sampling sites of São Domingos mining area during (a) the winter and (b) the summer seasons. “Others” represents the sum of all taxa that scored relative abundances below 1% (PDF 319 kb)
Rights and permissions
About this article
Cite this article
Ettamimi, S., Carlier, J.D., Cox, C.J. et al. A meta-taxonomic investigation of the prokaryotic diversity of water bodies impacted by acid mine drainage from the São Domingos mine in southern Portugal. Extremophiles 23, 821–834 (2019). https://doi.org/10.1007/s00792-019-01136-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-019-01136-1