Abstract
Angiogenesis is a biological process finely tuned by a plethora of pro- and anti-angiogenic molecules, among which vascular endothelial growth factors (VEGFs). Their biological activity is expressed through the interaction with three cognate receptor tyrosine kinases, VEGFR1, 2, and 3. VEGFR2 is the primary regulator of angiogenesis. Ligand-induced VEGFR2 dimerization and activation depend on direct ligand binding to extracellular domains 2 and 3 of receptor and in the establishment of interactions between proximal membrane domains. VEGFR2 domain 7 has been shown to play a crucial role in receptor dimerization and regulation, therefore, representing a convenient target for the allosteric modulation of VEGFR2 activity. The ability to prepare a functional VEGFR2D7 domain represents the starting point to the development of novel VEGFR2 binders acting as allosteric inhibitors of receptor activity. Here, we describe a robust and efficient procedure for the preparation in E. coli of the VEGFR2 domain 7. The protein was obtained with a good yield and was properly folded. It was investigated in a biochemical and structural study, providing information on its conformational arrangement and in solution properties.









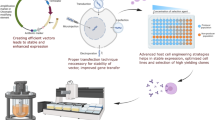
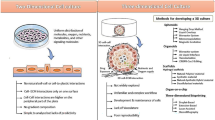
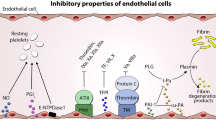
Similar content being viewed by others
Abbreviations
- VEGFs:
-
Vascular endothelial growth factors
- VEGFRs:
-
Vascular endothelial growth factor receptors
- RTKs:
-
Receptor tyrosine kinases
- TEMED:
-
Tetramethylethylenediamine
- APS:
-
Ammonium persulfate
- dNTPs:
-
Deoxynucleotide triphosphates
- IPTG:
-
Isopropyl β-d-1-thiogalactopyranoside
- O.D.600 :
-
Optical density at 600 nm
- Ni-NTA resin:
-
Nickel-charged nitrilotriacetic resin
- TEV protease:
-
Tobacco etch virus protease
- EDTA:
-
Ethylenediaminetetraacetic acid
- DTT:
-
Dithiothreitol
- RP-HPLC:
-
Reversed-phase high-performance liquid chromatography
- TFA:
-
Trifluoroacetic acid
- LC–MS:
-
Liquid chromatography–mass spectrometry
- Trp:
-
Tryptophan
- DSS:
-
4,4-Dimethyl-4-silapentane-1-sulfonic acid
References
Patel-Hett, S., & D’Amore, P. A. (2011). Signal transduction in vasculogenesis and developmental angiogenesis. The International Journal of Developmental Biology, 55, 353–363.
Ribatti, D. (2008). Judah Folkman, a pioneer in the study of angiogenesis. Angiogenesis, 11, 3–10.
Carmeliet, P. (2003). Angiogenesis in health and disease. Nature Medicine, 9, 653–660.
Shibuya, M. (2014). VEGF-VEGFR signals in health and disease. Biomolecules & Therapeutics, 22, 1–9.
Olsson, A. K., Dimberg, A., Kreuger, J., & Claesson-Welsh, L. (2006). VEGF receptor signalling-in control of vascular function. Nature Reviews Molecular Cell Biology, 7, 359–371.
Gerhardt, H., Golding, M., Fruttiger, M., Ruhrberg, C., Lundkvist, A., Abramsson, A., et al. (2003). VEGF guides angiogenic sprouting utilizing endothelial tip cell filopodia. The Journal of Cell Biology, 161, 1163–1177.
Takahashi, H., & Shibuya, M. (2005). The vascular endothelial growth factor (VEGF)/VEGF receptor system and its role under physiological and pathological conditions. Clinical Science, 109, 227–241.
Lania, G., Ferrentino, R., & Baldini, A. (2015). TBX1 represses Vegfr2 gene expression and enhances the cardiac fate of VEGFR2+ cells. PLoS ONE, 10, e0138525.
Smith, N. R., Baker, D., James, N. H., Ratcliffe, K., Jenkins, M., Ashton, S. E., et al. (2010). Vascular endothelial growth factor receptors VEGFR-2 and VEGFR-3 are localized primarily to the vasculature in human primary solid cancers. Clinical Cancer Research : An Official Journal of the American Association for Cancer Research, 16, 3548–3561.
Yamagishi, N., Teshima-Kondo, S., Masuda, K., Nishida, K., Kuwano, Y., Dang, D. T., et al. (2013). Chronic inhibition of tumor cell-derived VEGF enhances the malignant phenotype of colorectal cancer cells. BMC Cancer, 13, 229.
Chatterjee, S., Heukamp, L. C., Siobal, M., Schottle, J., Wieczorek, C., Peifer, M., et al. (2013). Tumor VEGF:VEGFR2 autocrine feed-forward loop triggers angiogenesis in lung cancer. The Journal of Clinical Investigation, 123, 1732–1740.
Ruch, C., Skiniotis, G., Steinmetz, M. O., Walz, T., & Ballmer-Hofer, K. (2007). Structure of a VEGF-VEGF receptor complex determined by electron microscopy. Nature Structural & Molecular Biology, 14, 249–250.
Kisko, K., Brozzo, M. S., Missimer, J., Schleier, T., Menzel, A., Leppanen, V. M., et al. (2011). Structural analysis of vascular endothelial growth factor receptor-2/ligand complexes by small-angle X-ray solution scattering. FASEB Journal : Official Publication of the Federation of American Societies for Experimental Biology, 25, 2980–2986.
Shinkai, A., Ito, M., Anazawa, H., Yamaguchi, S., Shitara, K., & Shibuya, M. (1998). Mapping of the sites involved in ligand association and dissociation at the extracellular domain of the kinase insert domain-containing receptor for vascular endothelial growth factor. The Journal of Biological Chemistry, 273, 31283–31288.
Dosch, D. D., & Ballmer-Hofer, K. (2010). Transmembrane domain-mediated orientation of receptor monomers in active VEGFR-2 dimers. FASEB Journal : Official Publication of the Federation of American Societies for Experimental Biology, 24, 32–38.
Brozzo, M. S., Bjelic, S., Kisko, K., Schleier, T., Leppanen, V. M., Alitalo, K., et al. (2012). Thermodynamic and structural description of allosterically regulated VEGFR-2 dimerization. Blood, 119, 1781–1788.
Hyde, C. A., Giese, A., Stuttfeld, E., Abram, Saliba J., Villemagne, D., Schleier, T., et al. (2012). Targeting extracellular domains D4 and D7 of vascular endothelial growth factor receptor 2 reveals allosteric receptor regulatory sites. Molecular and Cellular Biology, 32, 3802–3813.
Yang, Y., Xie, P., Opatowsky, Y., & Schlessinger, J. (2010). Direct contacts between extracellular membrane-proximal domains are required for VEGF receptor activation and cell signaling. Proceedings of the National academy of Sciences of the United States of America, 107, 1906–1911.
Sarabipour, S., Ballmer-Hofer, K., & Hristova, K. (2016). VEGFR-2 conformational switch in response to ligand binding. eLife, 5, e13876.
Thieltges, K. M., Avramovic, D., Piscitelli, C. L., Markovic-Mueller, S., Binz, H. K., & Ballmer-Hofer, K. (2018). Characterization of a drug-targetable allosteric site regulating vascular endothelial growth factor signaling. Angiogenesis, 21, 533–543.
Ellis L. M. (2005). Bevacizumab. Nature Reviews Drug Discovery, Suppl S8–9.
Krupitskaya, Y., & Wakelee, H. A. (2009). Ramucirumab a fully human mAb to the transmembrane signaling tyrosine kinase VEGFR-2 for the potential treatment of cancer. Current Opinion in Investigational Drugs, 10, 597–605.
D’Andrea, L. D., Del Gatto, A., De Rosa, L., Romanelli, A., & Pedone, C. (2009). Peptides targeting angiogenesis related growth factor receptors. Current Pharmaceutical Design, 15, 2414–2429.
Feng, S., Zou, L., Ni, Q., Zhang, X., Li, Q., Zheng, L., et al. (2014). Modulation, bioinformatic screening, and assessment of small molecular peptides targeting the vascular endothelial growth factor receptor. Cell Biochemistry and Biophysics, 70, 1913–1921.
Di Stasi, R., De Rosa, L., Romanelli, A., & D’Andrea, L. D. (2016). Peptides interacting with growth factor receptors regulating angiogenesis. Frontiers in Medicinal Chemistry, 9, 103–160.
De Rosa, L., Di Stasi, R., & D’Andrea, L. D. (2018). Pro-angiogenic peptides in biomedicine. Archives of Biochemistry and Biophysics, 660, 72–86.
D’Andrea, L. D., Romanelli, A., Di Stasi, R., & Pedone, C. (2010). Bioinorganic aspects of angiogenesis. Dalton Transactions, 39, 7625–7636.
D’Andrea, L. D., De Rosa, L., Vigliotti, C., & Cataldi, M. (2017). VEGF mimic peptides: Potential applications in central nervous system therapeutics. New Horizons in Translational Medicine, 3, 233–251.
Mendel, D. B., Laird, A. D., Xin, X., Louie, S. G., Christensen, J. G., Li, G., et al. (2003). In vivo antitumor activity of SU11248, a novel tyrosine kinase inhibitor targeting vascular endothelial growth factor and platelet-derived growth factor receptors: Determination of a pharmacokinetic/pharmacodynamic relationship. Clinical Cancer Research: An Official Journal of the American Association for Cancer Research, 9, 327–337.
Kendrew, J., Eberlein, C., Hedberg, B., McDaid, K., Smith, N. R., Weir, H. M., et al. (2011). An antibody targeted to VEGFR-2 Ig domains 4–7 inhibits VEGFR-2 activation and VEGFR-2-dependent angiogenesis without affecting ligand binding. Molecular Cancer Therapeutics, 10, 770–783.
Di Stasi, R., De Rosa, L., Diana, D., Fattorusso, R., & D’Andrea, L. D. (2019). Human recombinant VEGFR2D4 biochemical characterization to investigate novel anti-VEGFR2D4 antibodies for allosteric targeting of VEGFR2. Molecular Biotechnology, 61, 513–520.
Keunen, O., Johansson, M., Oudin, A., Sanzey, M., Rahim, S. A., Fack, F., et al. (2011). Anti-VEGF treatment reduces blood supply and increases tumor cell invasion in glioblastoma. Proceedings of the National academy of Sciences of the United States of America, 108, 3749–3754.
Chung, A. S., Kowanetz, M., Wu, X., Zhuang, G., Ngu, H., Finkle, D., et al. (2012). Differential drug class-specific metastatic effects following treatment with a panel of angiogenesis inhibitors. The Journal of Pathology, 227, 404–416.
Di Stasi, R., Diana, D., Capasso, D., Palumbo, R., Romanelli, A., Pedone, C., et al. (2010). VEGFR1(D2) in drug discovery: Expression and molecular characterization. Biopolymers, 94, 800–809.
Provencher, S. W., & Glockner, J. (1981). Estimation of globular protein secondary structure from circular dichroism. Biochemistry, 20, 33–37.
Manavalan, P., & Johnson, W. C., Jr. (1987). Variable selection method improves the prediction of protein secondary structure from circular dichroism spectra. Analytical Biochemistry, 167, 76–85.
Sreerama, N., & Woody, R. W. (2000). Estimation of protein secondary structure from circular dichroism spectra: Comparison of CONTIN, SELCON, and CDSSTR methods with an expanded reference set. Analytical Biochemistry, 287, 252–260.
Hwang, T. L., & Shaka, A. J. (1995). Water suppression that works. Excitation sculpting using arbitrary wave-forms and pulsed-field gradients. Journal of Magnetic Resonance Series A, 112, 275–279.
Dalvit, C. (1998). Efficient multiple-solvent suppression for the study of the interactions of organic solvents with biomolecules. Journal of Biomolecular NMR, 11, 437–444.
Goddard, T. D., & Kneller, D. G. (2004). SPARKY 3. California: University of San Francisco.
Keller, R. L. J. (2004). The computer aided resonance assignement tutorial. Newport Beach: CANTINA.
Morris, K. F., & Jr, Johnson C. S. (1992). Diffusion-ordered two-dimensional nuclear magnetic resonance spectroscopy. Journal of the American Chemical Society, 114, 3139–3141.
Price, W. S., Nara, M., & Arata, Y. (1997). A pulsed field gradient NMR study of the aggregation and hydration of parvalbumin. Biophysical Chemistry, 65, 179–187.
Garcia de la Torre, J., Huertas, M. L., & Carrasco, B. (2000). HYDRONMR: Prediction of NMR relaxation of globular proteins from atomic-level structures and hydrodynamic calculations. Journal of Magnetic Resonance, 147, 138–146.
Chemes, L. B., Alonso, L. G., Noval, M. G., & de Prat-Gay, G. (2012). Circular dichroism techniques for the analysis of intrinsically disordered proteins and domains. Methods in Molecular Biology, 895, 387–404.
Greenfield, N. J. (2006). Using circular dichroism spectra to estimate protein secondary structure. Nature Protocols, 1, 2876–2890.
Chen, Y., & Barkley, M. D. (1998). Toward understanding tryptophan fluorescence in proteins. Biochemistry, 37, 9976–9982.
Yang, Y., Zhang, Y., Cao, Z., Ji, H., Yang, X., Iwamoto, H., et al. (2013). Anti-VEGF- and anti-VEGF receptor-induced vascular alteration in mouse healthy tissues. Proceedings of the National academy of Sciences of the United States of America, 110, 12018–12023.
Gautier, B., Goncalves, V., Diana, D., Di Stasi, R., Teillet, F., Lenoir, C., et al. (2010). Biochemical and structural analysis of the binding determinants of a vascular endothelial growth factor receptor peptidic antagonist. Journal of Medicinal Chemistry, 53, 4428–4440.
Basile, A., Del Gatto, A., Diana, D., Di Stasi, R., Falco, A., Festa, M., et al. (2011). Characterization of a designed vascular endothelial growth factor receptor antagonist helical peptide with antiangiogenic activity in vivo. Journal of Medicinal Chemistry, 54, 1391–1400.
De Rosa, L., Diana, D., Basile, A., Russomanno, A., Isernia, C., Turco, M. C., et al. (2014). Design, structural and biological characterization of a VEGF inhibitor beta-hairpin-constrained peptide. European Journal of Medicinal Chemistry, 73, 210–216.
Wang, L., Coric, P., Broussy, S., Di Stasi, R., Zhou, L., D’Andrea, L. D., et al. (2019). Structural studies of the binding of an antagonistic cyclic peptide to the VEGFR1 domain 2. European Journal of Medicinal Chemistry, 169, 65–75.
Ruegg, C., Hasmim, M., Lejeune, F. J., & Alghisi, G. C. (2006). Antiangiogenic peptides and proteins: From experimental tools to clinical drugs. Biochimica et Biophysica Acta, 1765, 155–177.
Sulochana, K. N., & Ge, R. (2007). Developing antiangiogenic peptide drugs for angiogenesis-related diseases. Current Pharmaceutical Design, 13, 2074–2086.
Wilkins, D. K., Grimshaw, S. B., Receveur, V., Dobson, C. M., Jones, J. A., & Smith, L. J. (1999). Hydrodynamic radii of native and denatured proteins measured by pulse field gradient NMR techniques. Biochemistry, 38, 16424–16431.
Clarke, J., Cota, E., Fowler, S. B., & Hamill, S. J. (1999). Folding studies of immunoglobulin-like beta-sandwich proteins suggest that they share a common folding pathway. Structure, 7, 1145–1153.
Acknowledgements
LDR is supported by Fondazione Umberto Veronesi-Post-Doctoral Fellowship 2019. We would like to thank Mr Leopoldo Zona for technical assistance and Dr Luigi Russo for skillful help with HYDROPRO.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Di Stasi, R., Diana, D., De Rosa, L. et al. Biochemical and Conformational Characterization of Recombinant VEGFR2 Domain 7. Mol Biotechnol 61, 860–872 (2019). https://doi.org/10.1007/s12033-019-00211-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-019-00211-4