Abstract
Biological membranes display a staggering complexity of lipids and proteins orchestrating cellular functions. Superior analytical tools coupled with numerous functional cellular screens have enabled us to query their role in cellular signalling, trafficking, guiding protein structure and function–all of which rely on the dynamic membrane lipid properties indispensable for proper cellular functions. Alteration of these has led to emergence of various pathological conditions, thus opening an area of lipid-centric therapeutic approaches. This perspective is a short summary of the dynamic properties of membranes essential for proper cellular functions, dictating both protein and lipid functions, and mis-regulated in diseases. Towards the end, we focus on some challenges lying ahead and potential means to tackle the same, mainly underscored by multi-disciplinary approaches.
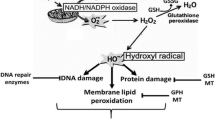
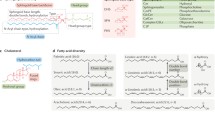
Graphic Abstract


adapted in part with permission from (Bigay and Antonny 2012)

Similar content being viewed by others
Abbreviations
- lo :
-
Liquid ordered domains
- lLd :
-
Liquid disordered domains/phase
- PALM:
-
Photoactivated localization microscopy
- STED:
-
Stimulated emission depletion
- NSOM:
-
Near-field scanning optical microscopy
- FRET:
-
Förster resonance energy transfer
- iSCAT:
-
Interferometric scattering microscopy
- SANS:
-
Small-angle neutron scattering
- SPT:
-
Single particle tracking
- FCS:
-
Fluorescence correlation spectroscopy
- svFCS:
-
Spot variation fluorescence correlation spectroscopy
- GUVs:
-
Giant unilamellar vesicles
- AFM:
-
Atomic force microscopy
- PH:
-
Pleckstrin homology
- COPI:
-
Coat protein I
- SM:
-
Sphingomyelin
- GPCR:
-
G protein-coupled receptor
- GTP:
-
Guanosine triphosphate
- PIP2:
-
Phosphatidyl inositol 4, 5 bisphosphate
- IgE:
-
Immunoglobulin E
- FCϵRI:
-
High affinity IgE receptor
- PS:
-
Phosphatidyl serine
- PE:
-
Phosphatidyl ethanolamine
- CD44:
-
Cluster of differentiation 44
- CD24:
-
Cluster of differentiation 24
- EGFR:
-
Epidermal growth factor receptor
- PI:
-
Phosphatidyl inositols
- Arp 2/3:
-
Actin related proteins 2/3
- WASP:
-
Wiskott–Aldrich syndrome protein
- TDM:
-
Trehalose Dimycolate
- MLT:
-
Membrane lipid therapy
- PI3K:
-
Phosphoinositide 3 kinase
- CDC42:
-
Cell division control protein 42
- RAC1:
-
Ras-related C3 botulinum toxin substrate 1
- Caspase:
-
Cysteine-aspartic proteases
- FADD:
-
Fas-associated death domain-containing protein
- DISC:
-
Death-inducing signalling complex
- PPARα:
-
Peroxisome proliferator-activated receptor alpha
- DAG:
-
Diacyl glycerol
References
Abankwa D, Gorfe AA, Hancock JF (2007) Ras nanoclusters: molecular structure and assembly. Semin Cell Dev Biol 18:599–607. https://doi.org/10.1016/j.semcdb.2007.08.003
Agrawal AG, Somani RR (2009) Farnesyltransferase inhibitor as anticancer agent. Mini Rev Med Chem 9:638–652
Ando J et al (2015) Sphingomyelin distribution in lipid rafts of artificial monolayer membranes visualized by Raman microscopy. Proc Natl Acad Sci USA 112:4558–4563. https://doi.org/10.1073/pnas.1418088112
Antonny B (2011) Mechanisms of membrane curvature sensing. Annu Rev Biochem 80:101–123. https://doi.org/10.1146/annurev-biochem-052809-155121
Atilla-Gokcumen GE et al (2014) Dividing cells regulate their lipid composition and localization. Cell 156:428–439. https://doi.org/10.1016/j.cell.2013.12.015
Babu M et al (2012) Interaction landscape of membrane-protein complexes in Saccharomyces cerevisiae. Nature 489:585–589. https://doi.org/10.1038/nature11354
Bacia K, Scherfeld D, Kahya N, Schwille P (2004) Fluorescence correlation spectroscopy relates rafts in model and native membranes. Biophys J 87:1034–1043. https://doi.org/10.1529/biophysj.104.040519
Barcelo-Coblijn G et al (2011) Sphingomyelin and sphingomyelin synthase (SMS) in the malignant transformation of glioma cells and in 2-hydroxyoleic acid therapy. Proc Natl Acad Sci USA 108:19569–19574. https://doi.org/10.1073/pnas.1115484108
Barrera NP, Zhou M, Robinson CV (2013) The role of lipids in defining membrane protein interactions: insights from mass spectrometry. Trends Cell Biol 23:1–8. https://doi.org/10.1016/j.tcb.2012.08.007
Berger EA (1997) HIV entry and tropism: the chemokine receptor connection. AIDS 11(Suppl A):S3–S16
Bigay J, Antonny B (2012) Curvature, lipid packing, and electrostatics of membrane organelles: defining cellular territories in determining specificity. Dev Cell 23:886–895. https://doi.org/10.1016/j.devcel.2012.10.009
Brown DA, Rose JK (1992) Sorting of GPI-anchored proteins to glycolipid-enriched membrane subdomains during transport to the apical cell surface. Cell 68:533–544
Chandra A et al (2011) The GDI-like solubilizing factor PDEdelta sustains the spatial organization and signalling of Ras family proteins. Nat Cell Biol 14:148–158. https://doi.org/10.1038/ncb2394
Chattopadhyay A, Jafurulla M (2012) Role of membrane cholesterol in leishmanial infection. Adv Exp Med Biol 749:201–213. https://doi.org/10.1007/978-1-4614-3381-1_14
Contreras FX et al (2012) Molecular recognition of a single sphingolipid species by a protein’s transmembrane domain. Nature 481:525–529. https://doi.org/10.1038/nature10742
Coskun U, Simons K (2011) Cell membranes: the lipid perspective. Structure 19:1543–1548. https://doi.org/10.1016/j.str.2011.10.010
Cywes C, Wessels MR (2001) Group A Streptococcus tissue invasion by CD44-mediated cell signalling. Nature 414:648–652. https://doi.org/10.1038/414648a
Dadhich R, Singh A, Menon AP, Mishra M, Athul CD, Kapoor S (2019) Biophysical characterization of mycobacterial model membranes and their interaction with rifabutin: towards lipid-guided drug screening in tuberculosis. Biochim Biophys Acta Biomembr 1861:1213–1227. https://doi.org/10.1016/j.bbamem.2019.04.004
de Wit G, Danial JS, Kukura P, Wallace MI (2015) Dynamic label-free imaging of lipid nanodomains. Proc Natl Acad Sci USA 112:12299–12303. https://doi.org/10.1073/pnas.1508483112
Dehio C, Meyer M, Berger J, Schwarz H, Lanz C (1997) Interaction of Bartonella henselae with endothelial cells results in bacterial aggregation on the cell surface and the subsequent engulfment and internalisation of the bacterial aggregate by a unique structure, the invasome. J Cell Sci 110(Pt 18):2141–2154
Dinic J, Ashrafzadeh P, Parmryd I (2013) Actin filaments attachment at the plasma membrane in live cells cause the formation of ordered lipid domains. Biochim Biophys Acta 1828:1102–1111. https://doi.org/10.1016/j.bbamem.2012.12.004
Doria ML, Cotrim CZ, Simoes C, Macedo B, Domingues P, Domingues MR, Helguero LA (2013) Lipidomic analysis of phospholipids from human mammary epithelial and breast cancer cell lines. J Cell Physiol 228:457–468. https://doi.org/10.1002/jcp.24152
Earl PL, Koenig S, Moss B (1991) Biological and immunological properties of human immunodeficiency virus type 1 envelope glycoprotein: analysis of proteins with truncations and deletions expressed by recombinant vaccinia viruses. J Virol 65:31–41
Eggeling C et al (2009) Direct observation of the nanoscale dynamics of membrane lipids in a living cell. Nature 457:1159–1162. https://doi.org/10.1038/nature07596
Emoto K, Toyama-Sorimachi N, Karasuyama H, Inoue K, Umeda M (1997) Exposure of phosphatidylethanolamine on the surface of apoptotic cells. Exp Cell Res 232:430–434. https://doi.org/10.1006/excr.1997.3521
Erazo T et al (2016) The new antitumor drug ABTL0812 inhibits the Akt/mTORC1 axis by upregulating tribbles-3 pseudokinase. Clin Cancer Res 22:2508–2519. https://doi.org/10.1158/1078-0432.CCR-15-1808
Escriba PV (2017) Membrane-lipid therapy: a historical perspective of membrane-targeted therapies - From lipid bilayer structure to the pathophysiological regulation of cells. Biochim Biophys Acta Biomembr 1859:1493–1506. https://doi.org/10.1016/j.bbamem.2017.05.017
Filipp D, Leung BL, Zhang J, Veillette A, Julius M (2004) Enrichment of lck in lipid rafts regulates colocalized fyn activation and the initiation of proximal signals through TCR alpha beta. J Immunol 172:4266–4274. https://doi.org/10.4049/jimmunol.172.7.4266
Finnegan CM, Rawat SS, Puri A, Wang JM, Ruscetti FW, Blumenthal R (2004) Ceramide, a target for antiretroviral therapy. Proc Natl Acad Sci USA 101:15452–15457. https://doi.org/10.1073/pnas.0402874101
Finnegan CM, Rawat SS, Cho EH, Guiffre DL, Lockett S, Merrill AH Jr, Blumenthal R (2007) Sphingomyelinase restricts the lateral diffusion of CD4 and inhibits human immunodeficiency virus fusion. J Virol 81:5294–5304. https://doi.org/10.1128/JVI.02553-06
Flieger A, Frischknecht F, Hacker G, Hornef MW, Pradel G (2018) Pathways of host cell exit by intracellular pathogens. Microb Cell 5:525–544. https://doi.org/10.15698/mic2018.12.659
Friedrichson T, Kurzchalia TV (1998) Microdomains of GPI-anchored proteins in living cells revealed by crosslinking. Nature 394:802–805. https://doi.org/10.1038/29570
Gajate C, Mollinedo F (2007) Edelfosine and perifosine induce selective apoptosis in multiple myeloma by recruitment of death receptors and downstream signaling molecules into lipid rafts. Blood 109:711–719. https://doi.org/10.1182/blood-2006-04-016824
Garcia-Saez AJ, Chiantia S, Schwille P (2007) Effect of line tension on the lateral organization of lipid membranes. J Biol Chem 282:33537–33544. https://doi.org/10.1074/jbc.M706162200
Gaus K, Gratton E, Kable EP, Jones AS, Gelissen I, Kritharides L, Jessup W (2003) Visualizing lipid structure and raft domains in living cells with two-photon microscopy. Proc Natl Acad Sci USA 100:15554–15559. https://doi.org/10.1073/pnas.2534386100
Ghosh S, Saha S, Goswami D, Bilgrami S, Mayor S (2012) Dynamic imaging of homo-FRET in live cells by fluorescence anisotropy microscopy. Methods Enzymol 505:291–327. https://doi.org/10.1016/B978-0-12-388448-0.00024-3
Grassme H, Jendrossek V, Bock J, Riehle A, Gulbins E (2002) Ceramide-rich membrane rafts mediate CD40 clustering. J Immunol 168:298–307. https://doi.org/10.4049/jimmunol.168.1.298
Guardiola-Serrano F et al (2015) The novel anticancer drug hydroxytriolein inhibits lung cancer cell proliferation via a protein kinase calpha- and extracellular signal-regulated kinase 1/2-dependent mechanism. J Pharmacol Exp Ther 354:213–224. https://doi.org/10.1124/jpet.114.222281
Hammond GR, Balla T (2015) Polyphosphoinositide binding domains: key to inositol lipid biology. Biochim Biophys Acta 1851:746–758. https://doi.org/10.1016/j.bbalip.2015.02.013
Heinemann F, Vogel SK, Schwille P (2013) Lateral membrane diffusion modulated by a minimal actin cortex. Biophys J 104:1465–1475. https://doi.org/10.1016/j.bpj.2013.02.042
Honigmann A, Mueller V, Ta H, Schoenle A, Sezgin E, Hell SW, Eggeling C (2014) Scanning STED-FCS reveals spatiotemporal heterogeneity of lipid interaction in the plasma membrane of living cells. Nat Commun 5:5412. https://doi.org/10.1038/ncomms6412
Huang Q, Shen HM, Ong CN (2005) Emodin inhibits tumor cell migration through suppression of the phosphatidylinositol 3-kinase-Cdc42/Rac1 pathway. Cell Mol Life Sci 62:1167–1175. https://doi.org/10.1007/s00018-005-5050-2
Kahya N, Scherfeld D, Bacia K, Poolman B, Schwille P (2003) Probing lipid mobility of raft-exhibiting model membranes by fluorescence correlation spectroscopy. J Biol Chem 278:28109–28115. https://doi.org/10.1074/jbc.M302969200
Kapoor S et al (2011) Temperature-pressure phase diagram of a heterogeneous anionic model biomembrane system: results from a combined calorimetry, spectroscopy and microscopy study. Biochim Biophys Acta 1808:1187–1195. https://doi.org/10.1016/j.bbamem.2011.01.011
Kapoor S, Triola G, Vetter IR, Erlkamp M, Waldmann H, Winter R (2012a) Revealing conformational substates of lipidated N-Ras protein by pressure modulation. Proc Natl Acad Sci USA 109:460–465. https://doi.org/10.1073/pnas.1110553109
Kapoor S, Weise K, Erlkamp M, Triola G, Waldmann H, Winter R (2012b) The role of G-domain orientation and nucleotide state on the Ras isoform-specific membrane interaction. Eur Biophys J 41:801–813. https://doi.org/10.1007/s00249-012-0841-5
Khanzada UK, Pardo OE, Meier C, Downward J, Seckl MJ, Arcaro A (2006) Potent inhibition of small-cell lung cancer cell growth by simvastatin reveals selective functions of Ras isoforms in growth factor signalling. Oncogene 25:877–887. https://doi.org/10.1038/sj.onc.1209117
Koldso H, Reddy T, Fowler PW, Duncan AL, Sansom MS (2016) Membrane compartmentalization reducing the mobility of lipids and proteins within a model plasma membrane. J Phys Chem B 120:8873–8881. https://doi.org/10.1021/acs.jpcb.6b05846
Krivanek R, Okoro L, Winter R (2008) Effect of cholesterol and ergosterol on the compressibility and volume fluctuations of phospholipid-sterol bilayers in the critical point region: a molecular acoustic and calorimetric study. Biophys J 94:3538–3548. https://doi.org/10.1529/biophysj.107.122549
Kumar GA, Jafurulla M, Chattopadhyay A (2016) The membrane as the gatekeeper of infection: cholesterol in host-pathogen interaction. Chem Phys Lipids 199:179–185. https://doi.org/10.1016/j.chemphyslip.2016.02.007
Kusumi A et al (2005) Paradigm shift of the plasma membrane concept from the two-dimensional continuum fluid to the partitioned fluid: high-speed single-molecule tracking of membrane molecules. Annu Rev Biophys Biomol Struct 34:351–378. https://doi.org/10.1146/annurev.biophys.34.040204.144637
Lemmon MA (2008) Membrane recognition by phospholipid-binding domains. Nat Rev Mol Cell Biol 9:99–111. https://doi.org/10.1038/nrm2328
Lingwood D et al (2011) Cholesterol modulates glycolipid conformation and receptor activity. Nat Chem Biol 7:260–262. https://doi.org/10.1038/nchembio.551
Lozano MM, Liu Z, Sunnick E, Janshoff A, Kumar K, Boxer SG (2013) Colocalization of the ganglioside G(M1) and cholesterol detected by secondary ion mass spectrometry. J Am Chem Soc 135:5620–5630. https://doi.org/10.1021/ja310831m
Manes S, del Real G, Martinez AC (2003) Pathogens: raft hijackers. Nat Rev Immunol 3:557–568. https://doi.org/10.1038/nri1129
Marki F, Hanni E, Fredenhagen A, van Oostrum J (1991) Mode of action of the lanthionine-containing peptide antibiotics duramycin, duramycin B and C, and cinnamycin as indirect inhibitors of phospholipase A2. Biochem Pharmacol 42:2027–2035
Martin-Gago P et al (2017) A PDE6delta-KRas inhibitor chemotype with up to seven h-bonds and picomolar affinity that prevents efficient inhibitor release by Arl2. Angew Chem Int Ed Engl 56:2423–2428. https://doi.org/10.1002/anie.201610957
McLaughlin PJ, Gooch JT, Mannherz HG, Weeds AG (1993) Structure of gelsolin segment 1-actin complex and the mechanism of filament severing. Nature 364:685–692. https://doi.org/10.1038/364685a0
Menendez JA, Lupu R (2007) Fatty acid synthase and the lipogenic phenotype in cancer pathogenesis. Nat Rev Cancer 7:763–777. https://doi.org/10.1038/nrc2222
Mikaty G et al (2009) Extracellular bacterial pathogen induces host cell surface reorganization to resist shear stress. PLoS Pathog 5:e1000314. https://doi.org/10.1371/journal.ppat.1000314
Mollinedo F, Gajate C (2015) Lipid rafts as major platforms for signaling regulation in cancer. Adv Biol Regul 57:130–146. https://doi.org/10.1016/j.jbior.2014.10.003
Moon S, Yan R, Kenny SJ, Shyu Y, Xiang L, Li W, Xu K (2017) Spectrally resolved, functional super-resolution microscopy reveals nanoscale compositional heterogeneity in live-cell membranes. J Am Chem Soc 139:10944–10947. https://doi.org/10.1021/jacs.7b03846
Mylvaganam SM, Grinstein S, Freeman SA (2018) Picket-fences in the plasma membrane: functions in immune cells and phagocytosis. Semin Immunopathol 40:605–615. https://doi.org/10.1007/s00281-018-0705-x
Nitenberg M et al (2018) The potent effect of mycolactone on lipid membranes. PLoS Pathog 14:e1006814. https://doi.org/10.1371/journal.ppat.1006814
Ortega-Arroyo J, Kukura P (2012) Interferometric scattering microscopy (iSCAT): new frontiers in ultrafast and ultrasensitive optical microscopy. Phys Chem Chem Phys 14:15625–15636. https://doi.org/10.1039/c2cp41013c
Overington JP, Al-Lazikani B, Hopkins AL (2006) How many drug targets are there? Nat Rev Drug Discov 5:993–996. https://doi.org/10.1038/nrd2199
Owen DM, Williamson D, Magenau A, Gaus K (2012a) Optical techniques for imaging membrane domains in live cells (live-cell palm of protein clustering). Methods Enzymol 504:221–235. https://doi.org/10.1016/B978-0-12-391857-4.00011-2
Owen DM, Williamson DJ, Magenau A, Gaus K (2012b) Sub-resolution lipid domains exist in the plasma membrane and regulate protein diffusion and distribution. Nat Commun 3:1256. https://doi.org/10.1038/ncomms2273
Parasassi T, Gratton E, Yu WM, Wilson P, Levi M (1997) Two-photon fluorescence microscopy of laurdan generalized polarization domains in model and natural membranes. Biophys J 72:2413–2429. https://doi.org/10.1016/S0006-3495(97)78887-8
Pathak P, London E (2015) The effect of membrane lipid composition on the formation of lipid ultrananodomains. Biophys J 109:1630–1638. https://doi.org/10.1016/j.bpj.2015.08.029
Pencer J, Mills TT, Kucerka N, Nieh MP, Katsaras J (2007) Small-angle neutron scattering to detect rafts and lipid domains. Methods Mol Biol 398:231–244. https://doi.org/10.1007/978-1-59745-513-8_16
Percherancier Y et al (2003) HIV-1 entry into T-cells is not dependent on CD4 and CCR71 localization to sphingolipid-enriched, detergent-resistant, raft membrane domains. J Biol Chem 278:3153–3161. https://doi.org/10.1074/jbc.M207371200
Pike LJ (2006) Rafts defined: a report on the Keystone Symposium on Lipid Rafts and Cell Function. J Lipid Res 47:1597–1598. https://doi.org/10.1194/jlr.E600002-JLR200
Pincet F, Adrien V, Yang R, Delacotte J, Rothman JE, Urbach W, Tareste D (2016) FRAP to characterize molecular diffusion and interaction in various membrane environments. PLoS ONE 11:e0158457. https://doi.org/10.1371/journal.pone.0158457
Plowman SJ, Muncke C, Parton RG, Hancock JF (2005) H-ras, K-ras, and inner plasma membrane raft proteins operate in nanoclusters with differential dependence on the actin cytoskeleton. Proc Natl Acad Sci USA 102:15500–15505. https://doi.org/10.1073/pnas.0504114102
Pralle A, Keller P, Florin EL, Simons K, Horber JK (2000) Sphingolipid-cholesterol rafts diffuse as small entities in the plasma membrane of mammalian cells. J Cell Biol 148:997–1008. https://doi.org/10.1083/jcb.148.5.997
Rawat SS et al (2004) Elevated expression of GM3 in receptor-bearing targets confers resistance to human immunodeficiency virus type 1 fusion. J Virol 78:7360–7368. https://doi.org/10.1128/JVI.78.14.7360-7368.2004
Rawat SS, Viard M, Gallo SA, Blumenthal R, Puri A (2006) Sphingolipids, cholesterol, and HIV-1: a paradigm in viral fusion. Glycoconj J 23:189–197. https://doi.org/10.1007/s10719-006-7924-4
Rawat SS, Zimmerman C, Johnson BT, Cho E, Lockett SJ, Blumenthal R, Puri A (2008) Restricted lateral mobility of plasma membrane CD4 impairs HIV-1 envelope glycoprotein mediated fusion. Mol Membr Biol 25:83–94. https://doi.org/10.1080/09687680701613713
Riethmuller J, Riehle A, Grassme H, Gulbins E (2006) Membrane rafts in host-pathogen interactions. Biochim Biophys Acta 1758:2139–2147. https://doi.org/10.1016/j.bbamem.2006.07.017
Rocks O et al (2005) An acylation cycle regulates localization and activity of palmitoylated Ras isoforms. Science 307:1746–1752. https://doi.org/10.1126/science.1105654
Rodriguez-Rivera FP, Zhou X, Theriot JA, Bertozzi CR (2017) Visualization of mycobacterial membrane dynamics in live cells. J Am Chem Soc 139:3488–3495. https://doi.org/10.1021/jacs.6b12541
Rotblat B, Prior IA, Muncke C, Parton RG, Kloog Y, Henis YI, Hancock JF (2004) Three separable domains regulate GTP-dependent association of H-ras with the plasma membrane. Mol Cell Biol 24:6799–6810. https://doi.org/10.1128/MCB.24.15.6799-6810.2004
Saha S, Raghupathy R, Mayor S (2015) Homo-FRET imaging highlights the nanoscale organization of cell surface molecules. Methods Mol Biol 1251:151–173. https://doi.org/10.1007/978-1-4939-2080-8_9
Saka SK, Honigmann A, Eggeling C, Hell SW, Lang T, Rizzoli SO (2014) Multi-protein assemblies underlie the mesoscale organization of the plasma membrane. Nat Commun 5:4509. https://doi.org/10.1038/ncomms5509
Saliba AE, Vonkova I, Gavin AC (2015) The systematic analysis of protein-lipid interactions comes of age. Nat Rev Mol Cell Biol 16:753–761. https://doi.org/10.1038/nrm4080
Sarkar-Banerjee S, Sayyed-Ahmad A, Prakash P, Cho KJ, Waxham MN, Hancock JF, Gorfe AA (2017) Spatiotemporal analysis of K-ras plasma membrane interactions reveals multiple high order homo-oligomeric complexes. J Am Chem Soc 139:13466–13475. https://doi.org/10.1021/jacs.7b06292
Scheidig AJ, Franken SM, Corrie JE, Reid GP, Wittinghofer A, Pai EF, Goody RS (1995) X-ray crystal structure analysis of the catalytic domain of the oncogene product p21H-ras complexed with caged GTP and mant dGppNHp. J Mol Biol 253:132–150. https://doi.org/10.1006/jmbi.1995.0541
Sengupta D, Chattopadhyay A (2015) Molecular dynamics simulations of GPCR-cholesterol interaction: an emerging paradigm. Biochim Biophys Acta 1848:1775–1782. https://doi.org/10.1016/j.bbamem.2015.03.018
Sengupta P, Jovanovic-Talisman T, Skoko D, Renz M, Veatch SL, Lippincott-Schwartz J (2011) Probing protein heterogeneity in the plasma membrane using PALM and pair correlation analysis. Nat Methods 8:969–975. https://doi.org/10.1038/nmeth.1704
Sezgin E, Schwille P (2011) Fluorescence techniques to study lipid dynamics. Cold Spring Harb Perspect Biol 3:a009803. https://doi.org/10.1101/cshperspect.a009803
Shen S, Yang L, Li L, Bai Y, Cai C, Liu H (2017) A plasma lipidomics strategy reveals perturbed lipid metabolic pathways and potential lipid biomarkers of human colorectal cancer. J Chromatogr B 1068–1069:41–48. https://doi.org/10.1016/j.jchromb.2017.10.004
Simons K, Ikonen E (1997) Functional rafts in cell membranes. Nature 387:569–572. https://doi.org/10.1038/42408
Simons K, Vaz WL (2004) Model systems, lipid rafts, and cell membranes. Annu Rev Biophys Biomol Struct 33:269–295. https://doi.org/10.1146/annurev.biophys.32.110601.141803
Sinensky M, Beck LA, Leonard S, Evans R (1990) Differential inhibitory effects of lovastatin on protein isoprenylation and sterol synthesis. J Biol Chem 265:19937–19941
Soubias O, Gawrisch K (2012) The role of the lipid matrix for structure and function of the GPCR rhodopsin. Biochim Biophys Acta 1818:234–240. https://doi.org/10.1016/j.bbamem.2011.08.034
Soubias O, Teague WE Jr, Hines KG, Gawrisch K (2015) Rhodopsin/lipid hydrophobic matching-rhodopsin oligomerization and function. Biophys J 108:1125–1132. https://doi.org/10.1016/j.bpj.2015.01.006
Spillane KM, Ortega-Arroyo J, de Wit G, Eggeling C, Ewers H, Wallace MI, Kukura P (2014) High-speed single-particle tracking of GM1 in model membranes reveals anomalous diffusion due to interleaflet coupling and molecular pinning. Nano Lett 14:5390–5397. https://doi.org/10.1021/nl502536u
Srivatsav AT, Mishra M, Kapoor S (2018) Small-molecule modulation of lipid-dependent cellular processes against cancer: fats on the gunpoint. Biomed Res Int 2018:6437371. https://doi.org/10.1155/2018/6437371
Steffens CM, Hope TJ (2004) Mobility of the human immunodeficiency virus (HIV) receptor CD4 and coreceptor CCR100 in living cells: implications for HIV fusion and entry events. J Virol 78:9573–9578. https://doi.org/10.1128/JVI.78.17.9573-9578.2004
Suzuki KG (2016) Single-molecule imaging of signal transduction via GPI-anchored receptors. Methods Mol Biol 1376:229–238. https://doi.org/10.1007/978-1-4939-3170-5_19
Teres S et al (2012) 2-Hydroxyoleate, a nontoxic membrane binding anticancer drug, induces glioma cell differentiation and autophagy. Proc Natl Acad Sci USA 109:8489–8494. https://doi.org/10.1073/pnas.1118349109
Urbancic I, Brun J, Shrestha D, Waithe D, Eggeling C, Chojnacki J (2018) Lipid composition but not curvature is the determinant factor for the low molecular mobility observed on the membrane of virus-like vesicles. Viruses 10(8):415. https://doi.org/10.3390/v10080415
van Zanten TS, Cambi A, Koopman M, Joosten B, Figdor CG, Garcia-Parajo MF (2009) Hotspots of GPI-anchored proteins and integrin nanoclusters function as nucleation sites for cell adhesion. Proc Natl Acad Sci USA 106:18557–18562. https://doi.org/10.1073/pnas.0905217106
Varshney P, Yadav V, Saini N (2016) Lipid rafts in immune signalling: current progress and future perspective. Immunology 149:13–24. https://doi.org/10.1111/imm.12617
Veatch SL, Keller SL (2003) Separation of liquid phases in giant vesicles of ternary mixtures of phospholipids and cholesterol. Biophys J 85:3074–3083. https://doi.org/10.1016/S0006-3495(03)74726-2
Visco I, Chiantia S, Schwille P (2014) Asymmetric supported lipid bilayer formation via methyl-beta-cyclodextrin mediated lipid exchange: influence of asymmetry on lipid dynamics and phase behavior. Langmuir 30:7475–7484. https://doi.org/10.1021/la500468r
Wang R et al (2013) Lipid raft regulates the initial spreading of melanoma A375 cells by modulating beta1 integrin clustering. Int J Biochem Cell Biol 45:1679–1689. https://doi.org/10.1016/j.biocel.2013.04.031
Wang YH, Bucki R, Janmey PA (2016) Cholesterol-dependent phase-demixing in lipid bilayers as a switch for the activity of the phosphoinositide-binding cytoskeletal protein gelsolin. Biochemistry 55:3361–3369. https://doi.org/10.1021/acs.biochem.5b01363
Wawrezinieck L, Rigneault H, Marguet D, Lenne PF (2005) Fluorescence correlation spectroscopy diffusion laws to probe the submicron cell membrane organization. Biophys J 89:4029–4042. https://doi.org/10.1529/biophysj.105.067959
Weise K, Triola G, Brunsveld L, Waldmann H, Winter R (2009) Influence of the lipidation motif on the partitioning and association of N-Ras in model membrane subdomains. J Am Chem Soc 131:1557–1564. https://doi.org/10.1021/ja808691r
Weise K et al (2011) Membrane-mediated induction and sorting of K-Ras microdomain signaling platforms. J Am Chem Soc 133:880–887. https://doi.org/10.1021/ja107532q
Weise K et al (2012) Dissociation of the K-Ras4B/PDEdelta complex upon contact with lipid membranes: membrane delivery instead of extraction. J Am Chem Soc 134:11503–11510. https://doi.org/10.1021/ja305518h
Wu H et al (2009) Effect of simvastatin on glioma cell proliferation, migration, and apoptosis. Neurosurgery 65:1087–1096. https://doi.org/10.1227/01.NEU.0000360130.52812.1D (discussion 1096–1087)
Yahi N, Aulas A, Fantini J (2010) How cholesterol constrains glycolipid conformation for optimal recognition of Alzheimer’s beta amyloid peptide (Abeta1-40). PLoS ONE 5:e9079. https://doi.org/10.1371/journal.pone.0009079
Yang L et al (2015) Comprehensive lipid profiling of plasma in patients with benign breast tumor and breast cancer reveals novel biomarkers. Anal Bioanal Chem 407:5065–5077. https://doi.org/10.1007/s00216-015-8484-x
Yu J, Fischman DA, Steck TL (1973) Selective solubilization of proteins and phospholipids from red blood cell membranes by nonionic detergents. J Supramol Struct 1:233–248. https://doi.org/10.1002/jss.400010308
Zalba S, Ten Hagen TL (2017) Cell membrane modulation as adjuvant in cancer therapy. Cancer Treat Rev 52:48–57. https://doi.org/10.1016/j.ctrv.2016.10.008
Zhang M et al (2005) CD45 signals outside of lipid rafts to promote ERK activation, synaptic raft clustering, and IL-2 production. J Immunol 174:1479–1490. https://doi.org/10.4049/jimmunol.174.3.1479
Zhou Y, Hancock JF (2018a) Deciphering lipid codes: K-Ras as a paradigm. Traffic 19:157–165. https://doi.org/10.1111/tra.12541
Zhou Y, Hancock JF (2018b) A novel prenyl-polybasic domain code determines lipid-binding specificity of the K-Ras membrane anchor. Small GTPases. https://doi.org/10.1080/21541248.2017.1379583
Zhou X et al (2012) Identification of plasma lipid biomarkers for prostate cancer by lipidomics and bioinformatics. PLoS ONE 7:e48889. https://doi.org/10.1371/journal.pone.0048889
Acknowledgements
We thank Dr. Rishikesh Narayan for comments on this manuscript. The work in our lab is supported by DBT-Ramalingaswami fellowship and DST Inspire Faculty Award awarded to SK. PA acknowledges DBT for PhD fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Research Involving Human and Animal Participants
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Adhyapak, P., Kapoor, S. Membrane Dynamics in Health and Disease: Impact on Cellular Signalling. J Membrane Biol 252, 213–226 (2019). https://doi.org/10.1007/s00232-019-00087-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-019-00087-0