Abstract
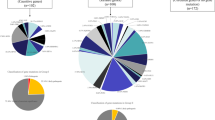
Rare missense variants play a crucial role in amyotrophic lateral sclerosis (ALS) pathophysiology. We report rare/novel missense variants from 154 Indian ALS patients, identified through targeted sequencing of 25 ALS-associated genes. As pathogenic variants could explain only a small percentage of ALS pathophysiology in our cohort, we investigated the frequency of tolerated and benign novel/rare variants, which could be potentially ALS susceptible. These variants were identified in 5.36% (8/149) of sporadic ALS (sALS) cases; with one novel variant each in ERBB4, SETX, DCTN1, and MATR3; four rare variants, one each in PON2 and ANG and two different rare variants in SETX. Identified variants were either absent or present at extremely rare frequencies (MAF < 0.01) in large population databases and were absent in 50 healthy controls sequenced through Sanger method. Furthermore, an oligogenic basis of ALS was observed in three sALS, with co-occurrence of intermediate-length repeat expansions in ATXN2 and a rare/novel variant in DCTN1 and SETX genes. Additionally, molecular dynamics and biochemical functional analysis of an angiogenin variant (R21G) identified from our cohort demonstrated loss of ribonucleolytic and nuclear translocation activities. Our findings suggest that rare variants could be potentially pathogenic and functional studies are warranted to decisively establish the pathogenic mechanisms associated with them.





Similar content being viewed by others
References
Renton AE, Chiò A, Traynor BJ (2014) State of play in amyotrophic lateral sclerosis genetics. Nat Neurosci
Al-Chalabi A, Hardiman O (2013) The epidemiology of ALS: a conspiracy of genes, environment and time. Nat Rev Neurol. https://doi.org/10.1038/nrneurol.2013.203
Cady J, Allred P, Bali T et al (2015) Amyotrophic lateral sclerosis onset is influenced by the burden of rare variants in known amyotrophic lateral sclerosis genes. Ann Neurol. https://doi.org/10.1002/ana.24306
Pang SYY, Hsu JS, Teo KC et al (2017) Burden of rare variants in ALS genes influences survival in familial and sporadic ALS. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2017.06.007
Krüger S, Battke F, Sprecher A et al (2016) Rare variants in neurodegeneration associated genes revealed by targeted panel sequencing in a German ALS cohort. Front Mol Neurosci. https://doi.org/10.3389/fnmol.2016.00092
Marangi G, Traynor BJ (2015) Genetic causes of amyotrophic lateral sclerosis: new genetic analysis methodologies entailing new opportunities and challenges. Brain Res
Wu D, Yu W, Kishikawa H et al (2007) Angiogenin loss-of-function mutations in amyotrophic lateral sclerosis. Ann Neurol. https://doi.org/10.1002/ana.21221
Padhi AK, Vasaikar SV, Jayaram B, Gomes J (2014) ANGDelMut—a web-based tool for predicting and analyzing functional loss mechanisms of amyotrophic lateral sclerosis-associated angiogenin mutations. F1000Research. https://doi.org/10.12688/f1000research.2-227.v3
Thiyagarajan N, Ferguson R, Subramanian V, Acharya KR (2012) Structural and molecular insights into the mechanism of action of human angiogenin-ALS variants in neurons. Nat Commun. https://doi.org/10.1038/ncomms2126
Boehringer A, Garcia-Mansfield K, Singh G et al (2017) ALS associated mutations in Matrin 3 alter protein-protein interactions and impede mRNA nuclear export. Sci Rep. https://doi.org/10.1038/s41598-017-14924-6
Marangi G, Lattante S, Doronzio PN et al (2017) Matrin 3 variants are frequent in Italian ALS patients. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2016.09.023
Padhi AK, Narain P, Dave U et al (2019) Insights into the role of ribonuclease 4 polymorphisms in amyotrophic lateral sclerosis. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2017.1419147
de Majo M, Topp SD, Smith BN et al (2018) ALS-associated missense and nonsense TBK1 mutations can both cause loss of kinase function. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2018.06.015
Higelin J, Catanese A, Semelink-Sedlacek LL et al (2018) NEK1 loss-of-function mutation induces DNA damage accumulation in ALS patient-derived motoneurons. Stem Cell Res. https://doi.org/10.1016/j.scr.2018.06.005
Padhi AK, Hazra S (2019) Insights into the role of d-amino acid oxidase mutations in amyotrophic lateral sclerosis. J Cell Biochem. https://doi.org/10.1002/jcb.27529
Narain P, Gomes J, Bhatia R et al (2017) C9orf72 hexanucleotide repeat expansions and Ataxin 2 intermediate length repeat expansions in Indian patients with amyotrophic lateral sclerosis. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2017.04.011
Narain P, Pandey A, Gupta S et al (2018) Targeted next-generation sequencing reveals novel and rare variants in Indian patients with amyotrophic lateral sclerosis. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2018.05.012
Ng PC, Henikoff S (2003) SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res
Adzhubei IA, Schmidt S, Peshkin L et al (2010) A method and server for predicting damaging missense mutations. Nat Methods. https://doi.org/10.1038/nmeth0410-248
Lek M, Karczewski KJ, Minikel EV et al (2016) Analysis of protein-coding genetic variation in 60,706 humans. Nature. https://doi.org/10.1038/nature19057
Padhi AK, Kumar H, Vasaikar SV et al (2012) Mechanisms of loss of functions of human angiogenin variants implicated in amyotrophic lateral sclerosis. PLoS One. https://doi.org/10.1371/journal.pone.0032479
Padhi AK, Banerjee K, Gomes J, Banerjee M (2014) Computational and functional characterization of angiogenin mutations, and correlation with amyotrophic lateral sclerosis. PLoS One. https://doi.org/10.1371/journal.pone.0111963
Al-Chalabi A, Van Den Berg LH, Veldink J (2017) Gene discovery in amyotrophic lateral sclerosis: implications for clinical management. Nat Rev Neurol
Sproviero W, Shatunov A, Stahl D et al (2017) ATXN2 trinucleotide repeat length correlates with risk of ALS. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2016.11.010
Abel O, Powell JF, Andersen PM, Al-Chalabi A (2012) ALSoD: a user-friendly online bioinformatics tool for amyotrophic lateral sclerosis genetics. Hum Mutat. https://doi.org/10.1002/humu.22157
Lin KP, Tsai PC, Liao YC et al (2015) Mutational analysis of MATR3 in Taiwanese patients with amyotrophic lateral sclerosis. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2015.02.008
Xu L, Li J, Tang L et al (2015) MATR3 mutation analysis in a Chinese cohort with sporadic amyotrophic lateral sclerosis. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2015.11.023
Leblond CS, Gan-Or Z, Spiegelman D et al (2016) Replication study of MATR3 in familial and sporadic amyotrophic lateral sclerosis. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2015.09.013
Brown RH, Al-Chalabi A (2017) Amyotrophic lateral sclerosis. Prog Med Chem. https://doi.org/10.1016/bs.pmch.2018.12.001
Couthouis J, Raphael AR, Daneshjou R, Gitler AD (2014) Targeted exon capture and sequencing in sporadic amyotrophic lateral sclerosis. PLoS Genet. https://doi.org/10.1371/journal.pgen.1004704
Kenna KP, McLaughlin RL, Byrne S et al (2013) Delineating the genetic heterogeneity of ALS using targeted high-throughput sequencing. J Med Genet. https://doi.org/10.1136/jmedgenet-2013-101795
Maruyama H, Morino H, Ito H et al (2010) Mutations of optineurin in amyotrophic lateral sclerosis. Nature. https://doi.org/10.1038/nature08971
Nakamura R, Sone J, Atsuta N et al (2016) Next-generation sequencing of 28 ALS-related genes in a Japanese ALS cohort. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2015.11.030
Takahashi Y, Fukuda Y, Yoshimura J et al (2013) Erbb4 mutations that disrupt the neuregulin-erbb4 pathway cause amyotrophic lateral sclerosis type 19. Am J Hum Genet. https://doi.org/10.1016/j.ajhg.2013.09.008
Zhang G, Li W, Li Z et al (2013) Association between paraoxonase gene and stroke in the Han Chinese population. BMC Med Genet. https://doi.org/10.1186/1471-2350-14-16
Valdmanis PN, Kabashi E, Dyck A et al (2008) Association of paraoxonase gene cluster polymorphisms with ALS in France, Quebec, and Sweden. Neurology. https://doi.org/10.1212/01.wnl.0000324997.21272.0c
Saeed M, Siddique N, Hung WY et al (2006) Paraoxonase cluster polymorphisms are associated with sporadic ALS. Neurology. https://doi.org/10.1212/01.wnl.0000227187.52002.88
Slowik A, Tomik B, Wolkow PP et al (2006) Paraoxonase gene polymorphisms and sporadic ALS. Neurology. https://doi.org/10.1212/01.wnl.0000219565.32247.11
Chen Y-Z, Bennett CL, Huynh HM et al (2004) DNA/RNA helicase gene mutations in a form of juvenile amyotrophic lateral sclerosis (ALS4). Am J Hum Genet. https://doi.org/10.1086/421054
Hirano M, Quinzii CM, Mitsumoto H et al (2011) Senataxin mutations and amyotrophic lateral sclerosis. Amyotroph Lateral Scler. https://doi.org/10.3109/17482968.2010.545952
Tripolszki K, Danis J, Padhi AK et al (2019) Angiogenin mutations in Hungarian patients with amyotrophic lateral sclerosis: clinical, genetic, computational, and functional analyses. Brain Behav. https://doi.org/10.1002/brb3.1293
Acknowledgments
Helpful suggestions from Prof B. Jayaram (IIT Delhi) are gratefully acknowledged.
Funding
The authors gratefully acknowledge the funding received from the Kusuma Trust UK and the financial and infrastructure support of the Indian Institute of Technology, Delhi for carrying out this research. DM, PN, and UD were supported by Research Fellowship from IIT Delhi, India.
Author information
Authors and Affiliations
Contributions
PN, AKP, VP, and JG conceived and designed the experiments. PN performed the experiments. AKP performed molecular dynamics simulations and analyses. PN, UD, and DM handled protein purification and assays. PN, AKP, UD, DM, VP, and JG analyzed the data. RB handled patient recruitment and clinical evaluation. PN, AKP, RB, VP, and JG contributed to the writing of the manuscript.
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Narain, P., Padhi, A.K., Dave, U. et al. Identification and characterization of novel and rare susceptible variants in Indian amyotrophic lateral sclerosis patients. Neurogenetics 20, 197–208 (2019). https://doi.org/10.1007/s10048-019-00584-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10048-019-00584-3