Abstract
Identification of key ionic channel contributors to the overall dynamics of a neuron is an important problem in experimental neuroscience. Such a problem is challenging since even in the best cases, identification relies on noisy recordings of membrane potential only, and strict inversion to the constituent channel dynamics is mathematically ill-posed. In this work, we develop a biophysically interpretable, learning-based strategy for data-driven inference of neuronal dynamics. In particular, we propose two optimization frameworks to learn and approximate neural dynamics from an observed voltage trajectory. In both the proposed strategies, the membrane potential dynamics are approximated as a weighted sum of ionic currents. In the first strategy, the ionic currents are represented using voltage dependent channel conductances and membrane potential in a parametric form, while in the second strategy, the currents are represented as a linear combination of generic basis functions. A library of channel activation/inactivation and time-constant curves describing prototypical channel kinetics are used to provide estimates of the channel variables to approximate the ionic currents. Finally, a linear optimization problem is solved to infer the weights/scaling variables in the membrane-potential dynamics. In the first strategy, the weights can be used to recover the channel conductances, and the reversal potentials while in the second strategy, using the estimated weights, active channels can be inferred and the trajectory of the gating variables are recovered, allowing for biophysically salient inference. Our results suggest that the complex nonlinear behavior of the neural dynamics over a range of temporal scales can be efficiently inferred in a data-driven manner from noisy membrane potential recordings.












Similar content being viewed by others
References
Ahrens, M., Paninski, L., Huys, Q.J. (2006). Large-scale biophysical parameter estimation in single neurons via constrained linear regression. In: Advances in neural information processing systems, pp. 25–32.
Berger, S.D., & Crook, S.M. (2015). Modeling the influence of ion channels on neuron dynamics in drosophila. Frontiers in computational neuroscience 9.
Buhry, L., Pace, M., Saïghi, S. (2012). Global parameter estimation of an hodgkin–huxley formalism using membrane voltage recordings: Application to neuro-mimetic analog integrated circuits. Neurocomputing, 81, 75–85.
Csercsik, D., Hangos, K.M., Szederkényi, G. (2012). Identifiability analysis and parameter estimation of a single hodgkin–huxley type voltage dependent ion channel under voltage step measurement conditions. Neurocomputing, 77(1), 178–188.
Dayan, P., & Abbott, L. (2001). Theoretical neuroscience: Computational and mathematical modeling of neural systems. Computational Neuroscience Series, Massachusetts Institute of Technology Press.
Doya, K., Selverston, A.I., Rowat, P.F. (1994). A hodgkin-huxley type neuron model that learns slow non-spike oscillation. In: Advances in neural information processing systems, pp. 566–573.
Drion, G., OLeary, T., Marder, E. (2015). Ion channel degeneracy enables robust and tunable neuronal firing rates. Proceedings of the National Academy of Sciences, 112(38), E5361–E5370.
Gerstner, W., & Naud, R. (2009). How good are neuron models? Science, 326(5951), 379–380. https://doi.org/10.1126/science.1181936.
Hamilton, F., Cressman, J., Peixoto, N., Sauer, T. (2014). Reconstructing neural dynamics using data assimilation with multiple models. EPL (Europhysics Letters), 107(6), 68,005.
Hodgkin, A.L., & Huxley, A.F. (1952). A quantitative description of membrane current and its application to conduction and excitation in nerve. The Journal of Physiology, 117(4), 500–544.
Izhikevich, E.M. (2003). Simple model of spiking neurons. IEEE Transactions on Neural Networks, 14(6), 1569–1572.
Izhikevich, E.M. (2007). Dynamical systems in neuroscience. MIT Press.
Lankarany, M., Zhu, W.P., Swamy, M. (2014). Joint estimation of states and parameters of hodgkin–huxley neuronal model using kalman filtering. Neurocomputing, 136, 289–299.
Lewis, F., Jagannathan, S., Yesildirak A. (1998). Neural network control of robot manipulators and non-linear systems. CRC Press.
Liao, F., Lou, X., Cui, B., Wu, W. (2016). State filtering and parameter estimation for hodgkin-huxley model. In: 2016 international joint conference on neural networks (IJCNN). IEEE, pp. 4658–4663.
Lynch, E.P., & Houghton, C.J. (2015). Parameter estimation of neuron models using in-vitro and in-vivo electrophysiological data. Frontiers in neuroinformatics 9.
Migliore, R., Lupascu, C.A., Bologna, L.L., Romani, A., Courcol, J.D., Antonel, S., Van Geit, W.A., Thomson, A.M., Mercer, A., Lange, S., et al. (2018). The physiological variability of channel density in hippocampal ca1 pyramidal cells and interneurons explored using a unified data-driven modeling workflow. PLoS Computational Biology, 14(9), e1006,423.
Narendra, K.S., & Annaswamy, A.M. (2012). Stable adaptive systems. Courier Corporation.
Narendra, K.S., & Parthasarathy, K. (1990). Identification and control of dynamical systems using neural networks. IEEE Transactions on Neural Networks, 1(1), 4–27.
Pospischil, M., Toledo-Rodriguez, M., Monier, C., Piwkowska, Z., Bal, T., Fregnac, Y., Markram, H., Destexhe, A. (2008). Minimal hodgkin–huxley type models for different classes of cortical and thalamic neurons. Biological Cybernetics, 99(4-5), 427–441.
Ritt, J.T., & Ching, S. (2015). Neurocontrol: Methods, models and technologies for manipulating dynamics in the brain. In: American control conference (ACC), 2015, IEEE, pp. 3765–3780.
Sastry, S., & Bodson, M. (2011). Adaptive control: stability, convergence and robustness. Courier Corporation.
Sinha, A., Schiff, S.J., Huebel, N. (2013). Estimation of internal variables from hodgkin-huxley neuron voltage. In: 2013 6th international IEEE/EMBS conference on neural engineering (NER), IEEE, pp. 194–197.
Ullah, G., & Schiff, S.J. (2009). Tracking and control of neuronal hodgkin-huxley dynamics. Physical Review E, 79(4), 040,901.
Van Geit, W., De Schutter, E., Achard, P. (2008). Automated neuron model optimization techniques: a review. Biological cybernetics, 99(4-5), 241–251.
Walch, O.J., & Eisenberg, M.C. (2016). Parameter identifiability and identifiable combinations in generalized hodgkin–huxley models. Neurocomputing, 199, 137–143.
Wang, G.J., & Beaumont, J. (2004). Parameter estimation of the hodgkin–huxley gating model: an inversion procedure. SIAM Journal on Applied Mathematics, 64(4), 1249–1267.
Willms, A.R., Baro, D.J., Harris-Warrick, R.M., Guckenheimer, J. (1999). An improved parameter estimation method for hodgkin-huxley models. Journal of computational neuroscience, 6(2), 145–168.
Acknowledgements
Funding: NIH 1R21-EY027590-01, NIH R01GM131403, NSF CMMI 1537015, NSF CMMI 1763070, NSF ECCS 1509342, NSF CAREER 1653589, ShiNung Ching Holds a Career Award at the Scientific Interface from the Burroughs-Wellcome Fund.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests
The authors declare that they have no conflict of interest.
Additional information
Action Editor: Gaute T. Einevoll
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix
Appendix
1.1 A.1 Computational procedure for strategy 1
We will describe strategy 1 using Example 1 without loss of generality. This procedure is also summarized as an algorithm (see Algorithm 1).
Library selection
We first constructed a library of the activation curves and the time-constant curves following Eq. (4) where the functional forms of \(\alpha _{n_{j}}(V)\) and \(\beta _{n_{j}}(V)\) were selected as given in Izhikevich (2007) with unknown parameters \(d,e,f,\bar {d},\bar {e},\bar {f}\). These functions are:
for the activation channels and
for the inactivation channels. The interval [Vb,Vth] is empirically observed to be [− 75, 30]mV with resting potential close to − 65mV (Fig. 13). Therefore, we constrained our design parameters
\(\bar {e} = e\), d ∈ (0, 5], f ∈ [1, 30], \(\bar {d}= d\), and \(\bar {f} = f\).
Schematic of Strategy I. a The voltage trace (red color) and its time derivative (blue color) are used to determine the function domain for the activation/inactivation and time-constant curves. b The voltage dependent functions \(\alpha _{n_{j}}(V)\) and \(\beta _{n_{j}}(V)\) are drawn from the literature and used to generate the library of voltage dependent activation curves and time-constant curves. Here, blue solid lines are the curves associated with activating channels while the red solid lines are the curves associated with inactivating channels. c The current Eq. (2) are forward simulated to produce candidate gating variable trajectories, i.e., m(t) and h(t). Here, blue solid lines are the curves associated with sampled gating variables obtained from the library while the red solid lines are the gating variables corresponding to the ground truth model of the 4th order HH model. d We sample a set of candidate gating variable trajectories. Regression is then performed to produce a candidate model
Channel pruning
The design parameters were uniformly randomly sampled from these intervals to form a set of \(\alpha _{n_{j}}(V)\), \(\beta _{n_{j}}(V)\). Because these functions are the channel opening and closing rates, we expect them to attain non-zero values over voltage ranges in which the observed voltage trace displays a non-zero derivative. In particular, we prune those functions that meet the criteria
and
After pruning, we are left with a set of nontrivial opening and closing rate functions. From these, we construct the final library of activation and time-constant via (4).
Gating variables
Using the constructed library, the channel dynamical Eq. (3) were forward simulated to obtain a set of corresponding gating variable trajectories (see Fig. 13). From this, we built a set of regression functions using these gating variable trajectories in the current Eq. (2). Here, the values of a,b in the current equations were constrained to be in the set {0, 1, 2, 3, 4}, and
Monte Carlo sampling and regression
To perform the regression and estimate the unknown parameters, we performed a Monte Carlo analysis. For Example 1, we used 150ms of (simulated) data with a sampling period of 0.01ms. Next, we picked in each iteration \(\phi (V,\hat {I}_{x},I) \in \mathbb {R}^{15000 \times 11}\) (l = 11), a subset of Φ(t), to make the regression problem sufficiently over-determined. These functions were picked randomly with uniform probability, at each iteration. Then we solved the linear regression problem as described in Eq. (11) using a standard SVD algorithm. During each iteration, the estimated parameters were recorded along with their residual/regression error (Rn) and the condition number (Cn) of the regression matrix (see Fig. 14). We performed this iterative procedure for 1000 iterations.
The procedure from Fig. 13 is repeated multiple times. a The Monte Carlo procedure produces many candidate models, each associated with a residual error Rn and condition number Cn. We select a model that minimizes error to within a standard deviation of the mean condition number, here indicated by the red arrow. b In the case of Example 1, we depict the final set of channels (red: passive; blue: active) among all candidate channels over Monte Carlo iterations
Model selection
The final model is selected as the one that minimizes the residual error to within one standard deviation of the mean condition number over all iterations.

1.2 A.2 Computational procedure for strategy 2
In strategy 2, the library selection step and the channel pruning steps are the same as described in Appendix A. We will describe the rest of the procedure next.
Gating variables
Using the library of activation/inactivation curves and time-constant curves, the channel dynamical Eq. (3) were forward simulated to obtain a set of corresponding gating variable trajectories. However, instead of using functional forms for the current equations as in strategy 1, we here use an (artificial) neural network architecture (Lewis et al. 1998) to transform the gating variable trajectories into currents.
Neural network (NN) design
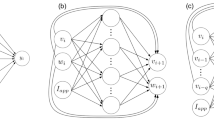
We use a random vector functional link network (Lewis et al. 1998) with random weights between the input and hidden layers. The weights from the hidden layer to the output layer Wx are free and to be regressed. The inputs to the NN were the voltage trajectory, the extrinsic current and the gating variables obtained from the previous step. We constructed the NN inputs in order to distinguish transient, persistent and leak currents. Specifically, we used three blocks of inputs : (i) {V (t),m(t),h(t)} (for approximating transient currents); (ii) {V (t),m(t)} (for approximating persistent currents); and (iii) V (t) (to approximate leak current) (see Fig. 10). The exogenous current I(t) was applied as the input to the output neuron scaled by a bias (B) which was free to be tuned.
Monte Carlo sampling and weight tuning
At each iteration, we picked six gating variables (five activation channels and one inactivation channel). We updated the weights of the hidden to output layer via linear regression as in strategy 1. We used 30 hidden layer neurons for each input block. We performed this analysis over 1500 iterations and retained the model that minimized the residual error.
1.3 A.3 Online learning rule and convergence analysis
Let the following equations describe the approximated neuronal dynamics as
where \(\hat {W}\) consists of the estimated parameters and ϕ is calculated using the estimated gating variables and measured voltage, \(\tilde {V}\) is the voltage estimation error, and \(\tilde {A}> 0\) is a design parameter. To bring the estimated parameters close to the true parameters, update the estimated parameters using the adaptation rule as follows: First define the performance measure as
Using the performance measure (20), define the learning rule as
with \(\alpha , \beta , h>0, \hat W(0)=\hat W_{0}\) and \(\tilde {V}(t)=V(t)-\hat {V}(t)\). Define the error in the estimated parameters as \(\tilde W(t) = W(t)-\hat {W}(t)\). To ensure persistency of excitation, the data (history) can be stacked to get the regression vector ϕ, voltage estimation error, \(\tilde {V}\) in the update equation (by using the historical data, V (0),…,V (t) and \(\hat {V}(0),\ldots , \hat {V}(t)\) ). Here, γ is the sigma modification term (Lewis et al. 1998), which performs the role of a regularizer, while avoiding parameter drift .
To analyse the convergence properties, consider a quadratic energy function,
Taking the time derivative, we get
Using the adaptation rule (21) and the approximated dynamcis (19), we have
This is simplified as
The slope of the energy function is thus bounded by
This implies that the voltage estimation error and the parameter estimation errors converge to a bound defined by the term \(\frac {\gamma ^{2}\epsilon }{2} W^{T}W\).
If we consider the voltage estimation error dynamics, \(\dot {\tilde {V}}(t)=\dot {V}(t)-\dot {\hat { V}}(t)\). Since we assumed that the exact channel dynamics are known in the analysis, the error dynamics is simplified as \(\dot {\tilde {V}}(t)=\tilde {W}^{T}(t)\phi (t)-\tilde {A}\tilde V(t)\). The linear term \(\tilde {A}\tilde V(t)\) is required to stabilize the model during the learning period. Furthermore, since the channel dynamics are not known, in general, we will have \(\dot {\tilde {V}}(t)=\tilde {W}^{T}(t)\phi (t)+\hat W(t)(\phi (t)-\hat {\phi }(t))-\tilde {A}\tilde V(t)\). Here the regression vector, \(\hat {\phi }(t)\), is calculated using a set of gating variables characterized by their activation curves, time-constants. As the errors in the gating variables converge, the regression vector, \(\hat {\phi }(t) \to \phi (t)\).
By representing the dynamics in parametric form, it can also be noticed that the current, I(t), is required to be a non-constant stimulus inorder for the matrix ϕ(t)ϕT(t) to be nonsingular for the existence of a solution to the linear regression problem (due to the presence of leak channel). This is a potential source for numerical errors in the linear regression problem as the regression matrix can be poorly conditioned.
In the second strategy, due to the approximation, there will be a reconstruction/approximation error, o(V,nj), i.e.,
Therefore, the voltage estimation error dynamics can be obtained in this case as, \(\dot {\tilde {V}}(t)=\tilde {W}^{T}(t)\phi (t)+o(V,n_{j})-\tilde {A}\tilde V(t)\). This approximation error will increase the error bound obtained earlier. By increasing the number of basis functions in the approximation, the approximation error can be reduced, however, at the expense of additional computations (Lewis et al. 1998). Since a random vector function link network is used, the regression problem is still linear.
It should also be noted that in the analysis above, the target parameters are assumed to be constants. The design parameters used in the learning algorithm (strategy 1) are α = 500,β = 750,γ = 0.0001, \(\tilde {A} = 0.01\) and α = 250,γ = 0.001,β = 350,A = 1 for strategy 2, yielded results that are similar to that presented in the Examples (but not shown).
Rights and permissions
About this article
Cite this article
Narayanan, V., Li, JS. & Ching, S. Biophysically interpretable inference of single neuron dynamics. J Comput Neurosci 47, 61–76 (2019). https://doi.org/10.1007/s10827-019-00723-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10827-019-00723-7