Abstract
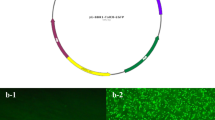
The prevalence of insect resistance against Bt toxins has led to the idea of enhancing demethylation from cell wall pectin by pectin methylesterase enzyme for overproduction of methanol which is toxic to insects pests. The AtPME and AnPME fragments ligated into pCAMBIA1301 vector were confirmed through restriction digestion with EcoR1 and BamH1. Excision of 3363 bp fragment from 11,850 bp vector confirmed the ligation of both fragments into pCAMBIA1301 vector. Transformation of pectin methylesterase-producing genes, i.e., AtPME and AnPME from Arabidopsis thaliana and Aspergillus niger cloned in plant expression vector pCAMBIA1301 under 35S promoter into cotton variety CEMB-33 harboring two Bt genes Cry1Ac and Cry2A, respectively, was done by using shoot apex-cut Agrobacterium-mediated transformation method. The plantlets were screened on MS medium supplemented with hygromycin on initial basis. Amplification of 412 and 543 bp, respectively, through gene-specific primer has been obtained which confirmed the successful introduction of pCAMBIA AtPME and AnPME genes into cotton variety CEMB 33. Relative expression of AtPME and AnPME genes through real-time PCR determined the expression level of both gene ranges between 3- and 3.5-fold in different transgenic cotton lines along with quantity of methanol ranging from 0.8 to 0.9% of maximum while 0.5% to 0.6% of minimum but no expression was obtained in negative non-transgenic control cotton plant with least quantity of methanol, i.e., 0.1%. Almost 100% mortality was observed in insect bioassay for Helicoverpa armigera on detached leaves bioassay and 63% for Pink Bollworm (Pectinophora gossypiella) on growing transgenic cotton bolls as compared to positive control transgenic cotton with double Bt genes where mortality was found to be 82% for H. armigera and 50% for P. gossypiella while 0% in negative control non-transgenic plants.










Similar content being viewed by others
References
Akram, A., Hussain, K., Nahid, N., Nasim, A. & Shaheen, S. (2017). Molecular characterization of a begomovirus and associated satellites from cotton (Gossypium hirsutum from Dera Ghazi Khan district of Pakistan. JAPS: Journal of Animal & Plant Sciences, 27(4), 1245–1255.
He, Z., Zhang, H., Tewolde, H. & Shankle, M. (2017). Chemical characterization of cotton plant parts for multiple uses. Agricultural & Environmental Letters. https://doi.org/10.2134/ael2016.11.0044.
Rani, S., Habib, N., Raza, I., & Zahra, N. (2017). Estimating compound growth rate, instability index and annual fluctuation of cotton in Pakistan. Asian Journal of Agriculture and rural Development, 7, 86–91.
Khan, M. J. (2017). Agriculture and textile sector: Challenges and way forward. https://peri.punjab.gov.pk/.
Akhtar, Z. R., Saeed, Z., Anjum, A. A., Hira, H., Ihsan, A., Noreen, A., & Salman, M. A. (2018). Pink boll worm resistance evaluation against organophosphate in Cry1Ac expressing transgenic cotton. Journal of Entomology and Zoology Studies, 6, 821–824.
Nareshkumar, B., Akbar, S. M., Sharma, H. C., Jayalakshmi, S. K., & Sreeramulu, K. (2018). Imidacloprid impedes mitochondrial function and induces oxidative stress in cotton bollworm, Helicoverpa armigera larvae (Hubner: Noctuidae). Journal of Bioenergetics and Biomembranes, 50, 21–32.
Akhtar, Z. R., Anjum, A. A., Saeed, Z., & Khalid, J. (2018). Resistance development in bollworms against Bt proteins deployed in genetically modified cotton. Journal of Entomology and Zoology Studies, 6, 1260–1264.
Rajendran, T., Birah, A., & Burange, P. S. (2018). Insect pests of cotton. Pests and their management (pp. 361–411). Berlin: Springer.
Kouser, S., & Qaim, M. (2015). Bt cotton, pesticide use and environmental efficiency in Pakistan. Journal of Agricultural Economics, 66, 66–86.
Verma, C., Tiwari, A. K. M., & Mishra, S. (2017). Biochemical and molecular characterization of cell wall degrading enzyme, pectin methylesterase versus banana ripening: An overview. Asian Journal of Biotechnology, 9, 1–23.
Carrière, Y., Williams, J. L., Crowder, D. W., & Tabashnik, B. E. (2018). Genotype-specific fitness cost of resistance to Bt toxin Cry1Ac in pink bollworm. Pest Management Science, 74, 2496–2503.
Gao, M., Wang, X., Yang, Y., Tabashnik, B. E., & Wu, Y. (2018). Epistasis confers resistance to Bt toxin Cry1Ac in the cotton bollworm. Evolutionary Applications, 11, 809–819.
Dev, S. (2017). Insecticides of natural origin. Abingdon: Routledge.
Dorokhov, Y. L., Shindyapina, A. V., Sheshukova, E. V., & Komarova, T. V. (2015). Metabolic methanol: Molecular pathways and physiological roles. Physiological Reviews, 95, 603–644.
Dixit, S., Upadhyay, S. K., Singh, H., Sidhu, O. P., Verma, P. C., & Chandrashekar, K. (2013). Enhanced methanol production in plants provides broad spectrum insect resistance. PLoS ONE, 8, e79664.
Hasunuma, T., Fukusaki, E.-I., & Kobayashi, A. (2003). Methanol production is enhanced by expression of an Aspergillus niger pectin methylesterase in tobacco cells. Journal of Biotechnology, 106, 45–52.
Gopal, A., & Benny, P. (2018). Selective comparison of repellent activity with Contact toxicity of certain essential oils against different insect orders. International Journal of Life Sciences, 6, 615–625.
Jirschitzka, J., Mattern, D. J., Gershenzon, J., & D’Auria, J. C. (2013). Learning from nature: New approaches to the metabolic engineering of plant defense pathways. Current Opinion in Biotechnology, 24, 320–328.
Von Dahl, C. C., Hävecker, M., Schlögl, R., & Baldwin, I. T. (2006). Caterpillar-elicited methanol emission: A new signal in plant–herbivore interactions? The Plant Journal, 46, 948–960.
Körner, E., von Dahl, C. C., Bonaventure, G., & Baldwin, I. T. (2009). Pectin methylesterase Na PME1 contributes to the emission of methanol during insect herbivory and to the elicitation of defence responses in Nicotiana attenuata. Journal of Experimental Botany, 60, 2631–2640.
Latif, A., Rao, A. Q., Khan, M. A. U., Shahid, N., Bajwa, K. S., Ashraf, M. A., et al. (2015). Herbicide-resistant cotton (Gossypium hirsutum) plants: An alternative way of manual weed removal. BMC Research Notes, 8, 453.
Gould, J. H., & Magallanes-Cedeno, M. (1998). Adaptation of cotton shoot apex culture to Agrobacterium-mediated transformation. Plant Molecular Biology Reporter, 16, 283.
Bajwa, K. S., Shahid, A. A., Rao, A. Q., Kiani, M. S., Ashraf, M. A., Dahab, A. A., et al. (2013). Expression of Calotropis procera expansin gene CpEXPA3 enhances cotton fibre strength. Australian Journal of Crop Science, 7, 206.
Puspito, A. N., Rao, A. Q., Hafeez, M. N., Iqbal, M. S., Bajwa, K. S., Ali, Q., et al. (2015). Transformation and evaluation of Cry1Ac + Cry2A and GTGene in Gossypium hirsutum L. Frontiers in plant science, 6, 943.
Muoki, R. C., Paul, A., Kumari, A., Singh, K., & Kumar, S. (2012). An improved protocol for the isolation of RNA from roots of tea (Camellia sinensis (L.) O. Kuntze). Molecular Biotechnology, 52, 82–88.
Stylianou, E., Harrington-Kandt, R., Beglov, J., Bull, N., Pinpathomrat, N., Swarbrick, G. M., Lewinsohn, D. A., Lewinsohn, D. M. & McShane, H. (2018). Identification and Evaluation of Novel Protective Antigens for the Development of a Candidate TB Subunit Vaccine. Infection and Immunity, IAI. 00014-00018.
Basit, M. (2018). Cotton pest control awareness in farmers of the Punjab, Pakistan and its impact on whitefly resistance against available insecticides. Phytoparasitica, 46, 183–195.
Sarwar, M., & Sattar, M. (2016). An analysis of comparative efficacies of various insecticides on the densities of important insect pests and the natural enemies of cotton, Gossypium hirsutum L. Pakistan Journal of Zoology, 48, 131–136.
Ramani, S. V., Thutupalli, A., El-Aroui, M.-A., & Kumar, P. (2017). MNE led new market creation in emerging countries: The case of Bt cotton. Multinational enterprises and sustainable development (pp. 131–150). Emerald Publishing Limited: Bingley.
Awan, M. F., Abbas, M. A., Muzaffar, A., Ali, A., Tabassum, B., Rao, A. Q., et al. (2015). Transformation of insect and herbicide resistance genes in cotton (Gossypium hirsutum L.). Journal of Agricultural Science & Technology, 17(2), 287–298.
Salisu, I. B., Shahid, A. A., Ali, Q., Rao, A. Q., & Husnain, T. (2018). Nutritional assessment of dietary Bt and CP4EPSPS proteins on the serum biochemical changes of rabbits at different developmental stages. Frontiers in Nutrition, 5, 49.
Ali, A., Ahmed, S., Nasir, I. A., Rao, A. Q., Ahmad, S., & Husnain, T. (2016). Evaluation of two cotton varieties CRSP1 and CRSP2 for genetic transformation efficiency, expression of transgenes Cry1Ac + Cry2A, GT gene and insect mortality. Biotechnology Reports, 9, 66–73.
Singh, A. K., & Pental, D. (2015). Selection and genetic transformation of a fast-growing cell line in cotton (Gossypium hirsutum) for transgene expression studies. Journal of Plant Biochemistry and Biotechnology, 24, 225–232.
Shahid, A. A. (2017). A Novel approach to study relationship between Cotton Leaf Curl Virus (CLCuV) and Beta Satellites. International Journal Biotechnology & Bioengineering, 3, 230–235.
Yasmeen, A., Kiani, S., Butt, A., Rao, A. Q., Akram, F., Ahmad, A., et al. (2016). Amplicon-based RNA interference targeting V2 gene of cotton leaf curl Kokhran Virus-Burewala strain can provide resistance in transgenic cotton plants. Molecular Biotechnology, 58, 807–820.
Khan, M., Bajwa, K. S., Husnain, T., Rao, A. Q., Puspito, A. N., Bakhsh, A., et al. (2013). Variation in expression of phytochrome B gene in cotton (Gossypium hirsutum L.). Journal of Agriculture Science and Technology, 15, 1033–1042.
Khan, A., Nasir, I. A., Tabassum, B., Aaliya, K., Tariq, M., & Rao, A. Q. (2017). Expression studies of chitinase gene in transgenic potato against Alternaria solani. Plant Cell, Tissue and Organ Culture (PCTOC), 128, 563–576.
Cseke, L. J., Kirakosyan, A., Kaufman, P. B., Warber, S., Duke, J. A., & Brielmann, H. L. (2016). Natural products from plants. Boca Raton: CRC Press.
Aslam, M. (2014). Emission and role of biogenic volatile organic compounds in biosphere. A Journal of Science for Development, 19, 69.
Matsui, K. (2016). A portion of plant airborne communication is endorsed by uptake and metabolism of volatile organic compounds. Current Opinion in Plant Biology, 32, 24–30.
Rao, A. Q., Irfan, M., Saleem, Z., Nasir, I. A., Riazuddin, S., & Husnain, T. (2011). Overexpression of the phytochrome B gene from Arabidopsis thaliana increases plant growth and yield of cotton (Gossypium hirsutum). Journal of Zhejiang University Science B, 12, 326–334.
Sivasupramaniam, S., Moar, W., Ruschke, L., Osborn, J., Jiang, C., Sebaugh, J., et al. (2014). Toxicity and characterization of cotton expressing Bacillus thuringiensis Cry1Ac and Cry2Ab2 proteins for control of lepidopteran pests. Journal of Economic Entomology, 101, 546–554.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zubair, M., Latif, A., Rao, A.Q. et al. A Combinational Approach of Enhanced Methanol Production and Double Bt Genes for Broad Spectrum Insect Resistance in Transgenic Cotton. Mol Biotechnol 61, 663–673 (2019). https://doi.org/10.1007/s12033-019-00192-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-019-00192-4