当前位置:
X-MOL 学术
›
ACS Chem. Biol.
›
论文详情
Our official English website, www.x-mol.net, welcomes your feedback! (Note: you will need to create a separate account there.)
Structural Characterization of Degrader-Induced Ternary Complexes Using Hydrogen–Deuterium Exchange Mass Spectrometry and Computational Modeling: Implications for Structure-Based Design
ACS Chemical Biology ( IF 4 ) Pub Date : 2021-09-28 , DOI: 10.1021/acschembio.1c00376 Scott J Eron 1 , Hongwei Huang 1 , Roman V Agafonov 1 , Mark E Fitzgerald 1 , Joe Patel 1 , Ryan E Michael 1 , Tobie D Lee 1 , Ashley A Hart 1 , Jodi Shaulsky 2 , Christopher G Nasveschuk 1 , Andrew J Phillips 1 , Stewart L Fisher 1 , Andrew Good 1
ACS Chemical Biology ( IF 4 ) Pub Date : 2021-09-28 , DOI: 10.1021/acschembio.1c00376 Scott J Eron 1 , Hongwei Huang 1 , Roman V Agafonov 1 , Mark E Fitzgerald 1 , Joe Patel 1 , Ryan E Michael 1 , Tobie D Lee 1 , Ashley A Hart 1 , Jodi Shaulsky 2 , Christopher G Nasveschuk 1 , Andrew J Phillips 1 , Stewart L Fisher 1 , Andrew Good 1
Affiliation
|
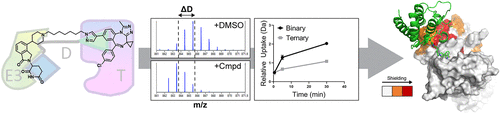
The field of targeted protein degradation (TPD) has grown exponentially over the past decade with the goal of developing therapies that mark proteins for destruction leveraging the ubiquitin–proteasome system. One common approach to achieve TPD is to employ a heterobifunctional molecule, termed as a degrader, to recruit the protein target of interest to the E3 ligase machinery. The resultant generation of an intermediary ternary complex (target–degrader–ligase) is pivotal in the degradation process. Understanding the ternary complex geometry offers valuable insight into selectivity, catalytic efficiency, linker chemistry, and rational degrader design. In this study, we utilize hydrogen–deuterium exchange mass spectrometry (HDX-MS) to identify degrader-induced protein–protein interfaces. We then use these data in conjunction with constrained protein docking to build three-dimensional models of the ternary complex. The approach was used to characterize complex formation between the E3 ligase CRBN and the first bromodomain of BRD4, a prominent oncology target. We show marked differences in the ternary complexes formed in solution based on distinct patterns of deuterium uptake for two degraders, CFT-1297 and dBET6. CFT-1297, which exhibited positive cooperativity, altered the deuterium uptake profile revealing the degrader-induced protein–protein interface of the ternary complex. For CFT-1297, the ternary complexes generated by the highest scoring HDX-constrained docking models differ markedly from those observed in the published crystal structures. These results highlight the potential utility of HDX-MS to provide rapidly accessible structural insights into degrader-induced protein–protein interfaces in solution. They further suggest that degrader ternary complexes exhibit significant conformation flexibility and that biologically relevant complexes may well not exhibit the largest interaction surfaces between proteins. Taken together, the results indicate that methods capable of incorporating linker conformation uncertainty may prove an important component in degrader design moving forward. In addition, the development of scoring functions modified to handle interfaces with no evolved complementarity, for example, through consideration of high levels of water infiltration, may prove valuable. Furthermore, the use of crystal structures as validation tools for novel degrader methods needs to be considered with caution.
中文翻译:

使用氢-氘交换质谱法和计算建模对降解剂诱导的三元配合物进行结构表征:对基于结构的设计的意义
靶向蛋白质降解 (TPD) 领域在过去十年中呈指数级增长,目标是开发利用泛素-蛋白酶体系统标记蛋白质以进行破坏的疗法。实现 TPD 的一种常见方法是使用称为降解剂的异双功能分子将感兴趣的蛋白质靶标招募到 E3 连接酶机制。由此产生的中间三元复合物(目标-降解剂-连接酶)在降解过程中至关重要。了解三元复杂的几何形状为选择性、催化效率、接头化学和合理的降解剂设计提供了宝贵的见解。在这项研究中,我们利用氢-氘交换质谱 (HDX-MS) 来识别降解剂诱导的蛋白质-蛋白质界面。然后,我们将这些数据与受约束的蛋白质对接结合使用来构建三元复合物的三维模型。该方法用于表征 E3 连接酶 CRBN 和 BRD4 的第一个溴结构域之间的复合物形成,BRD4 是一个突出的肿瘤学靶标。基于两种降解剂 CFT-1297 和 dBET6 的不同氘吸收模式,我们显示了溶液中形成的三元配合物的显着差异。CFT-1297 表现出积极的协同性,改变了氘吸收曲线,揭示了降解剂诱导的三元复合物的蛋白质-蛋白质界面。对于 CFT-1297,得分最高的 HDX 约束对接模型生成的三元配合物与已发表的晶体结构中观察到的明显不同。这些结果突出了 HDX-MS 的潜在实用性,可以为溶液中降解剂诱导的蛋白质-蛋白质界面提供快速可访问的结构洞察力。他们进一步表明,降解三元复合物表现出显着的构象灵活性,并且生物学相关的复合物可能不会表现出蛋白质之间最大的相互作用表面。总之,结果表明,能够结合连接器构象不确定性的方法可能被证明是降解器设计向前发展的重要组成部分。此外,改进评分函数的开发以处理没有进化互补性的界面,例如,通过考虑高水平的水渗透,可能证明是有价值的。此外,
更新日期:2021-11-19
中文翻译:

使用氢-氘交换质谱法和计算建模对降解剂诱导的三元配合物进行结构表征:对基于结构的设计的意义
靶向蛋白质降解 (TPD) 领域在过去十年中呈指数级增长,目标是开发利用泛素-蛋白酶体系统标记蛋白质以进行破坏的疗法。实现 TPD 的一种常见方法是使用称为降解剂的异双功能分子将感兴趣的蛋白质靶标招募到 E3 连接酶机制。由此产生的中间三元复合物(目标-降解剂-连接酶)在降解过程中至关重要。了解三元复杂的几何形状为选择性、催化效率、接头化学和合理的降解剂设计提供了宝贵的见解。在这项研究中,我们利用氢-氘交换质谱 (HDX-MS) 来识别降解剂诱导的蛋白质-蛋白质界面。然后,我们将这些数据与受约束的蛋白质对接结合使用来构建三元复合物的三维模型。该方法用于表征 E3 连接酶 CRBN 和 BRD4 的第一个溴结构域之间的复合物形成,BRD4 是一个突出的肿瘤学靶标。基于两种降解剂 CFT-1297 和 dBET6 的不同氘吸收模式,我们显示了溶液中形成的三元配合物的显着差异。CFT-1297 表现出积极的协同性,改变了氘吸收曲线,揭示了降解剂诱导的三元复合物的蛋白质-蛋白质界面。对于 CFT-1297,得分最高的 HDX 约束对接模型生成的三元配合物与已发表的晶体结构中观察到的明显不同。这些结果突出了 HDX-MS 的潜在实用性,可以为溶液中降解剂诱导的蛋白质-蛋白质界面提供快速可访问的结构洞察力。他们进一步表明,降解三元复合物表现出显着的构象灵活性,并且生物学相关的复合物可能不会表现出蛋白质之间最大的相互作用表面。总之,结果表明,能够结合连接器构象不确定性的方法可能被证明是降解器设计向前发展的重要组成部分。此外,改进评分函数的开发以处理没有进化互补性的界面,例如,通过考虑高水平的水渗透,可能证明是有价值的。此外,

























 京公网安备 11010802027423号
京公网安备 11010802027423号