当前位置:
X-MOL 学术
›
J. Phys. Chem. B
›
论文详情
Our official English website, www.x-mol.net, welcomes your feedback! (Note: you will need to create a separate account there.)
Systematic Detection and Characterization of Hydrogen Bonding in Proteins via Local Vibrational Modes
The Journal of Physical Chemistry B ( IF 3.3 ) Pub Date : 2021-03-05 , DOI: 10.1021/acs.jpcb.0c11392 Niraj Verma 1 , Yunwen Tao 1 , Elfi Kraka 1
The Journal of Physical Chemistry B ( IF 3.3 ) Pub Date : 2021-03-05 , DOI: 10.1021/acs.jpcb.0c11392 Niraj Verma 1 , Yunwen Tao 1 , Elfi Kraka 1
Affiliation
|
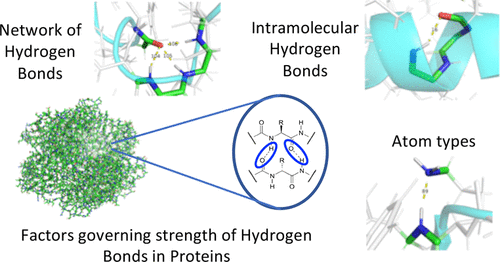
We introduce a new software, Efficient Detection of Hydrogen Bonds (EDHB), that systematically detects hydrogen bonds based on the nearest neighbors algorithm. EDHB classifies inter- and intramolecular hydrogen bonds as well as hydrogen bond networks. EDHB outperforms commonly used hydrogen bond detection methods in terms of speed of execution. An important additional feature of EDHB is that information from preceding quantum chemical studies (i.e., natural bond orbital analysis data and second energy derivative information) can be used to determine the electrostatic/covalent character of the hydrogen bonds and to calculate local-mode hydrogen bond force constants as a quantitative measure of their intrinsic strength. We applied EDHB to a diverse set of 163 proteins. We identified hydrogen bond networks forming intramolecular rings of different sizes as a common feature playing an important role for specific secondary structure orientations such as α-helixes and turns. However, these networks do not have a significant influence on the hydrogen bond strength. Our comprehensive local-mode analysis reveals the interesting result that the hydrogen bond angle is the governing factor determining the hydrogen bond strength in a protein. EDHB offers a broad range of application possibilities. In addition to proteins, EDHB can be generally used to detect and characterize hydrogen bonds in protein–ligand interactions, water clusters, and other systems where a hydrogen bond plays a critical role, as well as during molecular dynamics simulations. The program is freely available at https://github.com/ekraka/EDHB.
中文翻译:

通过局部振动模式对蛋白质中氢键的系统检测和表征
我们推出了一个新软件,氢键的有效检测(EDHB),可根据最近邻居算法系统地检测氢键。EDHB对分子间和分子内氢键以及氢键网络进行分类。EDHB在执行速度方面胜过常用的氢键检测方法。EDHB的另一个重要重要特征是,先前的量子化学研究信息(即自然键轨道分析数据和第二能量导数信息)可用于确定氢键的静电/共价特征并计算局部模式氢键力常数作为其内在强度的定量度量。我们将EDHB应用到了163种蛋白质中。我们确定形成不同大小的分子内环的氢键网络是一个共同的特征,对于特定的二级结构取向(例如α-螺旋和转角)起着重要的作用。但是,这些网络对氢键强度没有重大影响。我们全面的局部模式分析揭示了有趣的结果,即氢键角是决定蛋白质中氢键强度的决定因素。EDHB提供了广泛的应用可能性。除蛋白质外,EDHB通常还可用于检测和表征蛋白质-配体相互作用,水团簇和其他系统中氢键起关键作用的分子中的氢键,以及分子动力学模拟过程中的氢键。该程序可从https://github.com/ekraka/EDHB免费获得。
更新日期:2021-03-18
中文翻译:

通过局部振动模式对蛋白质中氢键的系统检测和表征
我们推出了一个新软件,氢键的有效检测(EDHB),可根据最近邻居算法系统地检测氢键。EDHB对分子间和分子内氢键以及氢键网络进行分类。EDHB在执行速度方面胜过常用的氢键检测方法。EDHB的另一个重要重要特征是,先前的量子化学研究信息(即自然键轨道分析数据和第二能量导数信息)可用于确定氢键的静电/共价特征并计算局部模式氢键力常数作为其内在强度的定量度量。我们将EDHB应用到了163种蛋白质中。我们确定形成不同大小的分子内环的氢键网络是一个共同的特征,对于特定的二级结构取向(例如α-螺旋和转角)起着重要的作用。但是,这些网络对氢键强度没有重大影响。我们全面的局部模式分析揭示了有趣的结果,即氢键角是决定蛋白质中氢键强度的决定因素。EDHB提供了广泛的应用可能性。除蛋白质外,EDHB通常还可用于检测和表征蛋白质-配体相互作用,水团簇和其他系统中氢键起关键作用的分子中的氢键,以及分子动力学模拟过程中的氢键。该程序可从https://github.com/ekraka/EDHB免费获得。

























 京公网安备 11010802027423号
京公网安备 11010802027423号